Abstract
Anaphase chromatin bridges can lead to chromosome breakage if not properly resolved before completion of cytokinesis. The NoCut checkpoint, which depends on Aurora B at the spindle midzone, delays abscission in response to chromosome segregation defects in yeast and animal cells. How chromatin bridges are detected, and whether abscission inhibition prevents their damage, remain key unresolved questions. We find that bridges induced by DNA replication stress and by condensation or decatenation defects, but not dicentric chromosomes, delay abscission in a NoCut-dependent manner. Decatenation and condensation defects lead to spindle stabilization during cytokinesis, allowing bridge detection by Aurora B. NoCut does not prevent DNA damage following condensin or topoisomerase II inactivation; however, it protects anaphase bridges and promotes cellular viability after replication stress. Therefore, the molecular origin of chromatin bridges is critical for activation of NoCut, which plays a key role in the maintenance of genome stability after replicative stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Holland, A. J. & Cleveland, D. W. Boveri revisited: chromosomal instability, aneuploidy and tumorigenesis. Nat. Rev. Mol. Cell Biol. 10, 478–487 (2009).
Ganem, N. J. & Pellman, D. Linking abnormal mitosis to the acquisition of DNA damage. J. Cell Biol. 199, 871–881 (2012).
Mankouri, H. W., Huttner, D. & Hickson, I. D. How unfinished business from S-phase affects mitosis and beyond. EMBO J. 32, 2661–2671 (2013).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Naim, V., Wilhelm, T., Debatisse, M. & Rosselli, F. ERCC1 and MUS81–EME1 promote sister chromatid separation by processing late replication intermediates at common fragile sites during mitosis. Nat. Cell Biol. 15, 1–8 (2013).
Ying, S. et al. MUS81 promotes common fragile site expression. Nat. Cell Biol. 15, 1001–1007 (2013).
Chan, K.-L., North, P. S. & Hickson, I. D. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. EMBO J. 26, 3397–3409 (2007).
Wang, L. H.-C., Schwarzbraun, T., Speicher, M. R. & Nigg, E. A. Persistence of DNA threads in human anaphase cells suggests late completion of sister chromatid decatenation. Chromosoma 117, 123–135 (2007).
Wang, L. H. C., Mayer, B., Stemmann, O. & Nigg, E. A. Centromere DNA decatenation depends on cohesin removal and is required for mammalian cell division. J. Cell Sci. 123, 806–813 (2010).
Cimini, D. et al. Merotelic kinetochore orientation is a major mechanism of aneuploidy in mitotic mammalian tissue cells. J. Cell Biol. 153, 517–527 (2001).
Thompson, S. L. & Compton, D. A. Chromosome missegregation in human cells arises through specific types of kinetochore-microtubule attachment errors. Proc. Natl Acad. Sci. USA 108, 17974–17978 (2011).
Stewénius, Y. et al. Structural and numerical chromosome changes in colon cancer develop through telomere-mediated anaphase bridges, not through mitotic multipolarity. Proc. Natl Acad. Sci. USA 102, 5541–5546 (2005).
Norden, C. et al. The NoCut pathway links completion of cytokinesis to spindle midzone function to prevent chromosome breakage. Cell 125, 85–98 (2006).
Mendoza, M. et al. A mechanism for chromosome segregation sensing by the NoCut checkpoint. Nat. Cell Biol. 11, 477–483 (2009).
Germann, S. M. et al. TopBP1/Dpb11 binds DNA anaphase bridges to prevent genome instability. J. Cell Biol. 204, 45–59 (2013).
Mendoza, M. & Barral, Y. Co-ordination of cytokinesis with chromosome segregation. Biochem. Soc. Trans. 36, 387–390 (2008).
Mullins, J. M. & Biesele, J. J. Terminal phase of cytokinesis in D-98s cells. J. Cell Biol. 73, 672–684 (1977).
Steigemann, P. et al. Aurora B-mediated abscission checkpoint protects against tetraploidization. Cell 136, 473–484 (2009).
Carlton, J. G., Caballe, A., Agromayor, M., Kloc, M. & Martin-Serrano, J. ESCRT-III governs the Aurora B-mediated abscission checkpoint through CHMP4C. Science 336, 220–225 (2012).
Baxter, J. & Diffley, J. F. X. Topoisomerase II inactivation prevents the completion of DNA replication in budding yeast. Mol. Cell 30, 790–802 (2008).
Cuylen, S., Metz, J., Hruby, A. & Haering, C. H. Entrapment of chromosomes by condensin rings prevents their breakage during cytokinesis. Dev. Cell 27, 469–478 (2013).
Lopez, V. et al. Cytokinesis breaks dicentric chromosomes preferentially at pericentromeric regions and telomere fusions. Genes Dev. 29, 322–336 (2015).
Hoffelder, D. R. et al. Resolution of anaphase bridges in cancer cells. Chromosoma 112, 389–397 (2004).
Janssen, A., van der Burg, M., Szuhai, K., Kops, G. J. P. L. & Medema, R. H. Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333, 1895–1898 (2011).
Fasulo, B. et al. Chk1 and Wee1 kinases coordinate DNA replication, chromosome condensation, and anaphase entry. Mol. Biol. Cell 23, 1047–1057 (2012).
Chan, K.-L., Palmai-Pallag, T., Ying, S. & Hickson, I. D. Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nat. Cell Biol. 11, 753–760 (2009).
Onishi, M., Ko, N., Nishihama, R. & Pringle, J. R. Distinct roles of Rho1, Cdc42, and Cyk3 in septum formation and abscission during yeast cytokinesis. J. Cell Biol. 202, 311–329 (2013).
Lippincott, J. & Li, R. Nuclear envelope fission is linked to cytokinesis in budding yeast. Exp. Cell Res. 260, 277–283 (2000).
Dobbelaere, J. & Barral, Y. Spatial coordination of cytokinetic events by compartmentalization of the cell cortex. Science 305, 393–396 (2004).
Zhao, X., Muller, E. G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998).
Holm, C., Goto, T., Wang, J. C. & Botstein, D. DNA topoisomerase II is required at the time of mitosis in yeast. Cell 41, 553–563 (1985).
Lavoie, B. D., Hogan, E. & Koshland, D. In vivo dissection of the chromosome condensation machinery: reversibility of condensation distinguishes contributions of condensin and cohesin. J. Cell Biol. 156, 805–815 (2002).
Schmidt, M., Bowers, B., Varma, A., Roh, D.-H. & Cabib, E. In budding yeast, contraction of the actomyosin ring and formation of the primary septum at cytokinesis depend on each other. J. Cell Sci. 115, 293–302 (2002).
Tully, G. H., Nishihama, R., Pringle, J. R. & Morgan, D. O. The anaphase-promoting complex promotes actomyosin-ring disassembly during cytokinesis in yeast. Mol. Biol. Cell 20, 1201–1212 (2009).
Pereira, G. & Schiebel, E. Separase regulates INCENP-Aurora B anaphase spindle function through Cdc14. Science 302, 2120–2124 (2003).
Neurohr, G. et al. A midzone-based ruler adjusts chromosome compaction to anaphase spindle length. Science 332, 465–468 (2011).
Yang, S. S., Yeh, E., Salmon, E. D. & Bloom, K. Identification of a mid-anaphase checkpoint in budding yeast. J. Cell Biol. 136, 345–354 (1997).
Woodruff, J. B., Drubin, D. G. & Barnes, G. Mitotic spindle disassembly occurs via distinct subprocesses driven by the anaphase-promoting complex, Aurora B kinase, and kinesin-8. J. Cell Biol. 191, 795–808 (2010).
Juang, Y. L. et al. APC-mediated proteolysis of Ase1 and the morphogenesis of the mitotic spindle. Science 275, 1311–1314 (1997).
Floyd, S. et al. Spatiotemporal organization of Aurora B by APC/CCdh1 after mitosis coordinates cell spreading through FHOD1. J. Cell Sci. 126, 2845–2856 (2013).
Ramaswamy, V., Williams, J. S., Robinson, K. M., Sopko, R. L. & Schultz, M. C. Global control of histone modification by the anaphase-promoting complex. Mol. Cell. Biol. 23, 9136–9149 (2003).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Louvion, J. F., Havaux-Copf, B. & Picard, D. Fusion of GAL4-VP16 to a steroid-binding domain provides a tool for gratuitous induction of galactose-responsive genes in yeast. Gene 131, 129–134 (1993).
Idrissi, F.-Z. et al. Distinct acto/myosin-I structures associate with endocytic profiles at the plasma membrane. J. Cell Biol. 180, 1219–1232 (2008).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Acknowledgements
We thank S. Oliferenko and N. Brownlow for suggestions and critical reading of the manuscript; Y. Barral and D. Pellman for helpful discussions; G. Filion for help with statistical analysis; T. Sanmartin for technical support; the CRG Advanced Light Microscopy Unit; and Y. Schwab (EMBL, Heidelberg), C. López-Iglesias and Y. Muela-Castro (CCIT—University of Barcelona) for assistance with electron microscopy. This research was supported by ‘La Caixa’ fellowships to N.A., G.N. and M.Maier, and grants from the Spanish Ministry of Economy and Competitivity (BFU2011-30185 and CDS2009-00016 to M.-I.G.; BFU2015-71308 and BFU2013-50245-EXP to J.T.-R.; and BFU2009-08213 and BFU2012-37162 to M.Mendoza), and from the European Research Council (ERC Starting Grant 260965 to M.Mendoza). We acknowledge support from the Spanish Ministry of Economy and Competitiveness, ‘Centro de Excelencia Severo Ochoa 2013-2017’, SEV-2012-0208.
Author information
Authors and Affiliations
Contributions
A.V. made the initial observations on ycg1 mutants leading to the results shown in Figs 3a, b and 4a and Supplementary Fig. 3c. G.N. made the initial observations on conditional dicentric chromosomes leading to Supplementary Fig. 5a. EM tomography and serial sectioning was performed by C.F. (Fig. 3d and Supplementary Fig. 3d), and by F.-Z.I. and M.-I.G. (Supplementary Fig. 3e). A.K. and M.Maier contributed time-lapse imaging of dicentric chromosomes (Fig. 5c) and of HU-induced bridges (Supplementary Fig. 1c, d). N.C. and J.T.-R. performed PFGE experiments (Supplementary Fig. 5d). N.A. performed all other experiments. M.Mendoza designed the experiments with the person(s) performing them. N.A. and M.Mendoza wrote the paper. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Chromosome segregation and abscission dynamics during replication stress.
(a) Wild type cells expressing Htb2-mCherry were treated with 100 mM HU for 2 h, and transferred to fresh media. The Htb2-mCherry signal was used to estimate the frequency of anaphase (with elongated nuclei) or telophase (with separated nuclei) cells at the indicated times (left). Two independent experiments were performed with similar results; results from one experiment are shown. The following number of cells n was scored for each time point: n = 170 (0 min), 146 (30 min), 160 (60 min), 156 (90 min), 164 (120 min), 124 (150 min), 149 (180 min). The micrographs show cells 120 min after HU washout stained with DAPI to visualize DNA (right). The arrows point to chromatin bridges. (b) Kinetics of nuclear division (visualized with Nsg1) relative to actomyosin ring contraction (Myo1) in wild type cells in log-phase or after exposure to HU at 30 °C, as in Fig. 2a. Asterisk marks the time of nuclear division. Numbers indicate time in minutes. The graph shows the time of nuclear division relative to the onset of ring contraction (time 0). Boxes include 50% of data points, whiskers 90%; medians are shown as lines and means as crosses. The kinetics between the two conditions were considered to be not statistically significant (P > 0.05, Mann-Whitney test). n = 32 untreated cells and 30 HU-treated cells, pooled from two independent experiments. (c) The time of chromosome segregation (Htb2-mCherry) relative to actomyosin ring contraction (Myo1-GFP) in wild-type cells in the absence or presence of 60 mM HU. HU-treated cells were imaged for 2 h immediately after HU addition. The graph shows the time of chromosome segregation relative to the onset of ring contraction (time 0). Median (lines) and mean (crosses) are shown (n = 80 untreated cells and 66 HU-treated cells, pooled from 3 independent experiments). (d) Segregation of the rDNA (Net1-GFP) and TEL12R (TetR-YFP) in a representative wild-type cell in the presence of 60 mM HU, as in (c). Arrowheads mark the spindle poles, an asterisk marks the nucleolus, and arrows mark TEL12R. Time is shown in minutes relative to anaphase onset; nucleolar segregation occurs at 10 min and TEL12 segregation follows at 12 min. The graph shows the time of segregation of the rDNA and TEL12R in the absence and presence of HU. n = 50 untreated and 42 HU-treated cells, pooled from 2 independent experiments (P > 0.05, Student’s t-test). (e–h) Abscission dynamics of untreated and HU-treated wild type cells (d), sml1Δ (e) and ahc1Δ (f) mutant cells, imaged as in (c). The graphs in (e,f) show the fraction of cells completing abscission relative to the time of membrane ingression; the median abscission times are shown in (h). WT and WT + HU from (d) are represented as dashed lines for comparison in (f–g). n is the total number of cells in each category, pooled from two independent experiments, and is shown in brackets in (h). Asterisk marks significant differences between WT treated and untreated cells (P < 0.0001, Mann-Whitney test). In (b–d), boxes include 50% of data points, whiskers 90%; medians are shown as lines and means as crosses. Scale bars in (a,b,d): 2 μm.
Supplementary Figure 2 Cytokinesis-dependent DNA damage after DNA replication stress.
(a) Cells of the indicated strains were treated with 100 mM HU for 2 h and imaged for 20 min. The percentage of cells with Mre11-GFP foci is shown. n = 41 (wild type); 46 (ipl1-321); 35 (ipl1-321 cyk3Δ); 40 (ahc1Δ); 40 (ahc1Δcyk3Δ) where n is the total number of cells from one experiment each. (b) After HU washout, cells treated as in (a) were imaged during nuclear elongation. n = 98 (WT Untreated). For HU-treated cells: n = 190 (wild type); 176 (ipl1-321); 134 (ipl1-321 cdc15-1); 122 (ipl1-321 cyk3Δ); 128 (ahc1Δ); 116 (ahc1Δ cyk3Δ), where n is the total number of cells, pooled from two experiments, except for HU-treated WT and ipl1-321 cells, which were pooled from 3 experiments each. (c) Time-lapse images of representative wild type, ipl1-321 and ipl1-321 cyk3Δ cells expressing Mre11-GFP, after cytokinesis following a HU pulse. The arrows point to nuclear Mre11 foci. Numbers indicate time in minutes where time 0 is the frame before cytokinesis. Frames were selected to show Mre11-GFP foci in the ipl1-321 mutant. No foci were detected between 18 and 42 min. Scale bars, 2 μm. For complete image series of each cell, see Supplementary Videos 1 –3 .
Supplementary Figure 3 Chromosome segregation and cytokinesis in ycg1-2 and top2-4 cells.
(a) Anaphase progression in top2-4 cells with fluorescently tagged histone (Htb2) and myosin (Myo1). Numbers indicate time in minutes. Scale bar is 2 μm. The graphs show the time of actomyosin ring contraction relative to nuclear elongation (left) and the duration of contraction (right), in the indicated cell types. Line represents the mean. n = 50 (wild type), 32 (ycg1-2), 34 (top2-4), where n = number of cells pooled from two experiments. (b) Time-lapse imaging was used to determine the percentage of cells with Mre11-GFP foci after nuclear elongation in the indicated strains. n = 98 cells (wild type), 46 (ycg1-2), 40 (top2-4), where n = number of cells pooled from two experiments. (c) Time of appearance of the next bud after cytokinesis (rebudding) relative to the time of membrane closure (GFP-CAAX). Number of cells (pooled from two independent experiments): n = 35 (wild type), 38 (ycg1-2), 52 (top2-4). The rebudding kinetics of wild type and mutant cells were considered significantly different (P < 0.0001, Mann-Whitney test). (d) Slices from a tomogram of the septum in a top2-4 cell. Arrowheads point to nuclear membrane entering the channel (left); Arrows point to plasma membrane underlying the channel (right). Scale bars are 0.2 μm. (e) Serial ultra-thin sections (60-70 nm) from wild type and ycg1-2 cells 3 h after release from a G1 block. Septum channels were present in all examined ycg1-2 mutant cells (9), whereas all wild-type cells showed intact septa (13). Continuous septum deposition might constrict these channels and lead to their delayed closure. Arrows point to lacunae traversing the septum of the mutant. PS, primary septum, SS, secondary septum. Scale bar is 0.5 μm.
Supplementary Figure 4 The role of Ahc1 and Ipl1 in abscission inhibition in ycg1-2 and top2-4 mutants.
(a) Fraction of GFP-CAAX cells of the indicated strains completing abscission relative to the time of membrane ingression. Wild-type and ycg1-2 data from Fig. 3c are shown for comparison. n = 53 (ycg1-2 ahc1Δ); 50 (top2-4); 52 (top2-4 ahc1Δ), where n = total number of cells pooled from two independent experiments. P = 0.31 (ycg1-2 versus ycg1-2 ahc1Δ) and P < 0.05 (top2-4 versus top2-4 ahc1Δ) (Mann-Whitney test). (b) Time series of membrane ingression and resolution of the indicated strains. Scale bar is 2 μm. The graphs show the GFP fluorescence intensity across the cleavage plane in the middle optical section, as in Fig. 2b. (c) Quantification of the time of chromosome segregation in the indicated strains. Boxes include 50% of data points, whiskers 90%; medians are shown as lines and means as crosses. WT, ycg1-2 and top2-4 from Fig. 3a are shown for comparison. Time 0 was defined as the frame before the onset of contraction of the myosin ring. n = 32 cells (ycg1-2 ipl1-321); 38 (ycg1-2 slk19Δ); 26 (top2-4 ipl1-321). Data were pooled from two independent experiments. Asterisk marks significant differences (P < 0.01, Mann Whitney test).
Supplementary Figure 5 Characterization of the conditional dicentric LC(IV:XII).
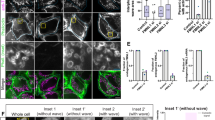
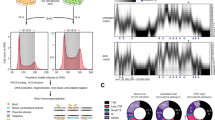
(a) Growth test of serial dilutions of the indicated cell types in rich medium plates with galactose (Gal) or glucose (Gluc). (b) Time of chromosome segregation for cells with the conditional dicentric chromosome in glucose, relative to full contraction of the myosin ring. Boxes include 50% of data points, whiskers 90%; medians are shown as lines and means as crosses. n = 20 cells (dicentric, no bridges); 19 (dicentric, with bridges), from one experiment. (c) Spindle tracks (left) and maximal spindle length (right) of cells with wild type chromosomes (WT) and the two categories of cells with dicentrics chromosomes (LC(IV:XII)). SPB distance is the linear distance between the two spindle pole bodies (SPBs) at each time point (Spc42-GFP). Both graphs show the mean and SD. Anaphase onset was defined when SPB distance was above 2.5 μm.n = 14 cells, pooled from two independent experiments, for each cell type or category. (d) PFGE analysis of dicentric chromosomes. Cells growing in galactose were arrested in G1 with alpha factor. The culture was split in two and glucose added to one half. Cells were released in glucose or galactose as indicated, and samples taken at times 0, 120 and 180 min for FACS, PFGE and Southern blot analysis. Asterisk marks position of broken dicentric molecules. The graph shows the amount of broken molecules in Southern blot, relative to intact dicentrics and arbitrarily set to 1 at time 0. (e) Abscission differences in the presence of distinct types of chromatin bridges are not due to differences in the amount of chromatin present at the cytokinesis site. The intensity of fluorescently labeled histones at the bud neck was low in condensin-deficient bridges (which inhibit abscission), whereas it was approximately 10-fold higher in both abscission-competent dicentric cells and in abscission-defective top2-4 mutants. The graph shows fluorescence intensities of Htb2-mCherry in the bud neck region at the onset of myosin ring contraction (Myo1-GFP), for the indicated cell types and conditions. Values are in arbitrary units (A. U.) and background-subtracted. Lines represent the mean. n = 15 cells (WT); 15 (Dicentrics, with bridges); 15 (ycg1-2); 15 (top2-4); where n = number of cells per strain, pooled from two independent experiments.
Supplementary Figure 6 The role of DNA damage and replication checkpoint in abscission dynamics in HU-treated, ycg1-2 and top2-4 cells.
(a,b) Mutation of MRE11, TEL1, RAD9, MRC1, CHK1 and RAD53 lead to abscission delays even in the absence of HU, perhaps due to endogenous replication stress. However, no significant differences in abscission are found between wild type and checkpoint-deficient HU-treated cells. Abscission dynamics of untreated and HU-treated cells of the indicated strains at 30 °C are shown. Cells were treated as in Supplementary Fig. 1c, d. Open inverted triangles represent untreated cells and filled inverted triangles represent HU-treated cells. WT and WT + HU from Supplementary Fig. 1e are represented as blue and green dashed lines for comparison. The median abscission times are represented for the indicated strains and conditions. n = total number of cells pooled from two independent experiments is represented in brackets. Asterisk represents significant differences relative to WT –HU (P < 0.05, Mann-Whitney test). ns, non significant. (c,d) Inactivation of the upstream checkpoint kinases Mec1 and Tel1 (homologues of mammalian ATM and ATR) does not restore abscission in top2-4 or ycg1-2 cells. Abscission dynamics in the indicated strains at 37 °C. GFP-CAAX cells were imaged and analyzed as in Fig. 3b, c. SML1 was deleted to allow viability of the mec1 mutant. n = total number of cells pooled from two independent experiments (except for ycg1-2 rad53-21, which is from 3 experiments) is represented in brackets. WT, ycg1-2 and top2-4 data from Fig. 3c are represented for comparison. Significant differences were found only for ycg1-2 rad53-21 and ycg1-2 mre11Δ relative to ycg1-2 (P < 0.0001, Mann-Whitney test).
Supplementary Figure 7 Spindle dynamics in HU-treated, ycg1-2 and top2-4 cells.
(a) Anaphase spindle disassembly visualized through tubulin (Tub1) and myosin (Myo1) dynamics in a top2-4 cell at 37 °C (top) and in a wild type cell after HU exposure at 30 °C (bottom). Asterisk marks full contraction of the myosin ring. Arrow points to depolymerizing spindle. Numbers indicate time in minutes. Scale bars are 2 μm. (b) Spindle tracks of wild type and ycg1-2 cells. SPB distance is the linear distance between the two spindle pole bodies (SPBs) at each time point. Mean and SD are shown. Anaphase onset was defined when SPB distance was above 2.5 μm. Arrows mark the time of spindle disassembly. n = 7 cells (wild type); 6 (ycg1-2). (c,d) Visualization of Cin8-GFP and Iqg-GFP dynamics during spindle elongation (Spc42-mCherry) in wild type and ycg1-2 cells; time interval is 1.5 min. All scale bars, 2 μm.
Supplementary Figure 8 Role of Cdh1 and Ahc1 in spindle stability and dicentric bridge detection.
(a) Anaphase spindle disassembly visualized through tubulin (Tub1-GFP) and myosin (Myo1-GFP) dynamics in a representative cdh1Δ cell. An asterisk marks full contraction of the myosin ring. 85% (26/30) of cdh1Δ cells disassembled the spindle after ring contraction. Scale bar, 2 μm. (b) Cumulative percentage of cells with resolved membranes (abscission) for cells of the indicated strains and categories. Wild-type and cdh1Δ dicentric cells (from Fig. 5d and Fig. 7a, b, respectively) are shown for comparison. cdh1Δ ahc1Δ, no bridges n = 34 cells; cdh1Δahc1Δ, with bridges n = 40 (pooled from two independent experiments). Abscission kinetics of cdh1Δ, with bridges and cdh1Δ ahc1Δ, with bridges were significantly different (P < 0.0005, Mann-Whitney test).
Supplementary information
Supplementary Information
Supplementary Information (PDF 1517 kb)
Cytokinesis and DNA damage after replication stress.
Time-lapse of a representative wild type cell expressing Mre11-GFP to visualize DNA damage, after cytokinesis following a HU pulse. DIC and GFP channels are shown. Numbers indicate time in minutes, time 0 is the frame before cytokinesis. This video is associated with Supplementary Fig. 2c. (AVI 132 kb)
Cytokinesis and DNA damage after replication stress.
Time-lapse of a representative ipl1-321 cell expressing Mre11-GFP to visualize DNA damage, after cytokinesis following a HU pulse. DIC and GFP channels are shown. Numbers indicate time in minutes, time 0 is the frame before cytokinesis. This video is associated with Supplementary Fig. 2c. (AVI 154 kb)
Cytokinesis and DNA damage after replication stress.
Time-lapse of a representative ipl1-321 cyk3Δ cell expressing Mre11-GFP to visualize DNA damage, after cytokinesis following a HU pulse. DIC and GFP channels are shown. Numbers indicate time in minutes, time 0 is the frame before cytokinesis. This video is associated with Supplementary Fig. 2c. (AVI 133 kb)
Inactivation of condensin inhibits abscission.
EM tomogram and 3D model of the membrane organization at the medial septum of a ycg1-2 cell. Plasma membrane is in purple, nuclear envelope in light gray, microtubules in green and vesicles in yellow. This video is associated with Fig. 3d. (AVI 33009 kb)
Inactivation of topoisomerase II inhibits abscission.
EM tomogram showing the membrane organization at the medial septum of a top2-4 cell. This video is associated with Supplementary Fig. 3d. (AVI 37761 kb)
Rights and permissions
About this article
Cite this article
Amaral, N., Vendrell, A., Funaya, C. et al. The Aurora-B-dependent NoCut checkpoint prevents damage of anaphase bridges after DNA replication stress. Nat Cell Biol 18, 516–526 (2016). https://doi.org/10.1038/ncb3343
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3343
This article is cited by
-
Topoisomerase II deficiency leads to a postreplicative structural shift in all Saccharomyces cerevisiae chromosomes
Scientific Reports (2021)
-
Budding yeast complete DNA synthesis after chromosome segregation begins
Nature Communications (2020)
-
Building bridges between chromosomes: novel insights into the abscission checkpoint
Cellular and Molecular Life Sciences (2019)
-
USP35 regulates mitotic progression by modulating the stability of Aurora B
Nature Communications (2018)
-
Myosin efflux promotes cell elongation to coordinate chromosome segregation with cell cleavage
Nature Communications (2017)