Abstract
Proper microtubule nucleation during cell division requires augmin, a microtubule-associated hetero-octameric protein complex. In current models, augmin recruits γ-tubulin, through the carboxyl terminus of its hDgt6 subunit to nucleate microtubules within spindles. However, augmin’s biochemical complexity has restricted analysis of its structural organization and function. Here, we reconstitute human augmin and show that it is a Y-shaped complex that can adopt multiple conformations. Further, we find that a dimeric sub-complex retains in vitro microtubule-binding properties of octameric complexes, but not proper metaphase spindle localization. Addition of octameric augmin complexes to Xenopus egg extracts promotes microtubule aster formation, an activity enhanced by Ran–GTP. This activity requires microtubule binding, but not the characterized hDgt6 γ-tubulin-recruitment domain. Tetrameric sub-complexes induce asters, but activity and microtubule bundling within asters are reduced compared with octameric complexes. Together, our findings shed light on augmin’s structural organization and microtubule-binding properties, and define subunits required for its function in organizing microtubule-based structures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Walczak, C. E. & Heald, R. Mechanisms of mitotic spindle assembly and function. Int. Rev. Cytol. 265, 111–158 (2008).
Meunier, S. & Vernos, I. Microtubule assembly during mitosis---from distinct origins to distinct functions? J. Cell Sci. 15, 2805–2814 (2012).
Teixidó-Travesa, N., Roig, J. & Lüders, J. The where, when and how of microtubule nucleation--one ring to rule them all. J. Cell Sci. 125, 4445–4456 (2012).
Lüders, J. & Stearns, T. Microtubule-organizing centres: a re-evaluation. Nat. Rev. Mol. Cell Biol. 8, 161–167 (2007).
Bettencourt-Dias, M. & Glover, D. M. Centrosome biogenesis and function: centrosomics brings new understanding. Nat. Rev. Mol. Cell Biol. 8, 451–463 (2007).
Kollman, J. M., Merdes, A., Mourey, L. & Agard, D. A. Microtubule nucleation by γ-tubulin complexes. Nat. Rev. Mol. Cell Biol. 12, 709–721 (2011).
Wadsworth, P. & Khodjakov, A. E pluribus unum: towards a universal mechanism for spindle assembly. Trends Cell Biol. 14, 413–419 (2004).
Walczak, C. E., Cai, S. & Khodjakov, A. Mechanisms of chromosome behaviour during mitosis. Nat. Rev. Mol. Cell Biol. 11, 91–102 (2010).
Dumont, J. & Desai, A. Acentrosomal spindle assembly and chromosome segregation during oocyte meiosis. Trends Cell Biol. 22, 241–249 (2012).
Karsenti, E. & Vernos, I. The mitotic spindle: a self-made machine. Science 294, 543–547 (2001).
Kelly, A. E. & Funabiki, H. Correcting aberrant kinetochore microtubule attachments: an Aurora B-centric view. Curr. Opin. Cell Biol. 21, 51–58 (2009).
Goshima, G., Mayer, M., Zhang, N., Stuurman, N. & Vale, R. D. Augmin: a protein complex required for centrosome-independent microtubule generation within the spindle. J. Cell Biol. 181, 421–429 (2008).
Uehara, R. et al. The augmin complex plays a critical role in spindle microtubule generation for mitotic progression and cytokinesis in human cells. Proc. Natl Acad. Sci. USA 106, 6998–7003 (2009).
Lawo, S. et al. HAUS, the 8-subunit human Augmin complex, regulates centrosome and spindle integrity. Curr. Biol. 19, 816–826 (2009).
Colombié, N., Głuszek, A. A., Meireles, A. M. & Ohkura, H. Meiosis-specific stable binding of augmin to acentrosomal spindle poles promotes biased microtubule assembly in oocytes. PLoS Genet. 9, e1003562 (2013).
Petry, S., Pugieux, C., Nédélec, F. J. & Vale, R. D. Augmin promotes meiotic spindle formation and bipolarity in Xenopus egg extracts. Proc. Natl Acad. Sci. USA 108, 14473–14478 (2011).
Petry, S., Groen, A. C., Ishihara, K., Mitchison, T. J. & Vale, R. D. Branching microtubule nucleation in Xenopus egg extracts mediated by augmin and TPX2. Cell 152, 768–777 (2013).
Kamasaki, T. et al. Augmin-dependent microtubule nucleation at microtubule walls in the spindle. J. Cell Biol. 202, 25–33 (2013).
Hayward, D., Metz, J., Pellacani, C. & Wakefield, J. G. Synergy between multiple microtubule-generating pathways confers robustness to centrosome-driven mitotic spindle formation. Dev. Cell 28, 81–93 (2014).
Wu, G. et al. Hice1, a novel microtubule-associated protein required for maintenance of spindle integrity and chromosomal stability in human cells. Mol. Cell. Biol. 28, 3652–3662 (2008).
Tan, S., Kern, R. C. & Selleck, W. The pST44 polycistronic expression system for producing protein complexes in Escherichia coli. Protein Expr. Purif. 40, 385–95 (2005).
Subramanian, R. et al. Insights into antiparallel microtubule crosslinking by PRC1, a conserved nonmotor microtubule binding protein. Cell 142, 433–443 (2010).
Trowitzsch, S., Bieniossek, C., Nie, Y., Garzoni, F. & Berger, I. New baculovirus expression tools for recombinant protein complex production. J. Struct. Biol. 172, 45–54 (2010).
Desai, A., Murray, A., Mitchison, T. J. & Walczak, C. E. The use of Xenopus egg extracts to study mitotic spindle assembly and function in vitro. Methods Cell Biol. 61, 385–412 (1999).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Hohn, M. et al. SPARX, a new environment for cryo-EM image processing. J. Struct. Biol. 157, 47–55 (2007).
Yang, Z., Fang, J., Chittuluru, J., Asturias, F. J. & Penczek, P. A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 20, 237–247 (2012).
Smith, M. B. et al. Interactive, computer-assisted tracking of speckle trajectories in fluorescence microscopy: application to actin polymerization and membrane fusion. Biophys. J. 101, 1794–1804 (2011).
Folta-Stogniew, E. & Williams, K. R. Determination of molecular masses of proteins in solution: implementation of an HPLC size exclusion chromatography and laser light scattering service in a core laboratory. J. Biomol. Tech. 10, 51–63 (1999).
Acknowledgements
We thank G. Goshima (Nagoya University, Japan) for the gift of UCHL5IP antibody; and L. Pelletier (University of Toronto, Canada) for the antibodies against C14orf94 and Cep27. The SEC-LS/UV/RI instrumentation used for light-scattering analysis was supported by a National Institutes of Health (NIH) award (1S10RR023748-01). TEM studies were conducted at the National Resource for Automated Molecular Microscopy, which is supported by the National Institute of General Medical Sciences (9 P41 GM103310). K-C.H. was supported by the Kimberly Lawrence-Netter Cancer Research Discovery Fund at The Rockefeller University and is supported by a Special Fellow Award from The Leukemia and Lymphoma Society. S.F. acknowledges postdoctoral support from an NIH National Research Service Award Fellowship (F32GM099380). R.A.M. acknowledges support from the NIH (GM-052468). T.M.K. acknowledges support from the NIH (GM-65933).
Author information
Authors and Affiliations
Contributions
K-C.H. carried out experiments in Figs 1–4 and assembled all figures. Q.H. helped purify and characterize proteins. Acquisition and interpretation of the electron microscopy and image analysis data shown in Fig. 5 were carried out by E.M.W-K., K-L.T. and R.A.M. Single-molecule assays in Fig. 3 were carried out by A.D. K-C.H. and Y.S. carried out experiments involving Xenopus egg extracts (Figs 6–8). S.F. quantitatively analysed aster morphology (Fig. 8). All authors helped write the manuscript. T.M.K. directed the project and helped design experiments and prepare the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1
Biochemical characterization of augmin sub-complexes. (a–c) Oligomeric states of Hice1⋅hDgt6Δ(433–955) (a), tetramer-I (Hice1⋅hDgt6 (1–432)⋅UCHL5IP⋅Cep27) (b), and tetramer-II (Hice1⋅hDgt6 (1–432)⋅His-C14orf94⋅Ccdc5) (c) were analysed by size exclusion chromatography coupled with light scattering. Recombinant sub-complexes elute earlier than expected for a globular protein of equivalent molecular mass. Elution volumes of molecular weight standards used in the size exclusion chromatography are indicated. The average molecular mass for sub-complexes analysed by light scattering are indicated (lines across the respective elution peaks). (d) Observed results from light scattering experiments (Obs) are presented beside molecular weights calculated using standards (Cal). (e) Purified hexamer (Hice1⋅hDgt6 (1–432)⋅UCHL5IP⋅Cep27⋅His-C14orf94⋅Ccdc5) was analysed by SDS-PAGE, followed by staining with Coomassie blue. (f) Western blot analysis of the hexameric complex with the indicated antibodies. (g) C14orf94 (a.a. 1–188) ⋅full length Ccdc5 hetero-dimer was examined by size exclusion chromatography (Superdex 200 16/60). Peak fractions (between 60 and 80 ml; volumes indicted) were analysed by SDS-PAGE (staining with Coomassie blue). Void volume (Vo) is indicated. The star indicates a GST-containing peak. Absorbance (a.u.) is 280 nm.
Supplementary Figure 2
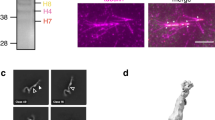
Analysis of subunits in the augmin holo-complex. (a) Recombinant holo-complex (Hice1⋅GFP-hDgt6⋅UCHL5IP⋅GFP-Cep27⋅C14orf94⋅Ccdc5⋅His-hDgt3⋅hDgt5) was analysed by SDS-PAGE, followed by staining with Coomassie blue. (b-d) Western blots were carried out using the indicated antibodies. GFP-tagged Cep27 within purified tetramer-I was also detected using anti-GFP antibody (b). The star indicates a GFP-containing degradation product of GFP-Cep27.
Supplementary Figure 3
Single molecule analysis of augmin complexes. (a–f) Fluorescence intensity average and distribution analysis of single particles for each complex adhered to a glass surface: (a) Hice1⋅hDgt6Δ(433–955) (intensity average 5,400 ± 1,800, N = 593 particles), (b) tetramer-I (GFP-Cep27) (intensity average 4,800 ± 1,800, N = 600 particles), (c) tetramer-I (GFP-hDgt6 (1–432)) (intensity average 5,400 ± 2,500, N = 631 particles), (d) octamer[hDgt6D(433–955)] (intensity average 4,500 ± 2,000, N = 626 particles), (e) holo-complex (8,300 ± 7,000, N = 662 particles). (f) Monomeric GFP (intensity average 4,800 ± 1,700, N = 577 particles) was used as reference. Intensity averages are reported as mean ± s.d. Three or more independent experiments were analysed. (g–k) The mean square displacements for Hice1⋅hDgt6Δ(433–955) (g), tetramer-I (GFP-Cep27) (h), tetramer-I (GFP-hDgt6 (1–432)) (i), octamer [hDgt6Δ(433–955)] (j), holo-complex (k) are shown. Diffusion coefficients (D) are calculated as half the slope of the fit line. Three or more independent experiments were analysed. (l) Image of GMPCPP-stabilized microtubules (X-rhodamine- and biotin-labelled) (top), GFP-tagged complexes (middle, maximum intensity projections from 300 images in the time-lapse sequence) and corresponding kymographs (below) are shown for GFP-tagged tetramer-I (GFP-hDgt6 (1–432)) that also has a GFP on hDgt6, as is the case for the dimer. Scale bars, horizontal, 2 μm; vertical, 2 s. (m) The region highlighted (box) in each kymograph is also shown in greater detail as a montage. Scale bar, 2 μm. (n) Binding of individual molecules to a microtubule was tracked to compute the CDF of the dwell time for the complex. Mean dwell time 〈t〉 and relative amplitude (in parentheses) were obtained by fitting to bi-exponential functions (grey curve). Data from three or more independent experiments were analysed.
Supplementary Figure 4
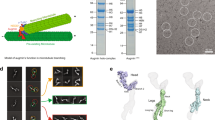
Electron microscopy analyses of augmin sub-complexes. (a,b) Negative stain electron micrographs of taxol stabilized microtubules in the presence of (a) Hice1⋅hDgt6Δ(433–955) and (b) GFP-tagged tetramer-I. Scale bar, 40 nm. Microtubule without tetramer-I binding is indicated by arrow. (c) SDS-PAGE analysis (stained with Coomassie blue) of purified tetramer-I with MBP-tagged Hice1 C-terminus. (d) Negative stain electron micrograph of the MBP-tagged tetramer-I (Hice1-MBP⋅hDgt6 (1–432)⋅UCHL5IP⋅Cep27) particles. Three representative class averages of individual particles (N = 82, 52 and 49) are shown. The position of the MBP-tag is indicated (arrow). Scale bar, 10 nm (e,f) Negative stain electron micrograph of tetramer-I (Hice1⋅hDgt6 (1–432)⋅UCHL5IP⋅Cep27) (e) and tetramer-II (Hice1⋅hDgt6 (1–432)⋅C14orf94⋅Ccdc5) (f) particles. A field of either tetramer-I or -II particles adsorbed onto a glow-discharged carbon grid was stained with 2% (w/v) uranyl acetate and processed for imaging. Scale bar, 100 nm.
Supplementary Figure 5
Analysis of metaphase spindle localization of augmin complexes. (a) Western blot analysis of crude Xenopus egg extracts (10-fold dilution in CSF-XB buffer, 7 ml) with anti-xCcdc5 antibody (lane 6). Equal volumes (7 ml) of serial dilutions of recombinant xCcdc5 (a.a. 1–204) were analysed by SDS-PAGE to determine endogenous xCccdc5 concentration (lane 1–5). Concentrations of recombinant xCcdc5 protein are indicated. (b) Localization of endogenous augmin in the metaphase spindles assembled in Xenopus egg extracts, fixed, spun down on coverslips and stained with anti-xCcdc5 and anti-α-tubulin antibodies. xCcdc5 antibody staining, left panels; and α-tubulin antibody staining (middle panels); and overlays (right panels; microtubules, red; xCcdc5, green; and DNA, blue), are shown. Scale bar, 10 μm. (c) Additional fluorescence images of metaphase spindles in the presence GFP-tagged octamer[hDgt6Δ(433–955). GFP (left panels); X-rhodamine (middle panels); and overlays (right panels; microtubules, red; GFP, green; and DNA, blue), are shown. Scale bar, 10 μm. (d,f) Fluorescence images of metaphase spindles in the presence of GFP-tagged Hice1⋅hDgt6Δ(433–955) (d) and GFP protein (f) (left panels, 15 nM); X-rhodamine (middle panels); and overlays (right panels; microtubules, red; GFP, green; and DNA, blue), are shown for spindles in d,f. Scale bar, 10 μm. (e,g) Linescans for GFP fluorescence (green) and X-rhodamine (red) along the long axis of the spindle are shown. (h) The intensity of the GFP image in the presence of GFP-tagged octamer[hDgt6Δ(433–955), Hice1Δ(1–140)] was adjusted to visualize GFP signal at the spindle poles. (i) Localization analysis of augmin holo-complex in metaphase spindles using Xenopus egg extracts. Recombinant GFP-tagged holo-complex (6 nM) (left panels); tubulin (X-rhodamine-labelled, middle panels); and overlays (right panels; tubulin, red; GFP, green; and DNA, blue), are shown. Scale bar, 10 μm.
Supplementary Figure 6
Microtubule aster formation in the presence of augmin complexes. (a,b) Aster assembly in the presence of Ran(Q69L) and GFP-tagged octamer[hDgt6Δ(433–955)] (a) or holo-complex (b) at 6 nM. (c) Immunodepletion of xCcdc5 from Xenopusegg extracts using anti-xCcdc5 antibody. xCcdc5 bands in crude (A), IgG-depleted (B) and xCcdc5-depleted (C) egg extracts were analysed by western blots. Equal volumes of egg extract samples with the same dilution were analysed by SDS-PAGE. Supernatant fractions (S) and bead bound fractions (P) are indicated. IgG and xCcdc5 bands are indicated. Three independent experiments were performed and depletion was found to be ∼60%. (d) A representative image of microtubule aster formed in the presence of Ran(Q69L) and GFP-tagged tetramer-II at 15 nM. Tubulin (X-rhodamine-labelled), left panel; and GFP fluorescence, right panel, are shown. Scale bar, 10 μm. (e) Analysis of microtubule (rhodamine signal) and augmin (GFP signal) levels in asters. Ratios of average fluorescence at 1 μm versus 5 mm radius for tetramer-II (15 nM) induced asters are shown. (N = 19 asters; s.d. was determined from data pooled from 3 independent experiments.) (f–h) Representative images after polar transformation of asters induced by RanQ69L alone (f) and in the presence of Ran(Q69L) and holo-complex (15 nM) (g) and octamer[hDgt6Δ(433–955)] (15 nM) (h). (i–m) Representative images of microtubule asters induced by Ran(Q69L) alone (i) and in the presence of Ran(Q69L) and GFP-tagged holo-complex (15 nM) (j), octamer[hDgt6Δ(433–955)] (15 nM) (k), octamer[hDgt6Δ(433–955)] (60 nM) (l) or tetramer-II (60 nM) (m). Three half-circle (180 degree) regions (at 4 and 8, and 12 μm radii) used for determination of coefficient of variation are indicated in first image for each panel (dashed white lines).
Supplementary Figure 7
Analysis of aster number and microtubule intensity in asters promoted by addition of recombinant augmin complexes to egg extracts. Individual fluorescence images of X-rhodamine-labelled microtubules were taken automatically using a motorized XY stage and then 400 captured images were stitched into one composite image using NIS-Element software. A representative single image from each composite image is shown in the bottom inset. Augmin complexes and Ran(Q69L) were added to extracts, incubated for 10 min (or as noted), fixed and processed. (a) Representative composite images of microtubule asters in the presence of buffer control, GFP-tagged octamer[hDgt6Δ(433–955)] and holo-complex. (b) Representative composite images of microtubule asters in the presence of GFP-tagged octamer[hDgt6Δ(433–955), Hice1Δ(1–140)] and octamer[hDgt6Δ(433–955)] with Ran(Q69L). (c) Representative composite images of microtubule asters in the presence of indicated GFP-tagged octamer[hDgt6Δ(433–955)] concentrations. (d) Representative composite images of microtubule asters in the presence of GFP-tagged octamer[hDgt6Δ(433–955)] without Ran(Q69L). Scale bars, 1 mm (composite image); 100 mm (single image). (e) Total microtubule intensity in asters induced in the presence of Ran(Q69L) and GFP-tagged tetramer-II or octamer[hDgt6Δ(433–955)] at 60 nM. (f) Total microtubule intensity in asters induced by of Ran(Q69L) alone and in the presence of Ran(Q69L) and GFP-tagged holo-complex or octamer[hDgt6Δ(433–955)] at 15 nM. s.d. was determined from data pooled from at least 3 independent experiments.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1245 kb)
Rights and permissions
About this article
Cite this article
Hsia, KC., Wilson-Kubalek, E., Dottore, A. et al. Reconstitution of the augmin complex provides insights into its architecture and function. Nat Cell Biol 16, 852–863 (2014). https://doi.org/10.1038/ncb3030
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3030
This article is cited by
-
Mechanisms underlying spindle assembly and robustness
Nature Reviews Molecular Cell Biology (2023)
-
Structural insights into how augmin augments the mitotic spindle
Nature Communications (2023)
-
Comprehensive analysis of the correlation of the pan-cancer gene HAUS5 with prognosis and immune infiltration in liver cancer
Scientific Reports (2023)
-
Sub-centrosomal mapping identifies augmin-γTuRC as part of a centriole-stabilizing scaffold
Nature Communications (2021)
-
Microtubule nucleation: beyond the template
Nature Reviews Molecular Cell Biology (2017)