Abstract
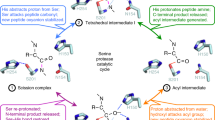
Intramembrane proteases hydrolyze peptide bonds within the membrane as a signaling paradigm universal to all life forms and with implications in disease. Deciphering the architectural strategies supporting intramembrane proteolysis is an essential but unattained goal. We integrated new, quantitative and high-throughput thermal light-scattering technology, reversible equilibrium unfolding and refolding and quantitative protease assays to interrogate rhomboid architecture with 151 purified variants. Rhomboid proteases maintain low intrinsic thermodynamic stability (ΔG = 2.1–4.5 kcal mol−1) resulting from a multitude of generally weak transmembrane packing interactions, making them highly responsive to their environment. Stability is consolidated by two buried glycines and several packing leucines, with a few multifaceted hydrogen bonds strategically deployed to two peripheral regions. Opposite these regions lie transmembrane segment 5 and connected loops that are notably exempt of structural responsibility, suggesting intramembrane proteolysis involves considerable but localized protein dynamics. Our analyses provide a comprehensive 'heat map' of the physiochemical anatomy underlying membrane-immersed enzyme function at, what is to our knowledge, unprecedented resolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Bowie, J.U. Solving the membrane protein folding problem. Nature 438, 581–589 (2005).
Booth, P.J. & Curnow, P. Folding scene investigation: membrane proteins. Curr. Opin. Struct. Biol. 19, 8–13 (2009).
De Strooper, B. & Annaert, W. Novel research horizons for presenilins and γ-secretases in cell biology and disease. Annu. Rev. Cell Dev. Biol. 26, 235–260 (2010).
Wolfe, M.S. Intramembrane proteolysis. Chem. Rev. 109, 1599–1612 (2009).
Brown, M.S., Ye, J., Rawson, R.B. & Goldstein, J.L. Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell 100, 391–398 (2000).
Urban, S. Making the cut: central roles of intramembrane proteolysis in pathogenic microorganisms. Nat. Rev. Microbiol. 7, 411–423 (2009).
Fluhrer, R., Steiner, H. & Haass, C. Intramembrane proteolysis by signal peptide peptidases: a comparative discussion of GXGD-type aspartyl proteases. J. Biol. Chem. 284, 13975–13979 (2009).
Greenblatt, E.J., Olzmann, J.A. & Kopito, R.R. Derlin-1 is a rhomboid pseudoprotease required for the dislocation of mutant α-1 antitrypsin from the endoplasmic reticulum. Nat. Struct. Mol. Biol. 18, 1147–1152 (2011).
Zettl, M., Adrain, C., Strisovsky, K., Lastun, V. & Freeman, M. Rhomboid family pseudoproteases use the ER quality control machinery to regulate intercellular signaling. Cell 145, 79–91 (2011).
Wang, Y., Zhang, Y. & Ha, Y. Crystal structure of a rhomboid family intramembrane protease. Nature 444, 179–180 (2006).
Wu, Z. et al. Structural analysis of a rhomboid family intramembrane protease reveals a gating mechanism for substrate entry. Nat. Struct. Mol. Biol. 13, 1084–1091 (2006).
Ben-Shem, A., Fass, D. & Bibi, E. Structural basis for intramembrane proteolysis by rhomboid serine proteases. Proc. Natl. Acad. Sci. USA 104, 462–466 (2007).
Baker, R.P., Young, K., Feng, L., Shi, Y. & Urban, S. Enzymatic analysis of a rhomboid intramembrane protease implicates transmembrane helix 5 as the lateral substrate gate. Proc. Natl. Acad. Sci. USA 104, 8257–8262 (2007).
Urban, S. Taking the plunge: integrating structural, enzymatic and computational insights into a unified model for membrane-immersed rhomboid proteolysis. Biochem. J. 425, 501–512 (2010).
Vinothkumar, K.R. Structure of rhomboid protease in a lipid environment. J. Mol. Biol. 407, 232–247 (2011).
Hong, H., Joh, N.H., Bowie, J.U. & Tamm, L.K. Methods for measuring the thermodynamic stability of membrane proteins. Methods Enzymol. 455, 213–236 (2009).
Curnow, P. & Booth, P.J. Combined kinetic and thermodynamic analysis of α-helical membrane protein unfolding. Proc. Natl. Acad. Sci. USA 104, 18970–18975 (2007).
Lau, F.W. & Bowie, J.U. A method for assessing the stability of a membrane protein. Biochemistry 36, 5884–5892 (1997).
Santoro, M.M. & Bolen, D.W. Unfolding free energy changes determined by the linear extrapolation method. 1. Unfolding of phenylmethanesulfonyl α-chymotrypsin using different denaturants. Biochemistry 27, 8063–8068 (1988).
Yeh, A.P., McMillan, A. & Stowell, M.H. Rapid and simple protein-stability screens: application to membrane proteins. Acta Crystallogr. D Biol. Crystallogr. 62, 451–457 (2006).
Senisterra, G.A. et al. Assessing the stability of membrane proteins to detect ligand binding using differential static light scattering. J. Biomol. Screen. 15, 314–320 (2010).
Choma, C., Gratkowski, H., Lear, J.D. & DeGrado, W.F. Asparagine-mediated self-association of a model transmembrane helix. Nat. Struct. Biol. 7, 161–166 (2000).
Zhou, F.X., Merianos, H.J., Brunger, A.T. & Engelman, D.M. Polar residues drive association of polyleucine transmembrane helices. Proc. Natl. Acad. Sci. USA 98, 2250–2255 (2001).
Joh, N.H. et al. Modest stabilization by most hydrogen-bonded side-chain interactions in membrane proteins. Nature 453, 1266–1270 (2008).
Curnow, P. & Booth, P.J. The contribution of a covalently bound cofactor to the folding and thermodynamic stability of an integral membrane protein. J. Mol. Biol. 403, 630–642 (2010).
Eilers, M., Shekar, S.C., Shieh, T., Smith, S.O. & Fleming, P.J. Internal packing of helical membrane proteins. Proc. Natl. Acad. Sci. USA 97, 5796–5801 (2000).
Oberai, A., Joh, N.H., Pettit, F.K. & Bowie, J.U. Structural imperatives impose diverse evolutionary constraints on helical membrane proteins. Proc. Natl. Acad. Sci. USA 106, 17747–17750 (2009).
Doura, A.K., Kobus, F.J., Dubrovsky, L., Hibbard, E. & Fleming, K.G. Sequence context modulates the stability of a GxxxG-mediated transmembrane helix-helix dimer. J. Mol. Biol. 341, 991–998 (2004).
Joh, N.H., Oberai, A., Yang, D., Whitelegge, J.P. & Bowie, J.U. Similar energetic contributions of packing in the core of membrane and water-soluble proteins. J. Am. Chem. Soc. 131, 10846–10847 (2009).
Russ, W.P. & Engelman, D.M. The GxxxG motif: a framework for transmembrane helix-helix association. J. Mol. Biol. 296, 911–919 (2000).
Schneider, D. & Engelman, D.M. Motifs of two small residues can assist but are not sufficient to mediate transmembrane helix interactions. J. Mol. Biol. 343, 799–804 (2004).
Urban, S. & Baker, R.P. In vivo analysis reveals substrate-gating mutants of a rhomboid intramembrane protease display increased activity in living cells. Biol. Chem. 389, 1107–1115 (2008).
Tastan, O., Klein-Seetharaman, J. & Meirovitch, H. The effect of loops on the structural organization of α-helical membrane proteins. Biophys. J. 96, 2299–2312 (2009).
Barrera, F.N. et al. Protein self-assembly and lipid binding in the folding of the potassium channel KcsA. Biochemistry 47, 2123–2133 (2008).
Veerappan, A., Cymer, F., Klein, N. & Schneider, D. The tetrameric α-helical membrane protein GlpF unfolds via a dimeric folding intermediate. Biochemistry 50, 10223–10230 (2011).
Urban, S. & Wolfe, M.S. Reconstitution of intramembrane proteolysis in vitro reveals that pure rhomboid is sufficient for catalysis and specificity. Proc. Natl. Acad. Sci. USA 102, 1883–1888 (2005).
Bondar, A.N., del Val, C. & White, S.H. Rhomboid protease dynamics and lipid interactions. Structure 17, 395–405 (2009).
Findlay, H.E., Rutherford, N.G., Henderson, P.J. & Booth, P.J. Unfolding free energy of a two-domain transmembrane sugar transport protein. Proc. Natl. Acad. Sci. USA 107, 18451–18456 (2010).
Otzen, D.E. Folding of DsbB in mixed micelles: a kinetic analysis of the stability of a bacterial membrane protein. J. Mol. Biol. 330, 641–649 (2003).
Fluhrer, R. et al. Intramembrane proteolysis of GXGD-type aspartyl proteases is slowed by a familial Alzheimer disease-like mutation. J. Biol. Chem. 283, 30121–30128 (2008).
Page, R.M. et al. Generation of Aβ38 and Aβ42 is independently and differentially affected by familial Alzheimer disease–associated presenilin mutations and γ-secretase modulation. J. Biol. Chem. 283, 677–683 (2008).
Osenkowski, P., Ye, W., Wang, R., Wolfe, M.S. & Selkoe, D.J. Direct and potent regulation of γ-secretase by its lipid microenvironment. J. Biol. Chem. 283, 22529–22540 (2008).
Santos, J.M., Ferguson, D.J., Blackman, M.J. & Soldati-Favre, D. Intramembrane cleavage of AMA1 triggers Toxoplasma to switch from an invasive to a replicative mode. Science 331, 473–477 (2011).
Acknowledgements
We are grateful to all members of the Urban lab and to D. Otzen for stimulating scientific discussions and to the Malaria Research Institute Biophysics Core for use of their CD spectropolarimeter. This work was supported by the Howard Hughes Medical Institute and the David and Lucile Packard Foundation.
Author information
Authors and Affiliations
Contributions
R.P.B. designed research, performed all biochemical experiments (except CD spectroscopy), analyzed data and prepared all figures; S.U. designed research, made all DNA constructs, performed CD spectroscopy, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 2060 kb)
Supplementary Data Set 1
Supplementary Dataset 1 (XLSX 53 kb)
Rights and permissions
About this article
Cite this article
Baker, R., Urban, S. Architectural and thermodynamic principles underlying intramembrane protease function. Nat Chem Biol 8, 759–768 (2012). https://doi.org/10.1038/nchembio.1021
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1021
This article is cited by
-
Ten catalytic snapshots of rhomboid intramembrane proteolysis from gate opening to peptide release
Nature Structural & Molecular Biology (2019)
-
Energy landscape underlying spontaneous insertion and folding of an alpha-helical transmembrane protein into a bilayer
Nature Communications (2018)
-
Structure formation during translocon-unassisted co-translational membrane protein folding
Scientific Reports (2017)
-
Steric trapping reveals a cooperativity network in the intramembrane protease GlpG
Nature Chemical Biology (2016)
-
Cytosolic extensions directly regulate a rhomboid protease by modulating substrate gating
Nature (2015)