Abstract
Macrophages are required for normal embryogenesis, tissue homeostasis and immunity against microorganisms and tumours1,2,3,4. Adult tissue-resident macrophages largely originate from long-lived, self-renewing embryonic precursors and not from haematopoietic stem-cell activity in the bone marrow4,5. Although fate-mapping studies have uncovered a great amount of detail on the origin and kinetics of fetal macrophage development in the yolk sac and liver6,7,8,9,10,11, the molecules that govern the tissue-specific migration of these cells remain completely unknown. Here we show that an endothelium-specific molecule, plasmalemma vesicle-associated protein (PLVAP), regulates the seeding of fetal monocyte-derived macrophages to tissues in mice. We found that PLVAP-deficient mice have completely normal levels of both yolk-sac- and bone-marrow-derived macrophages, but that fetal liver monocyte-derived macrophage populations were practically missing from tissues. Adult PLVAP-deficient mice show major alterations in macrophage-dependent iron recycling and mammary branching morphogenesis. PLVAP forms diaphragms in the fenestrae of liver sinusoidal endothelium during embryogenesis, interacts with chemoattractants and adhesion molecules and regulates the egress of fetal liver monocytes to the systemic vasculature. Thus, PLVAP selectively controls the exit of macrophage precursors from the fetal liver and, to our knowledge, is the first molecule identified in any organ as regulating the migratory events during embryonic macrophage ontogeny.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wynn, T. A., Chawla, A. & Pollard, J. W. Macrophage biology in development, homeostasis and disease. Nature 496, 445–455 (2013)
Davies, L. C., Jenkins, S. J., Allen, J. E. & Taylor, P. R. Tissue-resident macrophages. Nat. Immunol. 14, 986–995 (2013)
Ginhoux, F., Schultze, J. L., Murray, P. J., Ochando, J. & Biswas, S. K. New insights into the multidimensional concept of macrophage ontogeny, activation and function. Nat. Immunol. 17, 34–40 (2016)
Varol, C., Mildner, A. & Jung, S. Macrophages: development and tissue specialization. Annu. Rev. Immunol. 33, 643–675 (2015)
Sieweke, M. H. & Allen, J. E. Beyond stem cells: self-renewal of differentiated macrophages. Science 342, 1242974 (2013)
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010)
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012)
Gomez Perdiguero, E. et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature 518, 547–551 (2015)
Sheng, J., Ruedl, C. & Karjalainen, K. Most tissue-resident macrophages except microglia are derived from fetal hematopoietic stem cells. Immunity 43, 382–393 (2015)
Hoeffel, G. et al. C-Myb+ erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015)
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013)
Okabe, Y. & Medzhitov, R. Tissue biology perspective on macrophages. Nat. Immunol. 17, 9–17 (2016)
Amit, I., Winter, D. R. & Jung, S. The role of the local environment and epigenetics in shaping macrophage identity and their effect on tissue homeostasis. Nat. Immunol. 17, 18–25 (2016)
Palis, J., Robertson, S., Kennedy, M., Wall, C. & Keller, G. Development of erythroid and myeloid progenitors in the yolk sac and embryo proper of the mouse. Development 126, 5073–5084 (1999)
McGrath, K. E., Koniski, A. D., Malik, J. & Palis, J. Circulation is established in a stepwise pattern in the mammalian embryo. Blood 101, 1669–1676 (2003)
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013)
McGrath, K. E. et al. Distinct sources of hematopoietic progenitors emerge before HSCs and provide functional blood cells in the mammalian embryo. Cell Reports 11, 1892–1904 (2015)
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013)
Epelman, S. et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 40, 91–104 (2014)
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013)
Hoeffel, G. et al. Adult Langerhans cells derive predominantly from embryonic fetal liver monocytes with a minor contribution of yolk sac-derived macrophages. J. Exp. Med. 209, 1167–1181 (2012)
Kumaravelu, P. et al. Quantitative developmental anatomy of definitive haematopoietic stem cells/long-term repopulating units (HSC/RUs): role of the aorta-gonad-mesonephros (AGM) region and the yolk sac in colonisation of the mouse embryonic liver. Development 129, 4891–4899 (2002)
Bain, C. C. et al. Constant replenishment from circulating monocytes maintains the macrophage pool in the intestine of adult mice. Nat. Immunol. 15, 929–937 (2014)
Rantakari, P. et al. The endothelial protein PLVAP in lymphatics controls the entry of lymphocytes and antigens into lymph nodes. Nat. Immunol. 16, 386–396 (2015)
Haldar, M. et al. Heme-mediated SPI-C induction promotes monocyte differentiation into iron-recycling macrophages. Cell 156, 1223–1234 (2014)
Carlin, L. M. et al. Nr4a1-dependent Ly6Clow monocytes monitor endothelial cells and orchestrate their disposal. Cell 153, 362–375 (2013)
Stan, R. V. Endothelial stomatal and fenestral diaphragms in normal vessels and angiogenesis. J. Cell. Mol. Med. 11, 621–643 (2007)
Stan, R. V. et al. The diaphragms of fenestrated endothelia: gatekeepers of vascular permeability and blood composition. Dev. Cell 23, 1203–1218 (2012)
Tchaikovski, V., Fellbrich, G. & Waltenberger, J. The molecular basis of VEGFR-1 signal transduction pathways in primary human monocytes. Arterioscler. Thromb. Vasc. Biol. 28, 322–328 (2008)
Dejda, A. et al. Neuropilin-1 mediates myeloid cell chemoattraction and influences retinal neuroimmune crosstalk. J. Clin. Invest. 124, 4807–4822 (2014)
Thompson, L. F. et al. Crucial role for ecto-5′-nucleotidase (CD73) in vascular leakage during hypoxia. J. Exp. Med. 200, 1395–1405 (2004)
Stolen, C. M. et al. Absence of the endothelial oxidase AOC3 leads to abnormal leukocyte traffic in vivo. Immunity 22, 105–115 (2005)
Salmi, M. & Jalkanen, S. Cell-surface enzymes in control of leukocyte trafficking. Nat. Rev. Immunol. 5, 760–771 (2005)
Hayashi, S. & McMahon, A. P. Efficient recombination in diverse tissues by a tamoxifen-inducible form of Cre: a tool for temporally regulated gene activation/inactivation in the mouse. Dev. Biol. 244, 305–318 (2002)
Gordon, E. J., Gale, N. W. & Harvey, N. L. Expression of the hyaluronan receptor LYVE-1 is not restricted to the lymphatic vasculature; LYVE-1 is also expressed on embryonic blood vessels. Dev. Dyn. 237, 1901–1909 (2008)
Pham, T. H. et al. Lymphatic endothelial cell sphingosine kinase activity is required for lymphocyte egress and lymphatic patterning. J. Exp. Med. 207, 17–27 (2010)
Razani, B. et al. Caveolin-1 null mice are viable but show evidence of hyperproliferative and vascular abnormalities. J. Biol. Chem. 276, 38121–38138 (2001)
Herrnberger, L. et al. Lack of endothelial diaphragms in fenestrae and caveolae of mutant Plvap-deficient mice. Histochem. Cell Biol. 138, 709–724 (2012)
Squarzoni, P. et al. Microglia modulate wiring of the embryonic forebrain. Cell Reports 8, 1271–1279 (2014)
Clarke, R. L. et al. The expression of Sox17 identifies and regulates haemogenic endothelium. Nat. Cell Biol. 15, 502–510 (2013)
Kraal, G., Schornagel, K., Streeter, P. R., Holzmann, B. & Butcher, E. C. Expression of the mucosal vascular addressin, MAdCAM-1, on sinus-lining cells in the spleen. Am. J. Pathol. 147, 763–771 (1995)
Ganz, T. Macrophages and systemic iron homeostasis. J. Innate Immun. 4, 446–453 (2012)
Kohyama, M. et al. Role for Spi-C in the development of red pulp macrophages and splenic iron homeostasis. Nature 457, 318–321 (2009)
Swiers, G. et al. Early dynamic fate changes in haemogenic endothelium characterized at the single-cell level. Nat. Commun. 4, 2924 (2013)
Salmi, M., Karikoski, M., Elima, K., Rantakari, P. & Jalkanen, S. CD44 binds to macrophage mannose receptor on lymphatic endothelium and supports lymphocyte migration via afferent lymphatics. Circ. Res. 112, 1577–1582 (2013)
Hayashi, M., Schlesinger, D. H., Kennedy, D. W. & Yamada, K. M. Isolation and characterization of a heparin-binding domain of cellular fibronectin. J. Biol. Chem. 255, 10017–10020 (1980)
Vander Kooi, C. W. et al. Structural basis for ligand and heparin binding to neuropilin B domains. Proc. Natl Acad. Sci. USA 104, 6152–6157 (2007)
Ferrara, N. & Henzel, W. J. Pituitary follicular cells secrete a novel heparin-binding growth factor specific for vascular endothelial cells. Biochem. Biophys. Res. Commun. 161, 851–858 (1989)
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006)
Acknowledgements
We thank the following people for their expert technical assistance: E.-L. Väänänen, R. Sjöroos, S. Mäki, M. Pohjansalo, S. Tyystjärvi and P. Laasola. We thank J. Lilja and G. Jacquemet for advice. We acknowledge the Cell Imaging Core at the Turku Centre for Biotechnology, The Finnish Microarray and Sequencing Centre and the Laboratory of Electron Microscopy in University of Turku. The research was supported by grants from the Academy of Finland (to E.P., J.I. and M.S.), the Juselius Foundation (to P.R. and M.S.), the Cancer Foundation (to M.S.), the South-Western Regional Fund of the Finnish Cultural Foundation and the Foundation of Turku University (to P.R.) and the Satakunta Regional Fund of the Finnish Cultural Foundation (to N.J.).
Author information
Authors and Affiliations
Contributions
P.R. and N.J. contributed to the study design, and conducted most in vivo experiments and all FACS studies. E.L. and E.M. performed the whole-mount studies and the paraffin stainings, respectively. H.G. assisted with the in vivo experiments, and E.P. performed the ductal branching assays. J.I. planned, performed and analysed most biochemical experiments, and K.E. supervised and analysed the PLVAP–Fc generation and qPCR assays. K.A. conducted most of the confocal and all of the electron microscopy studies. M.S. conceived and supervised the study, planned experiments, analysed data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks D. Cheresh and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Full gating strategies for leukocyte subpopulations.
a, b, Gating strategies of adult (a) and fetal (b) macrophage and monocyte populations in the indicated tissues. The colour code of the final gates is the same as in Fig. 1 (orange, embryonic-derived macrophages in adults; black, bone-marrow-derived macrophages in adults; red, yolk-sac-derived macrophages in embryos; blue, fetal liver monocyte-derived macrophages in embryos; green, monocytes). The rightmost panels in adult lung (CD206 versus F4/80), adult peritoneal cavity (CD11b versus MHCII) and fetal liver (CD11b versus Ly6C) are validation stains for the indicated populations (not used for gating). The gating strategies for the other studied leukocyte populations are shown in Fig. 2c and Extended Data Fig. 5c (fetal liver macrophage–dendritic cell precursors, myeloid precursors, common monocyte progenitors, Ly6C+ and Ly6C− monocytes), Extended Data Fig. 2b (HSCs, common myeloid progenitors and common lymphoid progenitors in bone marrow), Extended Data Fig. 2d (Ly6Clow and Ly6Chigh bone marrow monocytes), Extended Data Fig. 2g (CD4+, CD8+ and B220+ lymphocytes in the adult organs), Extended Data Fig. 4b (EMPs and macrophages in the yolk sac), Extended Data Fig. 5d, e (EMPs and HSCs in fetal liver) and Extended Data Fig. 10d (mammary gland leukocytes). The rest of the validation gates are listed in Supplementary Table 1.
Extended Data Figure 2 Selective impairment in the accumulation of embryonic-derived tissue-resident macrophages in Plvap−/− mice.
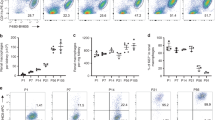
a, b, Flow cytometry analyses of adult bone-marrow-derived CD11b+F4/80intermediate tissue-resident macrophages (the black gate) in the colon and peripheral lymph nodes (PLN) (a), and HSCs (Lin−c-Kit+Sca-1+ cells), common myeloid (Lin−c-Kit+Sca-1−IL7R−; CMP) and common lymphoid (Lin−c-Kit+Sca-1lowIL7R+; CLP) progenitor cells in the bone marrow (BM) (b). c, Colony-forming assays on macrophage colony stimulating factor (M-CSF)-supplemented soft agar from bone marrow. d–g, Flow cytometry analyses of inflammatory (CD11b+Ly6Chigh) and patrolling (CD11b+Ly6Clow) monocytes in the bone marrow (d, green gates), recently entered tissue monocytes (CD11b+F4/80intermediateLy6Chigh and CD11b+F4/80intermediateLy6Clow cells, green gates) in the spleen (e) and liver (f) and CD4+, CD8+ and B220+ lymphocytes in the spleen, Peyer’s patches (PP), bone marrow (BM), liver, lung and blood (g) of adult mice. The flow cytometry data are shown as frequency of live-gated CD45+ cells (a, d), of live-gated Lin− cells (b, for HSC), of live-gated Lin−c-Kit+Sca-1−/low cells (b, for CMP and CLP), of live-gated CD45+B220−CD4−CD8−CD11b+F4/80intermediate cells (e, f), of live-gated CD45+ cells (g, for B cells) and of live-gated CD45+B220− cells (g, for CD4 and CD8 T cells). Each dot represents one mouse (pooled from 2–5 independent experiments, see Source Data), data are mean ± s.e.m. for each group.
Extended Data Figure 3 Normal seeding of embryonic-derived macrophages in Nt5e−/−, Aoc3−/− and Cav1−/− mice.
a, b, Flow cytometry analyses of embryonic-derived CD11b+F4/80high tissue-resident macrophages (orange gates) in the spleen, peritoneal cavity and liver, and of embryonic-derived CD11b+CD11chigh macrophages (orange gates) in the lungs of adult wild-type, Nt5e−/− and Aoc3−/− mice (a), and in wild-type and Cav1−/− mice (b). c, Electron micrographs of caveola in the fetal liver sinusoidal endothelium. Red arrow, a diaphragm-containing caveola; red arrowhead, a caveola without the diaphragm. d, Immunofluorescent stains of livers of wild-type, Cav1−/− and Plvap−/−mice at E16.5 for PLVAP and caveolin. e, Flow cytometry analyses of yolk-sac-derived (CD11b+F4/80high; red gates) and fetal liver-derived (CD11b+F4/80intermediate; blue gates) macrophages, and of CD11b+Ly6Chigh monocytes (green gate) in the blood in E16.5 wild-type and Cav1−/− mice. Shown are representative images (n = 2 (c) and n = 3 (d) biological replicates from 4 (d) independent stains). Scale bars, 50 nm (c) and 20 μm (d). The flow cytometry data are shown as frequency of live-gated CD45+B220−CD4−CD8− cells (a, b, adult tissues), and of CD45+B220− (b, adult blood and e). Each dot represents one mouse or embryo (pooled from 2 independent experiments and from 2–3 litters (e), see Source Data), data are mean ± s.e.m. for each group (*P < 0.05 by Mann–Whitney U-test).
Extended Data Figure 4 EMP and macrophage accumulation in the yolk sac and c-Kit+ cell accumulation in the AGM are intact in Plvap−/− mice.
a, Immunohistochemical analyses of PLVAP expression in whole-mounts of wild-type yolk sac at E8.5 and E10.5. Red arrows, PLVAP+ vascular endothelium. b, c, Flow cytometry analyses of live-gated CD41+CD45+c-KithighF4/80− EMP and CD45+c-Kit−F4/80+ macrophages (MAC) at E10.5 (b), and at E12.5 (c) in the yolk sac. d, Colony-forming assays on M-CSF-supplemented soft agar from E10.5 yolk sac. e, qPCR analyses of Plvap and Lyve1 expression in EMP of yolk sac in E10.5 wild-type mice (cells pooled from 30 embryos). Liver denotes mRNA from whole E12.5 fetal liver (a positive control). f, Whole-mount immunofluorescent stains of c-Kit, CD31 and PLVAP in the AGM region of E10.5 wild-type mice. CD31 is expressed in endothelial cells and HSC. White arrows, representative c-Kit+ cells. g, Whole-mount immunostains of c-Kit in the AGM region of E10.5 wild-type and Plvap−/− mice. Shown are representative images (a, f, g, n = 3 biological replicates from 2 independent stains). Scale bars, 50 μm (a, f, g). Each dot represents one embryo (pooled from 2–3 independent experiments and from 2–3 litters (b, c, d), see Source Data), data are mean ± s.e.m. for each group.
Extended Data Figure 5 Entry of monocyte progenitors to fetal liver and their differentiation to monocytes is PLVAP-independent.
a, Cytospin stains of sorted CD11b+F4/80high yolk-sac-derived macrophage-like cells and CD11b+F4/80intermediate fetal liver-derived monocyte-like cells from E14.5 fetal livers of wild-type and Plvap−/− mice (representative images from 4 embryos per genotype). Scale bars, 10 μm. b, qPCR analyses of sorted CD11b+F4/80high yolk-sac-derived macrophage-like cells (F4/80 Hi) and CD11b+F4/80intermediate fetal liver-derived monocyte-like cells (F4/80 Int) isolated from E16.5 livers of wild-type mice for cell-type signature genes Cx3cr1, Mrc1, Emr1, Flt3, Gata2, Ccr2, and Itgam (cells pooled from 10 embryos). c, The gating strategy to identify fetal liver CD11b−CSF-1R+c-Kit+Flt-3+Ly6C− macrophage-dendritic cell precursors (MDP, first P1, then the pink gate), CD11b−CSF-1R+c-Kit+Flt-3−Ly6C− fetal liver myeloid precursors (MP, first P2, then the blue gate), and CD11b−CSF-1R+c-Kit+Flt-3−Ly6C+ common monocyte progenitors (cMoP, first P2, then the violet gate) and their enumeration at E12.5, E13.5, E14.5, E15.5 and E16.5. The gating strategy to identify fetal liver CD11b+CSF-1R+c-Kit−Flt-3−Ly6C− monocytes (Ly6C− monocytes, first P3, then the brown gate) and CD11b+CSF-1R+c-Kit−Flt-3−Ly6C+ monocytes (Ly6C+ monocytes, first P3 then the green gate), and their enumeration at E13.5 and E15.5 livers is also shown (the quantification of these two monocyte types at E12.5, E14.5 and E16.5 livers are shown in Fig. 2d). The flow cytometry data are shown as frequency of live-gated CD45+ cells. d, Flow cytometry analyses of live-gated CD41+CD45+c-KithighF4/80− EMP cells at E12.5 in the livers. e, Flow cytometry analyses of live-gated Lin−c-Kit+Sca-1+ HSC at E12.5 in the livers. f, Colony forming assays on M-CSF supplemented soft agar from E12.5 liver single-cell suspensions. Each dot represents one embryo (2–3 independent experiments and from 2–3 litters (c–f), see Source Data), data are mean ± s.e.m. for each group (*P < 0.05 by Mann–Whitney U-test).
Extended Data Figure 6 Deletion of yolk-sac-derived, but not EMP and fetal liver monocyte-derived macrophages by an anti-CSF-1R antibody injection to E6.5 pregnant mice.
a, Flow cytometry analyses of CSF-1R (CD115) expression on live-gated CD41+CD45+c-KithighF4/80− early EMPs in E10.5 yolk sac (the two panels on the left) and on live-gated CD45+c-Kit−F4/80+ macrophages (MAC) in E10.5 yolk sac (the two panels on the right) in wild-type and Plvap−/− mice. Black histograms represent wild-type mice, red histograms represent Plvap−/− mice, grey histograms show isotype-matched negative-control antibodies. MFI, mean fluorescence intensity. b, The anti-CSF-1R (AFS) and control antibody (CTRL) treatment strategy and flow cytometry analyses of yolk-sac-derived, CD11b+F4/80high macrophages (red gates) and fetal liver-derived CD11b+F4/80intermediate macrophages (blue gates) at E14.5 in the brain and lung of wild-type and Plvap−/− embryos after a treatment of E6.5 pregnant Plvap+/− mice with a single dose of anti-CSF-1R antibody (AFS) or isotype-matched control antibody. c, The gating strategy and flow cytometry analyses of the frequencies of fetal liver macrophage–dendritic cell precursors, myeloid precursors, common monocyte progenitors, Ly6C+ and Ly6C− monocytes (gates as in Extended Data Fig. 5c and Supplementary Table 1) at E14.5 in the wild-type and Plvap−/− embryos of Plvap+/− dams, which were treated with the anti-CSF-1R antibody or isotype-matched control antibody at E6.5 of pregnancy. The flow cytometry data are shown as frequency of live-gated CD45+ cells (a), of live-gated CD45+B220− cells (b, lung), and of live-gated CD45+ cells (b, brain and c). Each dot represents one embryo (pooled from 2–3 independent experiments from 2–4 litters (a–c), see Source Data), data are mean ± s.e.m. for each group (*P < 0.05 by Mann–Whitney U-test).
Extended Data Figure 7 PLVAP in liver is induced early during fetal organogenesis and is selectively expressed in sinusoidal endothelial cells.
a, b, Immunohistological stains of formalin-fixed, paraffin-embedded fetal liver sections of wild-type (a) and Plvap−/− (b) mice with anti-PLVAP antibody MECA-32 at the indicated time points. White arrows, representative vessels (brown). c, d, qPCR analyses of Lyve1 expression in the liver in wild-type and Plvap−/− mice (c), and of Plvap and Lyve1 expression in sorted CD11b+F4/80intermediate and CD11b+F4/80high cells isolated from livers of E16.5 wild-type mice (cells pooled from 10 embryos). Liver denotes mRNA from whole E12.5 fetal liver (a positive control) (d). e–g, Immunofluorescent stains of E12.5, E14.5, and E16.5 fetal liver sections for PLVAP, F4/80 and LYVE-1 (e), E14.5 fetal liver for PLVAP and F4/80 (f), and E12.5, E14.5 and E16.5 liver for PLVAP and CD31 (another vascular endothelial marker) (g). Red arrow in f denotes an endothelial-penetrating protrusion of an F4/80+ myeloid cell. h, Immunofluorescent stains of B6;129 and BALB/c wild-type livers at E14.5 for PLVAP and LYVE-1. i, Immunofluorescent stains of human fetal liver (week 18) with an anti-PLVAP antibody (PAL-E). Shown are representative images (n ≥ 3 (a, b, e–h) and n = 1 (i) biological replicates from 2 independent stains). Scale bars, 20 μm (a, b, e–i). In c, each dot represents one embryo (pooled from 2 independent experiments and from 2–3 litters, see Source Data), data are mean ± s.e.m. of each group.
Extended Data Figure 8 Time-selective and cell type-selective knock-down of PLVAP results in accumulation of fetal liver monocytes.
a, The timing of single tamoxifen injections and tissue collections with PlvapF/F; CAGGCre-ERTM mice. b, Quantification of Plvap mRNA synthesis in fetal livers by qPCR 2 days after the tamoxifen treatments. c, Immunofluorescent stains of PLVAP and LYVE-1 in the fetal liver at E13.5, E14.5 and E15.5 (2 days after the tamoxifen pulse given on E11.5, E12.5 and E13.5, respectively). d, The gating strategy and flow cytometry analyses of the frequencies of fetal liver macrophage-dendritic cell precursors, myeloid precursors, common monocyte progenitors, Ly6C+ and Ly6C− monocytes, (gates as in Extended Data Fig. 5c and Supplementary Table 1) at E13.5, E14.5 and E15.5 in PlvapF/F (control) and PlvapF/F; CAGGCre-ERTM mice (in each case 2 days after the tamoxifen injection). e, Immunohistological analyses of PLVAP protein expression in the E14.5 fetal liver of PlvapF/F;Lyve1-Cre and control (PlvapF/F) mice. f, Flow cytometry analyses of macrophage–dendritic cell precursors, myeloid precursors, common monocyte progenitors, Ly6C+ and Ly6C− monocytes (defined as in d) at E13.5 and E14.5 in the fetal liver of the control (PlvapF/F) and PlvapF/F; Lyve1-Cre mice. Shown are representative images (n = 2 (c) and n = 4 (e) biological replicates from 2 independent stains). Scale bars, 20 μm (c, e). The flow cytometry data are shown as frequency of live-gated CD45+ cells. Each dot represents one embryo (pooled from 2–3 independent experiments and from 2–4 litters (b, d, f), see the Source Data), data are mean ± s.e.m. for each group (*P < 0.05 by Mann–Whitney U-test).
Extended Data Figure 9 Interactions of PLVAP with heparin and VEGF, and VEGF receptor expression on fetal liver monocytes.
a, Immunofluorescent analysis of VEGF and Plvap expression in E14.5 liver of wild-type mice (n = 3). Scale bar, 10 μm. b, Proximity ligation assays (PLAs) in E14.5 fetal livers. Shown are PLA signals (small dots, white) between VEGF and PLVAP (leftmost) and between a negative control protein (CO) and Plvap (third image from the left). In the merged images, total PLVAP expression determined by immunohistochemistry (immuno PLVAP, green) and nuclear stains (DAPI, blue) are displayed in addition to the PLA signals. Representative images from 3 independent experiments are shown. Scale bars, 10 μm. c, Schematic depiction of mouse PLVAP protein and the PLVAP–Fc fusion protein. Cyt, cytoplasmic domain; TM, transmembrane domain. Numbers represent amino acids. d, Pull-down assays analysing the affinity of binding of PLVAP–Fc and CD4–Fc (a negative control) fusion proteins to heparin (Hep)- and streptavidin (Sa)-affinity (negative control) beads. The bead-bound proteins were eluted in Laemmli’s sample buffer (the immunoblot on the left; this is the full image and different exposure of Fig. 3g), or in 1.0 M NaCl (the immunoblot on the right). The released proteins were separated in SDS–PAGE under reducing conditions and visualized using immunoblotting for the Fc-tail. Aliquots of PLVAP–Fc (loading Plvap–Fc) were used as loading controls. Representative blots from two independent assays are shown. e, Pull-down assays analysing the binding of PLVAP–Fc (20 μg) to heparin-beads in the absence (−) and presence of competing proteins fibronectin (FN, 100 μg) and collagen (ColI, 100 μg). The bound proteins were eluted, separated in SDS–PAGE under reducing conditions and visualized using immunoblotting for the Fc-tail. f, g Flow cytometry analyses of neuropilin-1 expression on live-gated CD41+CD45+c-KithighF4/80− EMP cells in E12.5 livers (f), and on macrophage-dendritic cell precursors, myeloid precursors, common monocyte progenitors, Ly6C+ and Ly6C− monocytes during monocytopoiesis in E14.5 livers (g) (gates as in Extended Data Fig. 5c and Supplementary Table 1) of wild-type and Plvap−/− mice. h, i, Flow cytometry analyses of VEGFR1 (h) and VEGFR2 (i) expression on cMoP, Ly6C+ and Ly6C− monocytes in E14.5 livers. The flow cytometry data are shown as frequency of live-gated (f) and live-gated CD45+ (h, i) cells. Each dot represents one embryo (pooled from 2–3 independent experiments and from 2–3 litters (f–i), see Source Data), data are mean ± s.e.m. for each group.
Extended Data Figure 10 Normal yolk-sac macrophage-dependent morphogenesis in Plvap−/− mice and the function of PLVAP during macrophage ontogenesis.
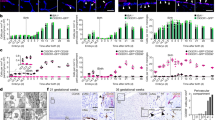
a, Haematoxylin–eosin stains of coronal sections of whole E14.5 embryos from wild-type and Plvap−/− mice. White arrows, neural tubes; red arrows, livers. Inset, the developing bronchial tree in the lungs. Scale bars, 2 mm (main image), 200 μm (inset). b, Macroscopic images of toes in wild-type and Plvap−/− embryos at E16.5. Scale bars, 1 mm. c, Prussian blue stains of livers, and quantification of Fe3+-containing cells (blue) in 5-week-old mice. Scale bars, 200 μm (main image), 50 μm (inset). d, The gating strategy for CD11b+F4/80+ cells in the mammary fat pad. e, Flow cytometry analyses of CD4+ T-helper cells and B220+ B lymphocytes in the mammary fat pads of wild-type and Plvap−/− mice. Shown are representative images (a–c; n = 3 biological replicates). Flow cytometry data are shown as frequency of live-gated CD45+ leukocytes. Each dot represents one mouse embryo (pooled from 2 independent experiments, see the Source Data), data are mean ± s.e.m. for each group. f, A schematic model depicting the organ-selective role of PLVAP in the seeding of fetal liver monocyte-derived tissue-resident macrophages. The yolk-sac-derived tissue-resident macrophages and the progenitors (EMP and HSC) of fetal liver monocytes develop and seed normally in the absence of PLVAP. By contrast, PLVAP supports the egress of fetal liver-derived monocytes to the blood, and thereby the seeding of fetal liver-derived tissue-resident macrophages in different tissues. PLVAP fibrils form the fenestral diaphragms (blue) in fetal liver sinusoidal endothelial cells (LSEC). The seeding of bone-marrow-derived monocytes and macrophages after birth is PLVAP-independent (not shown). In the inset, the molecular interactions of PLVAP with heparin, neuropilin-1 (NP-1) and VEGF in the fetal liver and the expression of neuropilin-1 and VEGFR1 by E14.5 fetal liver monocytes (MO) are depicted. Although it remains to be experimentally tested, it is likely that PLVAP–heparin complexes at the diaphrams of fetal LSEC have the potential to regulate monocyte egress by providing an adhesive substrate for fetal liver monocytes (for example, via neuropilin-1, and probably via other molecules as well), and/or by immobilizing chemoattractants (for example, VEGF, and possibly other heparin-binding chemotactic molecules). The possible selective role of PLVAP in the vasculature of target organs during the seeding of fetal liver-derived monocytes also remains to be tested.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 (gel source data) and Supplementary Tables 1-3. (PDF 1245 kb)
PLVAP is selectively expressed in the vasculature of E9.5 embryos.
Whole-mounts of E9.5 wild-type embryos were immunochemically stained (single stainings) for Plvap (white) and CD31 (red), optically cleared and imaged using spinning disk microscopy to produce z-stacks. The video shows horizontally rotated 3D reconstructions of the whole embryos (for Plvap the x-, y- and z-dimensions are 1298 μm, 1298 μm and 103 μm, respectively, and for CD31 1298 μm, 1298 μm and 183 μm, respectively). The videos show that PLVAP expression is restricted to the vasculature in a similar manner than that of another vascular marker CD31. The video is representative of three biological replicates with similar results. (AVI 10259 kb)
In fetal liver PLVAP is selectively expressed in sinusoidal endothelial cells
Whole-mounts of E14.5 wild-type livers were immunochemically stained for PLVAP (white) and LYVE-1 (red), optically cleared and imaged using spinning disk microscopy to produce z- stacks. The video shows horizontally and vertically rotated 3D econstructions of the fetal liver (the x-, y- and z-dimensions are 323 μm, 323 μm and 50 μm, respectively). The video is representative of three biological replicates with similar results. (AVI 55542 kb)
Source data
Rights and permissions
About this article
Cite this article
Rantakari, P., Jäppinen, N., Lokka, E. et al. Fetal liver endothelium regulates the seeding of tissue-resident macrophages. Nature 538, 392–396 (2016). https://doi.org/10.1038/nature19814
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19814
This article is cited by
-
The role of PLVAP in endothelial cells
Cell and Tissue Research (2023)
-
Oncofetal reprogramming in tumour development and progression
Nature Reviews Cancer (2022)
-
Plasmalemma vesicle-associated protein promotes angiogenesis in cholangiocarcinoma via the DKK1/CKAP4/PI3K signaling pathway
Oncogene (2021)
-
Embryonic macrophages function during early life to determine invariant natural killer T cell levels at barrier surfaces
Nature Immunology (2021)
-
Generation, localization and functions of macrophages during the development of testis
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.