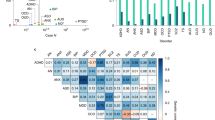
Abstract
The National Institute of Mental Health (NIMH) has made sustained investments in the development of genomic resources over the last two decades. These investments have led to the development of the largest biorepository for psychiatric genetics as a centralized national resource. In the realm of genomic resources, NIMH has been supporting large team science (TS) consortia focused on gene discovery, fine mapping of loci, and functional genomics using state-of-the-art technologies. The scientific output from these efforts has not only begun to transform our understanding of the genetic architecture of neuropsychiatric disorders, but it has also led to a broader cultural change among the investigator community towards deeper collaborations and broad pre-publication sharing of data and resources. The NIMH supported efforts have led to a vast increase in the amount of genetic and genomic resources available to the mental health research community. Here we provide an account of the existing resources and estimates of the scale and scope of what will be available in the near future. All biosamples and data described are intended for broad sharing with researchers worldwide, as allowed by the subject consent and applicable laws.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Bloom DE, Cafiero E, Jané-Llopis E, Abrahams-Gessel S, Bloom LR, Fathima S et al The Global Economic Burden of Noncommunicable Diseases. World Economic Forum and the Harvard School of Public Health, 2011 [online], Available at: http://www3.weforum.org/docs/WEF_Harvard_HE_GlobalEconomicBurdenNonCommunicableDiseases_2011.pdf.
Eaton WW, Martins SS, Nestadt G, Bienvenu OJ, Clarke D, Alexandre P . The burden of mental disorders. Epidemiol Rev 2008; 30: 1–14.
Lehner T, Senthil G, Addington AM . Convergence of advances in genomics, team science, and repositories as drivers of progress in psychiatric genomics. Biol Psychiatry 2015; 77: 6–14.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Moldin SO . NIMH Human Genetics Initiative: 2003 update. Am J Psychiatry 2003; 160: 621–622.
Parikshak NN, Luo R, Zhang A, Won H, Lowe JK, Chandran V et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 2013; 155: 1008–1021.
Shcheglovitov A, Shcheglovitova O, Yazawa M, Portmann T, Shu R, Sebastiano V et al. SHANK3 and IGF1 restore synaptic deficits in neurons from 22q13 deletion syndrome patients. Nature 2013; 503: 267–271.
Wang Y, Dolmetsch R . In vitro human corticogenesis. Neuron 2013; 77: 379–381.
He X, Sanders SJ, Liu L, De Rubeis S, Lim ET, Sutcliffe JS et al. Integrated model of de novo and inherited genetic variants yields greater power to identify risk genes. PLoS Genet 2013; 9: e1003671.
Ionita-Laza I, Lee S, Makarov V, Buxbaum JD, Lin X . Sequence kernel association tests for the combined effect of rare and common variants. Am J Hum Genet 2013; 92: 841–853.
Psychiatric GWAS Consortium Steering Committee. A framework for interpreting genome-wide association studies of psychiatric disorders. Mol Psychiatry 2009; 14: 10–17.
Buxbaum JD, Daly MJ, Devlin B, Lehner T, Roeder K, State MW et al. The autism sequencing consortium: large-scale, high-throughput sequencing in autism spectrum disorders. Neuron 2012; 76: 1052–1056.
De Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 2014; 515: 209–215.
Parikshak NN, Gandal MJ, Geschwind DH . Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat Rev Genet 2015; 16: 441–458.
Willsey AJ, Sanders SJ, Li M, Dong S, Tebbenkamp AT, Muhle RA et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 2013; 155: 997–1007.
Nichols L, Freund M, Ng C, Kau A, Parisi M, Taylor A et al. The National Institutes of Health Neurobiobank: a federated national network of human brain and tissue repositories. Biol Psychiatry 2014; 75: e21–e22.
PsychENCODE Consortium PsychENCODE Consortium Akbarian S, Liu C, Knowles JA, Vaccarino FM, Farnham PJ et al. The PsychENCODE project. Nat Neurosci 2015; 18: 1707–1712.
Shabani M, Knoppers BM, Borry P . From the principles of genomic data sharing to the practices of data access committees. EMBO Mol Med 2015; 7: 507–509.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016; 536: 285–291.
Acknowledgements
We thank Dr John Rice and his group at Washington University in St Louis; Drs Jay Tischfield and Linda Brzustowicz, and their group at Rutgers University; Dr Yigal Arens and his group at the Information Sciences Institute at University of Southern California, and the dbGaP and NDA staff for their help with data and sample accounting. We also acknowledge the efforts of several members of our TS and functional genomics consortia in providing up to date details on their data generation efforts, including Drs Pamela Sklar and Nenad Sestan.
Disclaimer
The views and opinions expressed in this article are those of the authors and should not be construed to represent the views of any of the sponsoring organization, agencies or the United States government.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
About this article
Cite this article
Senthil, G., Dutka, T., Bingaman, L. et al. Genomic resources for the study of neuropsychiatric disorders. Mol Psychiatry 22, 1659–1663 (2017). https://doi.org/10.1038/mp.2017.29
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2017.29
This article is cited by
-
A framework for the investigation of rare genetic disorders in neuropsychiatry
Nature Medicine (2019)
-
NIMH neuropsychiatric genomics: crucial foundational accomplishments and the extensive challenges that remain
Molecular Psychiatry (2017)