Abstract
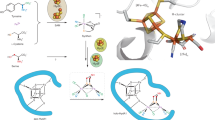
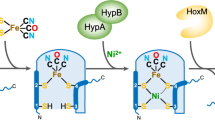
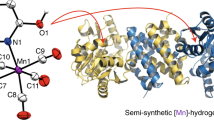
Hydrogenases catalyze the reversible oxidation of H2 into protons and electrons and are usually readily inactivated by O2. However, a subgroup of the [NiFe] hydrogenases, including the membrane-bound [NiFe] hydrogenase from Ralstonia eutropha, has evolved remarkable tolerance toward O2 that enables their host organisms to utilize H2 as an energy source at high O2. This feature is crucially based on a unique six cysteine-coordinated [4Fe-3S] cluster located close to the catalytic center, whose properties were investigated in this study using a multidisciplinary approach. The [4Fe-3S] cluster undergoes redox-dependent reversible transformations, namely iron swapping between a sulfide and a peptide amide N. Moreover, our investigations unraveled the redox-dependent and reversible occurence of an oxygen ligand located at a different iron. This ligand is hydrogen bonded to a conserved histidine that is essential for H2 oxidation at high O2. We propose that these transformations, reminiscent of those of the P-cluster of nitrogenase, enable the consecutive transfer of two electrons within a physiological potential range.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fontecilla-Camps, J.C., Volbeda, A., Cavazza, C. & Nicolet, Y. Structure/function relationships of [NiFe]- and [FeFe]-hydrogenases. Chem. Rev. 107, 4273–4303 (2007).
Fritsch, J., Lenz, O. & Friedrich, B. Structure, function and biosynthesis of O2-tolerant hydrogenases. Nat. Rev. Microbiol. 11, 106–114 (2013).
Schwartz, E., Fritsch, J. & Friedrich, B. in The Prokaryotes (eds. Eugene Rosenberg et al.) 119–199 (Springer Berlin Heidelberg, 2013).
Lenz, O. et al. H2 conversion in the presence of O2 as performed by the membrane-bound [NiFe]-hydrogenase of Ralstonia eutropha. ChemPhysChem 11, 1107–1119 (2010).
Bernhard, M., Benelli, B., Hochkoeppler, A., Zannoni, D. & Friedrich, B. Functional and structural role of the cytochrome b subunit of the membrane-bound hydrogenase complex of Alcaligenes eutrophus H16. Eur. J. Biochem. 248, 179–186 (1997).
Frielingsdorf, S., Schubert, T., Pohlmann, A., Lenz, O. & Friedrich, B. A trimeric supercomplex of the oxygen-tolerant membrane-bound [NiFe]-hydrogenase from Ralstonia eutropha H16. Biochemistry 50, 10836–10843 (2011).
Goris, T. et al. A unique iron-sulfur cluster is crucial for oxygen tolerance of a [NiFe]-hydrogenase. Nat. Chem. Biol. 7, 310–318 (2011).
Fritsch, J. et al. The crystal structure of an oxygen-tolerant hydrogenase uncovers a novel iron-sulphur centre. Nature 479, 249–252 (2011).
Shomura, Y., Yoon, K.S., Nishihara, H. & Higuchi, Y. Structural basis for a [4Fe-3S] cluster in the oxygen-tolerant membrane-bound [NiFe]-hydrogenase. Nature 479, 253–256 (2011).
Volbeda, A. et al. X-ray crystallographic and computational studies of the O2-tolerant [NiFe]-hydrogenase 1 from Escherichia coli. Proc. Natl. Acad. Sci. USA 109, 5305–5310 (2012).
Knüttel, K. et al. Redox properties of the metal centers in the membrane-bound hydrogenase from Alcaligenes eutrophus CH34. Bul. Pol. Acad. Sci. Chem. 42, 495–511 (1994).
Pandelia, M.E. et al. Characterization of a unique [FeS] cluster in the electron transfer chain of the oxygen tolerant [NiFe] hydrogenase from Aquifex aeolicus. Proc. Natl. Acad. Sci. USA 108, 6097–6102 (2011).
Schneider, K., Patil, D.S. & Cammack, R. ESR properties of membrane-bound hydrogenases from aerobic hydrogen bacteria. Biochim. Biophys. Acta 748, 353–361 (1983).
Cammack, R. “Super-reduction” of chromatium high-potential iron-sulphur protein in the presence of dimethyl sulphoxide. Biochem. Biophys. Res. Commun. 54, 548–554 (1973).
Cracknell, J.A., Wait, A.F., Lenz, O., Friedrich, B. & Armstrong, F.A. A kinetic and thermodynamic understanding of O2 tolerance in [NiFe]-hydrogenases. Proc. Natl. Acad. Sci. USA 106, 20681–20686 (2009).
Lauterbach, L. & Lenz, O. Catalytic production of hydrogen peroxide and water by oxygen-tolerant [NiFe]-hydrogenase during H2 cycling in the presence of O2 . J. Am. Chem. Soc. 135, 17897–17905 (2013).
Saggu, M. et al. Spectroscopic insights into the oxygen-tolerant membrane-associated [NiFe] hydrogenase of Ralstonia eutropha H16. J. Biol. Chem. 284, 16264–16276 (2009).
Ogata, H. et al. Activation process of [NiFe] hydrogenase elucidated by high-resolution X-ray analyses: conversion of the ready to the unready state. Structure 13, 1635–1642 (2005).
Rippers, Y., Horch, M., Hildebrandt, P., Zebger, I. & Mroginski, M.A. Revealing the absolute configuration of the CO and CN− ligands at the active site of a [NiFe] hydrogenase. ChemPhysChem 13, 3852–3856 (2012).
Ishikita, H. & Saito, K. Proton transfer reactions and hydrogen-bond networks in protein environments. J. R. Soc. Interface 11, 20130518 (2014).
Cross, M. et al. Removal of a cysteine ligand from rubredoxin: assembly of Fe2S2 and Fe(S-Cys)3(OH) centres. J. Biol. Inorg. Chem. 7, 781–790 (2002).
Sanders-Loehr, J. Resonance Raman-spectroscopy of iron-oxo and iron-sulfur clusters in proteins. ACS Symp. Ser. 372, 49–67 (1988).
Siebert, E. et al. Resonance Raman spectroscopy as a tool to monitor the active site of hydrogenases. Angew. Chem. Int. Edn. Engl. 52, 5162–5165 (2013).
Green, M.T. Application of Badger's rule to heme and non-heme iron-oxygen bonds: an examination of ferryl protonation states. J. Am. Chem. Soc. 128, 1902–1906 (2006).
Dikanov, S.A. & Tsvetkov, Y.D. Electron Spin Echo Envelope Modulation (ESEEM) Spectroscopy. (CRC Press, 1992).
Pandelia, M.E., Infossi, P., Stein, M., Giudici-Orticoni, M.T. & Lubitz, W. Spectroscopic characterization of the key catalytic intermediate Ni-C in the O2-tolerant [NiFe] hydrogenase I from Aquifex aeolicus: evidence of a weakly bound hydride. Chem. Commun. (Camb.) 48, 823–825 (2012).
Saggu, M. et al. Comparison of the membrane-bound [NiFe] hydrogenases from R. eutropha H16 and D. vulgaris Miyazaki F in the oxidized ready state by pulsed EPR. Phys. Chem. Chem. Phys. 12, 2139–2148 (2010).
Maly, T. & Prisner, T.F. Relaxation filtered hyperfine spectroscopy (REFINE). J. Magn. Reson. 170, 88–96 (2004).
Edmonds, D.T. & Summers, C.P. 14N pure quadrupole resonance in solid amino-acids. J. Magn. Reson. 12, 134–142 (1973).
Cammack, R. & MacMillan, F. in Metals in Biology Vol. 29 (eds. Hanson, G. & Berliner, L.) 11–44 (Springer New York, 2010).
Roessler, M.M., Evans, R.M., Davies, R.A., Harmer, J. & Armstrong, F.A. EPR spectroscopic studies of the Fe-S clusters in the O2-tolerant [NiFe]-hydrogenase Hyd-1 from Escherichia coli and characterization of the unique [4Fe-3S] cluster by HYSCORE. J. Am. Chem. Soc. 134, 15581–15594 (2012).
DeRose, V.J., Telser, J., Anderson, M.E., Lindahl, P.A. & Hoffman, B.M. A multinuclear ENDOR study of the C-cluster in CO dehydrogenase from Clostridium thermoaceticum: Evidence for HxO and histidine coordination to the [Fe4S4] center. J. Am. Chem. Soc. 120, 8767–8776 (1998).
Shergill, J.K. & Cammack, R. ESEEM and ENDOR studies of the Rieske iron-sulphur protein in bovine heart mitochondrial membranes. Biochim. Biophys. Acta 1185, 35–42 (1994).
Fritscher, J. Influence of hydrogen bond geometry on quadrupole coupling parameters: a theoretical study of imidazole-water and imidazole-semiquinone complexes. Phys. Chem. Chem. Phys. 6, 4950–4956 (2004).
Pelmenschikov, V. & Kaupp, M. Redox-dependent structural transformations of the [4Fe-3S] proximal cluster in O2-tolerant membrane-bound [NiFe]-hydrogenase: a DFT study. J. Am. Chem. Soc. 135, 11809–11823 (2013).
Ludwig, M., Cracknell, J.A., Vincent, K.A., Armstrong, F.A. & Lenz, O. Oxygen-tolerant H2 oxidation by membrane-bound [NiFe] hydrogenases of Ralstonia species. Coping with low level H2 in air. J. Biol. Chem. 284, 465–477 (2009).
Schubert, T., Lenz, O., Krause, E., Volkmer, R. & Friedrich, B. Chaperones specific for the membrane-bound [NiFe]-hydrogenase interact with the Tat signal peptide of the small subunit precursor in Ralstonia eutropha H16. Mol. Microbiol. 66, 453–467 (2007).
Abou Hamdan, A. et al. Understanding and tuning the catalytic bias of hydrogenase. J. Am. Chem. Soc. 134, 8368–8371 (2012).
Peters, J.W. et al. Redox-dependent structural changes in the nitrogenase P-cluster. Biochemistry 36, 1181–1187 (1997).
Huang, W. et al. Crystal structure of nitrile hydratase reveals a novel iron centre in a novel fold. Structure 5, 691–699 (1997).
Noveron, J.C., Olmstead, M.M. & Mascharak, P.K. Effect of carboxamido N coordination to iron on the redox potential of low-spin non-heme iron centers with N,S coordination: relevance to the iron site of nitrile hydratase. Inorg. Chem. 37, 1138–1139 (1998).
Abou Hamdan, A. et al. O2-independent formation of the inactive states of NiFe hydrogenase. Nat. Chem. Biol. 9, 15–17 (2013).
Carepo, M. et al. 17O ENDOR detection of a solvent-derived Ni-(OHx)-Fe bridge that is lost upon activation of the hydrogenase from Desulfovibrio gigas. J. Am. Chem. Soc. 124, 281–286 (2002).
Bruska, M.K., Stiebritz, M.T. & Reiher, M. Analysis of differences in oxygen sensitivity of Fe-S clusters. Dalton Trans. 42, 8729–8735 (2013).
Johnson, R.E., Papaefthymiou, G.C., Frankel, R.B. & Holm, R.H. Effects of secondary bonding interactions on the [Fe4S4]2+ core of ferredoxin site analogs: [Fe4S4(SC6H4-o-OH)4]2−, a distorted cubane-type cluster with one five-coordinate iron atom. J. Am. Chem. Soc. 105, 7280–7287 (1983).
Pandelia, M.E. et al. Electronic structure of the unique [4Fe-3S] cluster in O2-tolerant hydrogenases characterized by 57Fe Mössbauer and EPR spectroscopy. Proc. Natl. Acad. Sci. USA 110, 483–488 (2013).
Lanzilotta, W.N., Christiansen, J., Dean, D.R. & Seefeldt, L.C. Evidence for coupled electron and proton transfer in the [8Fe-7S] cluster of nitrogenase. Biochemistry 37, 11376–11384 (1998).
Matias, P.M. et al. [NiFe] hydrogenase from Desulfovibrio desulfuricans ATCC 27774: gene sequencing, three-dimensional structure determination and refinement at 1.8 Å and modelling studies of its interaction with the tetrahaem cytochrome c3 . J. Biol. Inorg. Chem. 6, 63–81 (2001).
Ogata, H., Kellers, P. & Lubitz, W. The crystal structure of the [NiFe] hydrogenase from the photosynthetic bacterium Allochromatium vinosum: characterization of the oxidized enzyme (Ni-A state). J. Mol. Biol. 402, 428–444 (2010).
Wallbank, B., Johnson, C.E. & Main, I.G. Surface reduction of solid 3d transition metal compounds during X-ray photoelectron spectroscopy. J. Electron Spectrosc. Relat. Phenom. 4, 263–269 (1974).
Ball, E.G. Studies on oxidation-reduction XXIII. Ascorbic acid. J. Biol. Chem. 118, 219–239 (1937).
Mueller, U. et al. Facilities for macromolecular crystallography at the Helmholtz-Zentrum Berlin. J. Synchrotron Radiat. 19, 442–449 (2012).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
CCP4. The CCP4 suite programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Kleihues, L., Lenz, O., Bernhard, M., Buhrke, T. & Friedrich, B. The H2 sensor of Ralstonia eutropha is a member of the subclass of regulatory [NiFe] hydrogenases. J. Bacteriol. 182, 2716–2724 (2000).
Schäfer, C., Friedrich, B. & Lenz, O. Novel, oxygen-insensitive group 5 [NiFe]-hydrogenase in Ralstonia eutropha. Appl. Environ. Microbiol. 79, 5137–5145 (2013).
Stoll, S. & Schweiger, A. EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J. Magn. Reson. 178, 42–55 (2006).
Vignais, P.M. H/D exchange reactions and mechanistic aspects of the hydrogenases. Coord. Chem. Rev. 249, 1677–1690 (2005).
Acknowledgements
We are grateful to D. von Stetten and A. Royant of the ID29S-Cryobench (European Synchrotron Radiation Facility (ESRF), Grenoble) and the scientific staff of the ESRF at beamlines ID14-1, ID 14-4, ID29, ID 23-1 and ID23-2 and to U. Müller, M. Weiss and the scientific staff of the BESSY-MX/Helmholtz-Zentrum Berlin für Materialien und Energie at beamlines BL 14.1, BL 14.2 and BL 14.3 operated by the Joint Berlin MX-Laboratory at the BESSY II electron storage ring (Berlin-Adlershof, Germany), where the data were collected, for continuous support. We thank C. Müller-Dieckmann for helpful advice in collecting anomalous data at the ID29 (ESRF, Grenoble). J. Hamann is acknowledged for excellent technical support in molecular biology. J. Newie, S. Wohlfahrt and T. Goris are acknowledged for their help in constructing MBH variants. This work was supported by the German Research Foundation (DFG) Cluster of Excellence 'Unifying Concepts in Catalysis' (to P.S., O.L., P.H., V.P. and R.B.). P.S. acknowledges K.P. Hofmann and his advanced investigator European Research Council grant (ERC-2009/249910-TUDOR) for support.
Author information
Authors and Affiliations
Contributions
S.F. and J.F. prepared MBH samples. P.S., J.F. and S.F. optimized MBH crystallization under different conditions and performed soaking experiments. P.S. collected the X-ray diffraction data, performed data processing and solved the MBH structures. A.S., M.H., J.K. and P.S. refined the crystal structures. J.L., C.T., F.L. and R.B. recorded and analyzed EPR data. E.S., I.Z. and P.H. recorded and analyzed RR spectroscopy data. V.P., Y.R. and M.K. performed and analyzed the DFT calculations. S.F. performed and analyzed H-D exchange measurements. T.J. and O.L. constructed the MBH mutant proteins. S.F., J.F., O.L. and P.S. wrote the paper with contributions from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–14, Supplementary Tables 1–5 and Supplementary Notes 1–3. (PDF 12530 kb)
Rights and permissions
About this article
Cite this article
Frielingsdorf, S., Fritsch, J., Schmidt, A. et al. Reversible [4Fe-3S] cluster morphing in an O2-tolerant [NiFe] hydrogenase. Nat Chem Biol 10, 378–385 (2014). https://doi.org/10.1038/nchembio.1500
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1500
This article is cited by
-
Stepwise assembly of the active site of [NiFe]-hydrogenase
Nature Chemical Biology (2023)
-
A personal account on 25 years of scientific literature on [FeFe]-hydrogenase
JBIC Journal of Biological Inorganic Chemistry (2023)
-
Aerobic hydrogen-oxidizing bacteria in soil: from cells to ecosystems
Reviews in Environmental Science and Bio/Technology (2022)
-
Double-flow focused liquid injector for efficient serial femtosecond crystallography
Scientific Reports (2017)
-
Metagenomic Sequencing Unravels Gene Fragments with Phylogenetic Signatures of O2-Tolerant NiFe Membrane-Bound Hydrogenases in Lacustrine Sediment
Current Microbiology (2015)