Abstract
In the present study we investigated the molecular mechanisms regulating the expression of RAR-related orphan receptor gamma t (RORγt), the central factor controlling interleukin (IL)-17 transcription and Th17 differentiation. In key studies, we found that cells from mice with major deletions of E-protein transcription factors, E2A and HEB, display greatly reduced RORγt/IL-17 expression and that E-protein-deficient mice exhibit greatly diminished IL-17-dependent inflammation in experimental allergic encephalitis models. In additional studies, we unexpectedly found that cells from mice with deletion of Id3, a protein that inhibits E-protein binding to DNA, display diminished RORγt/IL-17 expression and mice deficient in this protein exhibit decreased Th17-mediated inflammation in a cell-transfer colitis model. The explanation of these initially paradoxical findings came from studies showing that Id3 deficiency leads to increased IL-4-induced GATA-3 expression, the latter a negative regulator of RORγt transcription; thus, increased Id3 expression likely has a net positive effect on RORγt expression via its inhibition of IL-4 production. Finally, we found that both E-proteins and Id3 are upregulated in tandem by the cytokines that induce Th17 differentiation, transforming growth factor-β, and IL-6, implying that these transcription factors are critical regulators of Th17 induction.
Similar content being viewed by others
Introduction
In recent years, there has been considerable progress in our understanding of the molecular factors regulating the Th17 response. However, most of this understanding centers around the regulation of interleukin (IL)-17 transcription by RAR-related orphan receptor gamma t (RORγt) and relatively little around the regulation of RORγt itself. Thus, while it is known that Runx1,1 Stat3,2, 3 and perhaps BATF4 exert a positive influence on RORγt transcription, it is not clear how these factors are related to the two cytokines known to upregulate RORγt, namely transforming growth factor (TGF)-β and IL-6. The same can be said for hypoxia-inducible factor-1α, a factor with a positive effect on RORγt transcription that is induced by stress stimuli that do not necessarily include TGF-β and IL-6.5, 6, 7
One class of transcription factors that has not been thoroughly explored as to their possible role in Th17 differentiation are the E-proteins, a family of proteins whose relation to Th17 is suggested by studies showing that members of this family regulate RORγt expression during thymocyte development.8, 9 E-proteins are homo- or heterodimeric transcriptional activators (or repressors) whose function is downregulated by interaction with any of the four Id proteins that inhibit their binding to DNA and thus their transcriptional activity.10, 11, 12 In studies of thymic development it has been shown that interplay between E-proteins and Id proteins have important roles in the regulation of thymocyte differentiation at various thymocyte differentiation checkpoints and in shaping the fate of naive cells.13, 14, 15 In part, such a regulation involves E-protein effects on RORγt expression as it has been shown that RORγt is decreased in E-protein-deficient CD4+/CD8+ thymocytes.9 In addition, pre-T cell receptor signaling induces E-protein-dependent RORγt expression and this inhibits cell proliferation while promoting gene rearrangement.8 With respect to peripheral effects of E-proteins, it has been shown that Id2-mediated suppression of E-proteins is necessary for lymphoid tissue inducing cell and mature natural killer cell development;16 in addition, E-proteins may regulate the development of effector-memory CD8+ T cells as the latter are deficient in mice lacking Id3.17 However, there is no evidence presently available indicating that these negative peripheral effects of E-proteins involve RORγt. In the present study we show that E-proteins (E2A and HEB), acting in conjunction with Id3, positively regulate RORγt expression in peripheral T cells and that such regulation is a critical factor in the induction of Th17 differentiation.
Results
E-proteins are required for RORγt promoter–reporter activity
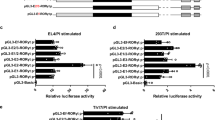
Prompted by the fact that RORγt expression is decreased in thymocytes of E2A-null mice,9 in initial studies we explored how E-proteins affect RORγt promoter activity in luciferase assays. To this end, we generated a PGL4 luciferase reporter plasmid containing RORγt promoter fragments of 2 kb (−2 kb—+109), 1.4 kb(−1.4 kb—+109), and 0.3 kb(−0.3 kb—+109) and then transfected the reporter constructs into EL4 (LAF) cells or primary CD4+ T cells. We found that while the 0.3- and 1.4-kb fragments gave rise to minimal promoter-reporter activity, the 2-kb fragment gave rise to high activity in both EL4 cells (Figure 1a) and primary CD4+ T cells (Figure 1b).
E-protein is required for RORγt promoter luciferase reporter activity. (a, b) A 2-kb mouse RORγt promoter fragment is required for maximal promoter activity. RORγt promoter fragments of various sizes inserted into luciferase reporter constructs were co-transfected with Renilla luciferase plasmid into EL4 (LAF) cells (a) or primary CD4+ T cells (b); following transfection, cells were stimulated with PMA/ionomycin (P/I) (a) or anti-CD3/anti-CD28 (b) overnight after which luciferase activity was measured and normalized by Renilla luciferase activity. (c) E-box binding sites are essential for RORγt promoter–reporter activity. A wild-type (WT) 2-kb RORγt promoter construct without and with mutated E-box binding sites at −1.8 or −1.6 kb were transfected into EL4 cells. Following transfection, cells were stimulated overnight with P/I after which luciferase activity was measured and normalized as above. (d) E-box binding sites E3 and E4 are also essential for RORγt promoter–reporter activity. A WT 2-kb RORγt promoter construct without and with mutated E-box binding sites E3 and E4 were transfected into EL4 cells. Following transfection, cells were stimulated overnight with P/I after which luciferase activity was measured and normalized as above. (e) E-protein upregulates RORγt promoter–reporter activity. The indicated RORγt promoter–reporter constructs were co-transfected with E47 or HEB expression plasmids or empty vector into EL4 cells and cultured overnight after which luciferase activity was measured and normalized as above. Data shown are derived from mean±s.d. of duplicate transfections in each experiment and are representative of at least three independent experiments. (f, g) E-proteins bind to the RORγt promoter in primary Th17 cells. Chromatin from purified naive CD4+ T cells, CD4+ T cells polarized under Th0 conditions or Th17 conditions were subjected to chromatin immunoprecipitation analysis for the detection of binding of E-proteins E47 (g) and HEB (f) with anti-E47 and anti-HEB, respectively. Results are representative of three independent experiments in each case. Data shown are mean±s.d. of duplicate PCRs.
DNA sequencing analysis of the RORγt promoter region revealed several consensus E-protein binding sites (E-boxes) that are conserved among mouse, rat, and human RORγt promoters.1 We found that mutation of the E-box at −1.6 kb had no effect on promoter activity, whereas mutation of the E-box at −1.8 kb reduced promoter activity to the level seen with the 0.3-kb promoter fragment (Figure 1c). In addition, mutation of the E3 and/or E4 E-boxes 1 close to the transcription start site also led to reduced promoter activity of the 2-kb fragment (Figure 1d, Supplementary Figure 1a online). Thus, several E-boxes act in tandem to support optimal RORγt promoter-reporter activity.
In further studies, we co-transfected reporter constructs with E-protein expression vectors. We found that co-transfection of the 2- and 0.3-kb RORγt promoter–reporter constructs along with either an E47 or HEB expression vector led to substantial increases in luciferase signal as compared with co-transfection of empty vector (Figure 1e). In contrast, co-transfection of E-protein expression vectors with a 2-kb promoter–reporter bearing various E-box binding site mutations failed to reverse the complete or partial loss of the luciferase signal (Supplementary Figure 1b). Collectively, the data derived from the above promoter–reporter studies suggested that E-proteins do have the potential to regulate RORγt transcription.
E2A and HEB act as transcription factors that regulate RORγt and IL-17 expression
The results of the above luciferase reporter studies led us to investigate if E-proteins act as RORγt transcription factors in developing Th17 cells. In initial studies we performed chromatin immunoprecipitation assays in CD4+ T cells and found that both E2A and HEB bind to the RORγt promoter in Th17 cells but not in naive CD4+ cells or Th0 cells (Figure 1f). Thus, E-proteins do in fact bind to the RORγt promoter in physiologic cells, but only in the context of Th17 differentiation.
To determine if deletion of E-proteins affected RORγt transcription, we next performed gene knockdown studies using small interfering RNA (siRNA). Three different siRNAs for each gene were employed, of which at least two resulted in substantial reduction of mRNA expression (Supplementary Figure 2a,b). We found that CD4+ T cells cultured under Th17 conditions transfected with E2A or HEB siRNA (but not scrambled siRNA) resulted in substantial attenuation of RORγt transcription and IL-17 expression at both the RNA (Figure 2a) and protein level (Figure 2b). These findings are in agreement with previous studies of the regulation of RORγt transcription and IL-17 expression by E2A siRNA.18
E-proteins are required for Th17 cell differentiation. (a, b) Purified naive CD4+ T cells were transfected with E2A small interfering RNA (siRNA) or control scrambled siRNA. Cells were then cultured under Th17 conditions after which RORγt (a) expression and interleukin (IL)-17 production were measured by reverse transcriptase (RT)–PCR and flow cytometry (a, b). (c, d) Purified naive CD4+ T cells were transfected with HEB siRNA and control scrambled siRNA. Cells were cultured under Th17 conditions after which RORγt (c) and IL-17(c, d) were measured as indicated above. Data above are representative of at least three independent experiments. (e) ER-Cre+ and wild-type (WT) mice were administered tamoxifen by gavage six times over a period of 30 days. Purified naive CD4+ T cells were cultured under Th17 conditions after which they were analyzed for E2A, HEB, RORγt, and IL-17 mRNA expression by real time RT–PCR. Data are representative of three independent experiments. Error bars represent s.d. of duplicated PCRs. (f) ER-Cre+ and WT mice were administered tamoxifen by gavage six times over a 1- month period. Purified naive CD4+ T cells obtained from these mice 1 week after the final dose of tamoxifen were labeled with CSFE and cultured under Th17 conditions for 3 days after which IL-17 expression was measured by flow cytometry. Data are representative of three independent experiments. (g) Summary percentage of IL-17 decrease and percentage of E-protein decrease in Th17 cells from mice subjected to tamoxifen administration regimens. Data are derived from at least three independent experiments. In each case, the percentage of IL-17+ cells for WT mice were set as 100% and each dot represents data from one mouse. Bar represents mean value. ***P<0.001. (h) Purified CD4+ T cells from WT Cre+ or E2Af/+HEBf/fCD4Cre+ mice treated with tamoxifen as in Figure 2e were cultured under Th17 conditions; IL-17 expression was measured by flow cytometry. Data are representative of two independent experiments. (i) Purified naive CD4+ T cells from WT or E2Af/fHEBf/fCD4Cre mice were cultured under Th17 conditions for 3 days; IL-17 expression was measured by flow cytometry. Data are representative of two independent experiments.
Next, in more definitive studies of the affect of E-protein deletion, we examined RORγt and IL-17 expressions in ER-Cre-inducible E2A and HEB conditional knockout mice (E2Af/fHEBf/fER-Cre mice).13 Accordingly, we administered tamoxifen by gavage to ER-Cre+ and wild-type (WT) mice every other day for 10 days (five times) and then, 1 week after the final tamoxifen administration, we obtained purified naive CD4+ T cells from the mice and cultured them under Th17 conditions in the presence of 4-OH tamoxifen to evaluate the effects of deletion on Th17 differentiation. We found that cells from tamoxifen-treated ER-Cre+ mice exhibited about a 50% decrease in RORγt and IL-17 mRNA levels (Supplementary Figure 2c,e) as well as similar reduction in the frequency of IL-17 producing cells (Supplementary Figure 2d) as compared with tamoxifen-treated WT mice. Parenthetically, deletion of E2A or HEB had no effect on Foxp3 expression in the cells cultured under Th17 conditions (Supplementary Figure 2d).
To achieve more profound E-protein deletion, we next subjected E-protein conditional KO and WT mice to relatively long-term tamoxifen administration. Accordingly, we administered tamoxifen six times over a period of 1 month and then, 6 days after the last injection, again cultured purified naive CD4+ spleen T cells under Th17 conditions to evaluate effects of deletion on Th17 differentiation. Indeed, we found that this regimen led to a ∼75–80% decrease in the expression of E2A and HEB mRNA, accompanied by similar decreases in RORγt and IL-17 mRNA in cells from ER-Cre+ mice compared with WT mice (Figure 2e). In addition, the frequency of IL-17-producing cells was also reduced by about 75–80% in the conditional KO mice (Figure 2f). Finally, it should be noted that E-protein mRNA decreases had a slightly greater effect on IL-17 expression than on RORγt expression; this could be due to the fact that IL-17 transcription is highly sensitive to decreases in RORγt.
Comparison of data derived from individual E-protein floxed mice (with partial or complete E2A or HEB deletions) in the long and short tamoxifen administration regimens described above (and shown in Figure 2e as well as in Supplementary Figure 2c,e) revealed that the percentage decrease in functional E-protein (a heterodimer of E2A and HEB that reflects both E2A and HEB mRNA levels) correlated with the percentage decrease in the number of IL-17-expressing cells. Thus, these the data strongly suggest that E-protein expression is in fact necessary for RORγt and subsequent IL-17 expression.
It has been reported that both E-proteins and RORγt are involved in regulating cell proliferation. To investigate this possibility, purified naive CD4+ T cells from mice subjected to tamoxifen treatment were labeled with carboxyfluorescein succinimidyl ester and cultured under Th17 conditions. We found no significant changes in cell proliferation between WT and conditional KO cells (Figure 2f lower panel), whereas IL-17 expression underwent a very substantial decrease (Figure 2f upper panel). These results thus indicate that the effect of E-protein deletion on IL-17 expression is not due to an effect on cell proliferation. Finally, the above data on tamoxifen-mediated Cre-induced deletion of E-proteins was not an artifact of Cre toxicity as IL-17-producing cells were greatly reduced in E2Af/fHEBf/fCre+ Th17 cells compared with that of WT Cre+ Th17 cells (Figure 2h).
Th17 differentiation in E2Af/fHEBf/fCD4Cre mice
To verify the findings described above in mice with conditional E2A/HEB deletions, we then determined Th17 differentiation in E2Af/fHEBf/fCD4Cre mice. We found that purified naive CD4+ T cells obtained from such mice and cultured under Th17 conditions give rise to greatly decreased numbers of IL-17-producing cells as compared with that of WT mice (Figure 2i). Importantly, the deletion efficiency of E2A and HEB in CD4Cre+ mice (75–80%) was similar to that obtained with the 1-month tamoxifen injection regimen (Supplementary Figure 2f). These results again indicate that deletion of E2A and HEB leads to impairment of Th17 differentiation.
E-protein control of RORγt transcription is regulated by Id proteins
As mentioned above, Id proteins are a family of transcription cofactors that have been shown to bind to E-proteins and thereby inhibit the transcription activity of the latter by interfering with their DNA-binding activity.11, 12 Id2 and Id3 are the main Id family members involved in the regulation of E-protein function during lymphoid development19, 20 and indeed we found that both of these Id proteins are expressed in Th17 cells (data not shown). We therefore conducted studies of Id protein effects on RORγt expression–utilizing cells from mice with Id protein deletion, focusing mainly on mice with Id3 deletion (Id3−/− mice) as it has been shown that mice with Id2 deletion exhibit Id3 upregulation and could thus yield equivocal results.17
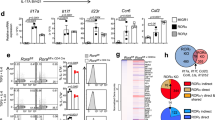
In initial studies along these lines, naive CD4+ T cells purified from Id3−/− and WT control mice were cultured under Th17 conditions for 3 days at which time IL-17 expression levels were measured by flow cytometry. Surprisingly, we observed that IL-17 expression in the Id3−/− Th17 cells was decreased rather than increased compared with WT Th17 cells (Figure 3a left panels and Figure 3b). We reasoned that this paradoxical result could have been due to the fact that the absence of Id3 led to the induction of an inhibitory factor normally suppressed by Id3 that has a countervailing negative effect on RORγt and IL-17 expression. One such possible inhibitory factor is IL-4, as production of the latter has been shown to be increased in Id3−/− mice13 and, in addition, has been shown to downregulate IL-17 production.21 Indeed, we found that cells from Id3−/− mice cultured under Th17 conditions in the presence of anti-IL-4 exhibited increased IL-17 expression (Figure 3a). Furthermore, RORγt (Figure 3c) and IL-17 mRNA (Figure 3d) were also significantly increased in Id3−/− Th17 cells containing anti-IL-4, whereas GATA-3 levels were significantly decreased (Figure 3a right panels and Figure 3e). These results thus suggested that IL-4 itself or GATA-3 induced by IL-4 does in fact inhibit RORγt expression in the absence of Id3.
Id3 regulates Th17 cell differentiation. (a) Naive CD4+ T cells purified from Id3−/− and wild-type (WT) littermate control mice were cultured under Th17 conditions or Th17 conditions plus the addition of anti-IL-4 (5 μg ml−1, 10 μg ml−1) for 3 days. Interleukin (IL)-17 was analyzed by flow cytometry. Data are representative of three independent experiments. (b) Naive CD4+ T cells purified from individual Id3−/− and WT littermate control mice were cultured under Th17 conditions or Th17 conditions plus anti-IL-4 (5 μg ml−1, 10 μg ml-1) for 3 days; IL-17 was analyzed by flow cytometry after which the percentage of IL-17+ cells for each mouse was plotted. Each dot represents data from one mouse Data are pooled from two independent experiments. (c–e) Naive CD4+ T cells purified from Id3−/− and WT littermate control mice were cultured under Th17 conditions or Th17 conditions plus the addition of anti-IL-4 (5 μg ml-1, 10 μg ml-1) for 3 days. RORγt (c), IL-17 (d), and GATA-3 (e) expression were measured by real-time reverse transcriptase–PCR. ***P<0.001. Data are representative of three independent experiments. (f–h) Development of colitis in Rag2−/− mice transferred with naive CD4+ T cells from WT or Id3−/− mice. (f) Change in body weight after cell transfer; (g) hematoxylin and eosin staining of colon tissue sections from Rag2−/− mice 5 weeks after cell transfer; (h) IL-17 and interferon-γ production by mesenteric lymph node cells of Rag2−/− mice stimulated with anti-CD3/28; cytokines were measured in culture supernatants 48 h after stimulation by ELISA.
In further studies to investigate the effect of IL-4 in Id3−/− mice on Th17 cell differentiation, we found that naive CD4+ T cells from Id3−/− mice produced increased amounts of IL-4 mRNA as compared with WT cells, but this increase was not accompanied by an increase in GATA-3 mRNA expression (Supplementary Figure 3a). In contrast, CD4+ T cells from Id3−/− mice cultured (and activated) under both Th0 and Th17 conditions expressed increased amounts of IL-4 mRNA compared with WT cells (Supplementary Figure 3b upper panels) and in this case GATA-3 mRNA was also expressed (Supplementary Figure 3a right panel and Figure 3b lower panels).
To determine if the above effects of Id3 deletion on Th17 responses also obtain in vivo, we next examined IL-17 expression in Id3−/− mice with cell-transfer colitis. Accordingly, we monitored colitis development in RAG2−/− mice transferred purified CD4+/CD45Rbhi CD4+ T cells from Id3−/− or WT mice. We found that recipients of Id3−/− CD4+ T cells developed less severe colitis than recipients of WT T cells as indicated by both weight loss and colonic morphology (Figure 3f). In addition, anti-CD3/CD28-stimulated T cells from the mesenteric nodes of mice transferred WT cells produced significantly greater amounts of IL-17 than T cells from mice transferred Id3−/− cells, whereas, in contrast, these T cell populations produced equivalent amounts of interferon-γ (Figure 3h). With the caveat that Id3 has multiple effects on lymphoid cell development in addition to those on Th17 differentiation, these results thus provided in vivo validation of our in vitro observation that cells from Id3−/− mice produce lower amounts of IL-17 than cells from WT mice.
In additional studies focusing on the effects of IL-4 and GATA- on Th17 differentiation, we found first that WT CD4+ T cells cultured under Th17 conditions in the presence of exogenous IL-4 exhibited decreased RORγt and IL-17 mRNA expression and increased GATA-3 mRNA expression (Figure 4a–c), as well as a decreased frequency of IL-17-producing cells (Figure 4d). Second, developing Th17 cells transduced with a retrovirus expressing GATA-3 have a much lower frequency of IL-17-producing cells and RORγt expression as compared with cells transduced with empty vector (Figure 4e). Third, RORγt promoter–reporter activity but not IL-17 promoter–reporter activity was inhibited by co-transfection of a GATA-3 expression plasmid (Figure 4g). These studies thus provide further support for the idea that IL-4 and GATA-3 are negative regulators of E-protein/Id protein–mediated Th17 differentiation.
GATA-3 inhibits Th17 differentiation. (a–d) Purified CD4+ T cells from wild-type (WT) mice were cultured under Th17 conditions or Th17 conditions plus interleukin (IL)-4 (10, 20, and 40 ng ml−1) for 3 days. RORγt (a), IL-17 (b), GATA-3 (c) were measured by reverse transcriptase (RT)–PCR and IL-17 expression was analyzed by flow cytometry (d). Data are representative of three independent experiments. (e, f) CD4+ T cells were activated and transduced with green fluorescent protein (GFP)-empty (MIG) vector or GFP-GATA3 following which cells were cultured under Th17 conditions and analyzed 3 days after transduction. IL-17 was analyzed by flow cytometry (e) and RORγt expression was measured by RT-PCR (f). GFP+ cells were gated, numbers indicate percentage of IL-17 positive cells. Data are representative of two independent experiments. (g) A 2-kb RORγt promoter construct was co-transfected with empty vector or a GATA-3 expression vector into EL4 cells after which the cells were stimulated overnight with PMA/Iono. RORγt promoter–reporter activity was measured and normalized by Renilla luciferase activity. Data are representative of three independent experiments. **P<0.01. (h) A 2-kb IL-17 promoter construct was co-transfected with empty vector or a GATA-3 expression vector into Jurkat cells after which the cells were stimulated overnight with PMA/Iono. IL-17 promoter–reporter activity was measured and normalized by Renilla luciferase activity. Data are representative of three independent experiments. (i) Naive CD4+ T cells purified from Id3+/− and WT littermate control mice were cultured under Th17 conditions or Th17 conditions plus anti-IL-4 (5 μg ml−1, 10 μg ml−1) for 3 days; IL-17 vs. Foxp3 expression was then analyzed by flow cytometry. Data are representative of two independent experiments. (j) Naive CD4+ T cells purified from individual Id3+/− and WT littermate control mice were cultured under Th17 conditions or Th17 conditions plus anti-IL-4 (5 μg ml−1, 10 μg ml−1) for 3 days; IL-17 was analyzed by flow cytometry after which the percentage of IL-17+ cells for each mouse was plotted. Each dot represents data from one mouse. Data are pooled from two independent experiments.
Finally, we determined the effect of partial deletion of Id3 on Th17 differentiation with studies in which we cultured CD4+ T cells from Id3+/− mice under Th17 conditions. We found that partial deletion of Id3, in contrast to complete deletion, was associated with a modest increase in IL-17 expression compared with WT T cells that was further increased by addition of anti-IL-4 (Figure 4i). To investigate this discrepancy further, we compared the capacity of T cells from Id3+/− mice and Id3−/− mice to produce IL-4 and express GATA-3, and found that after T cell receptor activation under Th0 or Th17 conditions, Id3+/− cells produce substantially less of these factors than Id3−/− cells (Supplementary Figure 3c). These findings thus indicate that in mice with decreased but not absent amounts of Id3 protein (Id3+/− mice), there is more E-protein available to drive Th17 differentiation but not enough GATA-3 to block such differentiation; in contrast, in mice with total absence of Id3 protein (Id3−/− mice), high levels of IL-4 and GATA3 are produced that block the positive effect of E-protein on Th17 differentiation. Thus, RORγt expression requires calibrated upregulation of Id3 to achieve an E-protein/Id3 protein ratio that balances the positive and negative effects of Id3.
In a separate series of studies we also determined the effect of Id2 deletion on RORγt and IL-17 mRNA expression despite the above-mentioned concern about the possible effect of Id2 deletion of Id3 levels. We found that cells from Id2−/− mice stimulated under Th17 conditions for 3 days also expressed decreased RORγt and IL-17 mRNA as well as increased GATA-3 mRNA, but these effects were less than those observed in cells from Id3−/− mice (Supplementary Figure 3d,e). Thus it is likely that Id2, under Th17 conditions, has a similar but lesser effect on Th17 differentiation as Id3.
Synchronous regulation of E-proteins and Id proteins by TGF-β and IL-6 signaling
We next determined whether TGF-β and IL-6, the two major cytokines driving Th17 cell differentiation, act through E-proteins. To address this question, purified CD4+ T cells from WT mice were activated with anti-CD3/anti-CD28 and cultured with different combinations of cytokines. Cells were harvested on days 1, 2, and 3, at which time E-protein (E2A or HEB), Id3, RORγt, and IL-17 expression were assessed by real-time reverse transcriptase–PCR (Figure 5a–f). We found that on days 1 and 2, there is no significant change of E-protein expression under all of the cytokine conditions. However, on day 3, cells cultured in the presence of TGF-β and IL-6 (i.e., Th17 conditions) exhibited a significant increase in both E2A and HEB expression as compared with cells cultured with no added cytokines (i.e., neutral conditions), or cells cultured with TGF-β or IL-6 alone (Figure 5a). Interestingly, Id3 was also significantly increased on day 3 under Th17 culture conditions but not under other conditions (Figure 5c). In additional studies focusing on RORγt and IL-17, we found that while small increases of these proteins were seen on days 1 and 2, large increases were seen only on day 3 accompanying changes in E-protein and Id3 expression (Figure 5d). Thus, E-proteins and Id3 are synchronously upregulated by cytokines that have been identified as the main inducers of the Th17 response and appear to be necessary for the induction of this response.
Kinetic studies of E-protein and Id protein expression under various cytokine conditions. Purified CD4+ T cells from wild-type (WT) mice were cultured with various combinations of cytokines involved in Th17 differentiation. Cells were harvested on d1, d2 and d3 respectively, after which expression levels of E2A (a), HEB (b), Id3 (c), RORγt (d), and interleukin (IL)-17 (e) were analyzed by reverse transcriptase–PCR. Data are representative of at least three independent experiments.
In parallel studies, we measured effect of IL1-β and IL-23 on HEB, Id3, RORγt, and IL-17 levels. We again found that IL-6 and TGF-β (in combination) gave rise to increased levels of these factors on the day 3 of culture. However, IL-1β and IL-23, alone or in combination (or both in combination with IL-6) were not associated with an increase in HEB and Id3 (Supplementary Figure 4a–d).
E-proteins are required for Th17-mediated autoimmune inflammatory disease
To determine whether E-protein is also essential for RORγt expression and Th17 differentiation in an in vivo setting, we first investigated the role of E-proteins in the generation of IL-17-producing T cells in the un-inflamed small intestine, a mucosal site previously shown to harbor such cells during commensal colonization.22 Accordingly, ER-Cre+ and WT mice were administered tamoxifen by gavage every other day over a period of 10 days and 4 weeks later the frequency of IL-17-producing cells in the small intestinal lamina propria was assessed by flow cytometry. We found that ER-Cre+ mice exhibited a much lower level of RORγt and IL-17 mRNA (Figure 6a) and lower frequency of IL-17 cells (Figure 6b) than WT mice.
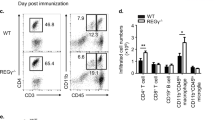
E-proteins regulate Th17 responses in vivo. (a, b) ER-Cre+ and wild-type (WT) mice were administered tamoxifen by gavage five times every other day. Four weeks after the final tamoxifen administration, lamina propria cells were purified from the small intestine; some of the cells obtained were directly analyzed by reverse transcriptase (RT)–PCR for RORγt or interleukin (IL)-17 (a), some of the cells were stimulated with PMA/Iono for 4 h with Golgi stop in the last 3 h and then analyzed for interleukin (IL)-17-expressing cells by flow cytometry gated on CD4+ cells (b). Data are representative of two independent experiments. (c, d) ER-Cre+ and WT were orally administered with tamoxifen every other day for five times, One week after the final dose, mice were orally inoculated with C. rodentium and were weighed at the indicated time points (d) and followed for survival. Data are representative of two independent experiments. (e) Colonic tissue obtained from WT- or knockout (KO)-infected mice at 14 days after initiation of infection were subjected to real-time RT-PCR analysis of RORγt and IL-17 expression. Data are representative of two independent experiments. (f) Experimental allergic encephalitis (EAE) disease course in WT and E2Af/+HEBf/fCre+ (KO) mice that had been treated with tamoxifen as above. Data are representative of two independent experiments; n=5 (WT), n=6 (KO). (g) EAE disease course in Rag2-deficient mice reconstituted with 20 × 106 purified CD4+ T cells from WT and E2Af/+HEBf/fCre+ (KO) mice that had been treated with tamoxifen as above; n=6 (WT), n=6 (KO). Data are representative of two independent experiments. (h) Cytokine production by cells isolated from the spinal cords of WT and E2Af/+HEBf/fCre+ (KO) mice on day 23 after EAE induction. The cells were stimulated for 4 h with PMA/Iono in the presence of Golgi stop and then analyzed for surface and intracellular cytokines by flow cytometry gated on CD4+ cells. Clinical score are shown in parentheses. Data are representative of two independent experiments. Tabulated results from two independent experiments: Total: all IL-17+ cells; interferon (IFN)-γ+:IL-17+IFN-γ+ cells; IFN-γ−:IL17+IFN-γ− cells. Error bars represent standard deviation; **P<0.02, *P<0.05. (i) Cytokine production by cells isolated from the spinal cord of Rag2-deficient mice reconstituted with tamoxfen-treated WT or KO purified CD4+ T cells on day 20 after EAE induction. The cells were stimulated for 4 h with PMA/Iono in the presence of Golgi stop and then analyzed for surface and intracellular cytokines by flow cytometry gated on CD4+ cells. Clinical scores are shown in parentheses. Data shown are representative of two independent experiments. Tabulated results are from two independent experiments. Total: all IL-17+ cells; IFN-γ+:IL-17/IFN-γ+ cells; IFN-γ−:IL17+IFN-γ− cells. Error bars represent s.d.; **P<0.01, *P<0.05.
In further in vivo studies, we determined the effect of E-proteins on the Th17 response of mice subjected to small intestinal infection with Citrobacter rodentium, an infection in which an intact IL-23–IL-17 axis has been shown to be essential for host protection.23 In these studies, ER-Cre+ and WT mice were administered tamoxifen by gavage as described above and 1 week later orally inoculated with C. rodentium. Whereas infected WT had a transient loss of weight followed by full recovery of weight by 10 days after inoculation, infected KO mice exhibited persistent loss of weight over the 16-day observation period (Figure 6c); furthermore, blinded studies of colons at day 14 after inoculation showed that C. rodentium-infected KO mice displayed greater mucosal inflammation, ulceration, and fibrotic reaction than WT mice (data not shown) and, in pooled experiments, whereas ∼20% of the Cre+ mice had died by day 14, none of the WT mice had died by this time point (Figure 6d). These findings correlated with the fact that colonic tissues from KO mice exhibited decreased levels of RORγt and IL-17 compared with tissues from WT mice. These in vivo data, taken together, strongly suggest that E-protein deficiency leads to decreased IL-17 responses during both commensal colonization and pathogenic bacterial infection of the intestine.
We next investigated the role of E-protein in experimental allergic encephalitis (EAE), a model of inflammation previously shown to be associated with a robust Th17 response.24 In an initial study, WT and E2Af/+HEBf/fCre+ (KO) mice were injected with tamoxifen six times as in Figure 2e. EAE was induced in WT and KO mice by injecting MOG35-55 in complete Freund’s adjuvant and pertussis toxin (PTX) at days 0 and 2. WT mice exhibited initial evidence of disease on day 12 and evidence of severe disease (score 3) on days 18–20. In contrast, only about 50% of the E2Af/+HEBf/fCre+ mice exhibited disease and this consisted of mild disease (maximum score 1) on days 16–20, after which they recovered; the other 50% of mice in this group did not develop any disease (Figure 6c).
In complementary studies, we employed an EAE model in which EAE is induced in RAG2-deficient mice (RAG2−/−) after the latter are reconstituted with purified CD4+ T cells from WT or E-protein conditional KO mice that have been treated with tamoxifen as described above. Transfer of 20 × 106 CD4+ T cells from WT mice to RAG2−/− mice, followed by induction of EAE 48 h later led to development of disease by 14 days, which peaked at a score of 3. In contrast, transfer of the same number of CD4+ T cells from E-protein conditional KO mice resulted in delayed disease: most of the mice with disease displayed a disease score of 1, and only a few mice displayed a score of 2 (Figure 6d).
Although E-protein conditional KO mice and Rag2−/− mice that received CD4+ T cells from E-protein conditional KO mice developed highly attenuated EAE, the number of infiltrating CD4+ T cells present in the spinal cords of these mice were similar to those in mice reconstituted with WT cells (data not shown). In both cases, interferon-γ-producing cells were present in a similar frequency in WT and E-protein conditional KO mouse spinal cords; however, the frequency of infiltrating T cells producing IL-17 was much lower in E-protein conditional KO mice (Figure 6e) compared with WT mice. Together, these results indicated that E-proteins not only regulate IL-17 expression in vitro, they also regulate IL-17 in vivo.
Discussion
Previous studies of the molecular mechanisms governing Th17 differentiation have focused mainly on the factors regulating IL-17 transcription, particularly as the latter relates to the transcriptional activity of RORγt, the Th17 lineage-defining factor known to be under the control of Th17-inducing cytokines such as TGF-β and IL-6.25, 26 In the present study, we address a perhaps more fundamental Th17 molecular mechanism, namely the mechanism by which RORγt itself is regulated. We show that E-proteins and Id3 are factors induced by TGF-β and IL-6 during Th17 differentiation that have distinct yet interactive roles in RORγt expression.
Our initial findings derived from luciferase reporter studies established that there are several E-protein binding sites in the RORγt promoter that when individually mutated, display greatly reduced promoter–reporter activity. Then, in more definitive studies utilizing ER-Cre conditional E2A and HEB double-floxed mice, we showed that a tamoxifen-induced 75–80% reduction in E2A/HEB mRNA levels led to comparable reductions in RORγt and IL-17 expression levels. In addition, we showed that CD4+ T cells from E2Af/fHEBf/fCD4Cre mice with E2A/HEB deletion due to constitutive expression of Cre in CD4+ T cells exhibited a similar level of RORγt/IL-17 deletion. Finally, we demonstrated that E-proteins induction has a major impact on the capacity to mount Th17 responses in vivo under both noninflammatory and inflammatory conditions. Taken together, these studies offer strong proof that RORγt is highly regulated by E-protein during in vitro Th17 differentiation. As such, they corroborate earlier preliminary studies showing that E-protein mRNA knockdown in vitro inhibits RORγt transcription and IL-17 expression.18
Id proteins are known to dimerize with E-proteins and thus block their ability to bind to DNA.11, 12 This suggested to us that the interaction between Id protein and E-proteins might be involved in RORγt transcription, and led us to conduct extensive studies of Th17 differentiation in Id3-deficient mice. Unexpectedly, we found that Id3−/− CD4+ T cells manifest decreased RORγt expression and IL-17 transcription rather than increased expression of these parameters. This initially paradoxical result was subsequently explained by the finding that Id3−/− cells cultured under Th17 conditions greatly upregulate their production of a factor that has a strong negative effect on RORγt transcription, GATA-3, and such upregulation, in turn, was explained by the finding that Id3−/− cells also increase their production of IL-4, a GATA-3 inducer previously shown to inhibit Th17 differentiation.21 These in vitro results were subsequently confirmed in vivo in a cell-transfer colitis model in which Id3−/− naive cells were found to induce less colitis and less IL-17 expression than WT naive cells.
It should be noted that in a previous study it was found that Id3−/− cells exhibit increased rather than decreased Th17 differentiation as reported in the present study.18 In extensive unpublished in vitro studies bearing on this issue, we found that cells with partial deletion of Id3, i.e., Id3+/− cells, do in fact exhibit increased Th17 differentiation relative to WT cells because Id3+/− cells do not produce IL-4 at a level sufficient to induce inhibitory GATA-3 under Th17 conditions; thus, in this case increased E-protein RORγt transcriptional activity due to decreased Id3 is unopposed by increased GATA-3-mediated inhibition of RORγt transcription. Whether or not these data can explain previous findings awaits additional study; we can say with certainty, however, that the conclusions relative to Id3 effects reported here are based on reproducible in vitro experiments that were subsequently corroborated by in vivo studies of Id3−/− mice.
Finally, the importance of both E-proteins and Id3 in the regulation of RORγt and IL-17 expression was highlighted by the fact that these proteins undergo upregulation by TGF-β and IL-6 in CD4+ T cells during Th17 differentiation, but hardly at all in cultures containing either of these cytokines alone. This assertion follows from the fact that E-proteins and Id3 are the only factors so far shown to be involved in Th17 differentiation that are responsive to the key cytokines inducing such differentiation, other than RORγt itself whose transcription is being regulated by these proteins. The fact that E-proteins and Id3 upregulation are synchronized also supports this assertion because, as we have seen, both factors are necessary to successfully execute the role of E-proteins in Th17 differentiation.
In conclusion, the upregulation of E-proteins and Id proteins in nascent Th17 cells by TGF-β and IL-6 suggests that these transcription factors act in tandem to serve as a molecular ‘switch’ that turns on Th17 differentiation. Given their upregulation by factors specifically involved in the induction of the Th17 response, E-proteins and Id proteins appear to have a specific relation to this response.
Methods
Mice and tamoxifen administration. C57BL/6 WT mice were from the Jackson Laboratory (Bar Harbor, ME). E2Af/fHEBf/fER-Cre mice and Id3−/− mice (all B6 background) were obtained from Dr Yuan Zhuang, Duke University; all mice were bred in an NIAID animal facility under specific pathogen-free conditions. To induce Cre expression in E2Af/fHEBf/fER-Cre mice, the mice were administered tamoxifen (Sigma, St Louis, MO, Cat#H7904) dissolved in corn oil per os five times every other day (3 mg/mice) or six times every 5 days. Cell culture or EAE studies were conducted one week after the final tamoxifen administration. All animal studies were performed under an animal use protocol approved by the Animal Use Committee of the NIAID.
Transient transfection and reporter assays. The RORγt promoter was obtained by PCR using mouse genomic DNA as a template. The PCR product was cloned into the pGL4.10 basic luciferase vector (Promega, Madison, WI) using the NheI and BglII entry sites. Promoter mutations were introduced using a Quickchange II site-directed mutagenesis kit (Stratagene, Grand Island, NY). PCR primers for these mutations are listed in Supplementary Table 1 online or for the E3 and E4 mutations as previously published.8 All constructs were verified by sequencing. EL4 cells were transfected using a Nucleofector II machine (Amaxa, Cologne, DEU), then stimulated and subjected to luciferase assay as previously described.1
RNA interference. CD4+ T cells were obtained using a naive CD4 T cell purification kit (Miltenyi, Cologne, DEU), and then transfected by nucleofection with Stealth E2A- or HEB-specific siRNA or control (scrambled) siRNA (Invitrogen, Grand Island, NY) (Supplementary Table 1) as previously described.1
In vitro Th17 cell differentiation and flow cytometric analysis. CD4+ T cells were purified by magnetic sorting using mouse anti-CD4 beads (Miltenyi) or by naive CD4 T cell purification kit (Miltenyi). Purified cells were cultured under Th17 condition as previously described.1 Some of the cells were cultured in the presence of 1 μM 4-OH-tamoxifen (4-OHT, Sigma cat #T5648). Flow cytometric analysis of intracellular cytokine was performed as previously described.1 The following antibodies were used for flow cytometry: anti-CD4-PE or Percp, anti-IL-17-PE or APC (BD Bioscience, San Jose, CA), anti-Foxp3-FITC or -APC, anti-GATA3-PE (eBioscience, San Diego, CA).
mRNA isolation and quantitative real-time PCR. mRNA isolation and quantitative real-time PCR were performed as previously described.1 List of primers and probes from Applied Biosystems (Grand Island, NY) is provided in Supplementary Table 1.
Chromatin immunoprecipitation assay. Chromatin was obtained by using a ChIP-IT Express Enzymatic kit (Active Motif cat# 53009). The chromatin was incubated with anti-E47 (Santa Cruz, Dallas, TE, cat# sc-763), anti-HEB (Santa Cruz cat# sc-357) or normal Rabbit IgG antibody as previously described.1 The DNA obtained was subjected to real-time PCR using primer pairs and probes listed in Supplementary Table 1.
Retroviral transduction. Empty vector and GATA-3 retroviral transduction were performed as previously described.1 The cells were cultured under Th17 condition after transduction and analysis as previously described.1
Transfer colitis. CD4+CD45RBhi naive T cells obtained from WT or Id3−/− mice were purified by FACS sorting following which 4 × 105 purified cells were transferred into Rag2−/− mice. The mice were then monitored for weight loss, and tissue was collected for analysis at the time indicated.
C. rodentium -induced colitis. ER-Cre+ mice and ER-Cre− mice were administered tamoxifen every other day for 10 days (five times) as described above. C. rodentium (ATCC51459) was prepared by incubation with shaking at 37 °C for 6 h in lysogeny broth. The relative concentration of bacteria was assessed by measuring absorbance at OD600. Mice were inoculated with 1 × 109 c.f.u. bacteria by oral administration 1 week after the final tamoxifen treatment. Tissue was collected post inoculation for histology and/or cytokine analysis at the times indicated.
Isolation of lamina propria lymphocytes. Purified small intestinal or large intestinal lymphocytes cells were obtained as previously described.27
EAE induction. EAE was induced by subcutaneous immunization on day 0 with MOG35-55 peptide (0.2 mg per mouse, Biosynthesis) emulsified in CFA (CFA supplemented with 8 mg ml−1 Mycobacterium tuberculosis, Gibco, Grand Island, NY) and intraperitoneal injection on days 0 and 2 with pertussis toxin (200 ng per mouse, Calbiochem, Darmstadt, DEU). Scoring system criteria: 0=no disease, 1=flaccid tail, 2=weak/partially paralyzed hind legs, 3=completely paralyzed hind legs, and 4=completely paralyzed hind legs and partially paralyzed front legs.
CD4+ T cell transfers. CD4+ T cells were purified by anti-CD4 magnetic microbeads (purity >95%) from WT Cre+ and E2A/HEB/Cre mice that had been treated with tamoxifen as described above, and were transferred into RAG2−/− mice (20 × 106 cells per mouse) intravenously. Forty-eight hours after adoptive transfer, the mice were subjected to EAE induction.
Isolation of lymphocytes from spinal cords. Spinal cords were extracted from dissected spinal columns by flushing the latter with phosphate-buffered saline using a 19G needle. The columns were then cut into small pieces and were subjected to lymphocyte extraction using a Neutral Tissue Dissociation Kit (Miltenyi, Cat #130-092-628). The cell suspension obtained was then washed with phosphate-buffered saline, resuspended in 10 ml of ice cold Hank's balanced salt solution containing 0.9 M sucrose and centrifuged at 850 g for 10 min; finally, the pelleted cells were washed with media, restimulated with PMA/Iono, and analyzed for intracellular cytokines by flow cytometry as above.
References
Zhang, F, Meng, G & Strober, W. Interactions among the transcription factors Runx1, RORgammat and Foxp3 regulate the differentiation of interleukin 17-producing T cells. Nat Immunol 9, 1297–1306 (2008).
Wei, L, Laurence, A, Elias, K.M. & O'Shea, J.J. IL-21 is produced by Th17 cells and drives IL-17 production in a STAT3-dependent manner. J Biol Chem 48, 34605–34610 (2007).
Zhou, L. et al. IL-6 programs T(H)-17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nat Immunol 8, 967–974 (2007).
Schraml, B.U. et al. The AP-1 transcription factor Batf controls T(H)17 differentiation. Nature 460, 405–409 (2009).
Rius, J. et al. NF-kappaB links innate immunity to the hypoxic response through transcriptional regulation of HIF-1alpha. Nature 453, 807–811 (2008).
Dang, E.V. et al. Control of T(H)17/T(reg) balance by hypoxia-inducible factor 1. Cell 146, 772–784 (2011).
Shi, L.Z. et al. HIF1alpha-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J Exp Med 208, 1367–1376 (2011).
Xi, H, Schwartz, R, Engel, I, Murre, C & Kersh, G.J. Interplay between RORgammat, Egr3, and E proteins controls proliferation in response to pre-TCR signals. Immunity 24, 813–826 (2006).
Jones, M.E. & Zhuang, Y. Acquisition of a functional T cell receptor during T lymphocyte development is enforced by HEB and E2A transcription factors. Immunity 27, 860–870 (2007).
Murre, C. Helix-loop-helix proteins and lymphocyte development. Nat Immunol 6, 1079–1086 (2005).
Kee., B.L. E and ID proteins branch out. Nat Rev Immunol 9, 175–184 (2009).
Benezra, R, Davis, R.L., Lockshon, D, Turner, D.L. & Weintraub, H. The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell 61, 49–59 (1990).
Miyazaki, M, Rivera, R.R., Miyazaki, K, Lin, Y.C., Agata, Y & Murre, C. The opposing roles of the transcription factor E2A and its antagonist Id3 that orchestrate and enforce the naive fate of T cells. Nat Immunol 12, 992–1001 (2011).
Braunstein, M & Anderson, M.K. HEB-deficient T-cell precursors loss T-cell potential and adopt and alternate pathway of differentiation. Mol Cell Biol 31, 971–982 (2011).
Jones-Mason, M.E., Zhao, X, Kappes, D, Lasorella, A, Iavorone, A & Zhuang, Y. E protein transcription factors are required for the development of CD4(+) lineage T cells. Immunity 36, 348–361 (2012).
Boos, M.D., Yokota, Y, Eberl, G & Kees, B.L. Mature natural killer cell and lymphoid tissue-inducing cell development requires Id2-mediated suppression of E protein activity. J Exp Med 204, 1119–1130 (2007).
Yang, C.Y. et al. The transcriptional regulators Id2 and Id3 control the formation of distinct memory CD8+ T cell subsets. Nat Immunol 12, 1221–1229 (2011).
Maruyama, T. et al. Control of the differentiation of regulatory T cells and T(H)17 cells by the DNA-binding inhibitor Id3. Nat Immunol 12, 86–95 (2011).
Yokota, Y. et al. Development of peripheral lymphoid organs and natural killer cells depends on the helix-loop-helix inhibitor Id2. Nature 397, 702–706 (1999).
Rivera, R.R., Johns, C.P., Quan, J, Johnson, R.S. & Murre, C. Thymocyte selection is regulated by the helix-loop-helix inhibitor protein, Id3. Immunity 12, 17–26 (2000).
Harrington, L.E. et al. Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat Immunol 6, 1123–1132 (2005).
Ivanov, I.I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Symonds, E.L., Riedel, C.U., O'Mahoney, D.O., Lapthorne, S, O'Mahoney, L & Shanahan, F. Involvement of T helper type 17 and regulatory T cell activity in Citrobacter rodentium invasion and inflammatory damage. Clin Exp Immunol 157, 148–154 (2009).
Zepp, J, Wu, L & Li, X. IL-17 receptor signaling and T helper 17-mediated autoimmune demyelinating disease. Trends Immunol 32, 232–239 (2011).
Mangan, P.R. et al. Transforming growth factor-beta induces development of the T(H)17 lineage. Nature 441, 231–234 (2006).
Veldhoen, M, Hocking, R.J., Atkins, C.J., Locksley, R.M. & Stockinger, B. TGFbeta in the context of an inflammatory cytokine milieu supports de novo differentiation of IL-17-producing T cells. Immunity 19, 179–189 (2006).
Yang, Z. et al. NOD2 transgenic mice exhibit enhanced MDP-mediated down-regulation of TLR2 responses and resistance to colitis induction. Gastroenterology 133, 1510–1521 (2007).
Acknowledgements
We thank Dr Yuan Zhuang (Duke University) for E2Af/fHEBf/fER-Cre mice and Id3−/− mice, Dr William Paul (NIAID/NIH) for a GATA3 construct, and Dr Kersh Gilbert (Emory University) for E47, HEB, and Id3 constructs. We also thank Dr Qian Chen (NIH/NIAID) for help with the EAE model.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
Author contribution
F.P. planned and performed studies and wrote/edited the manuscript; I.F. performed studies and managed animal breeding; Z.Y. performed studies; W.S. planned studies and edited the manuscript.
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Rights and permissions
About this article
Cite this article
Zhang, F., Fuss, I., Yang, Z. et al. Transcription of RORγt in developing Th17 cells is regulated by E-proteins. Mucosal Immunol 7, 521–532 (2014). https://doi.org/10.1038/mi.2013.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2013.69
This article is cited by
-
NFAT primes the human RORC locus for RORγt expression in CD4+ T cells
Nature Communications (2019)
-
Oxysterol levels and metabolism in the course of neuroinflammation: insights from in vitro and in vivo models
Journal of Neuroinflammation (2018)
-
HEB is required for the specification of fetal IL-17-producing γδ T cells
Nature Communications (2017)
-
Jieduquyuziyin prescription suppresses IL-17 production and Th17 activity in MRL/lpr mice by inhibiting expression of Ca2+/calmodulin-dependent protein kinase-4
Journal of Natural Medicines (2015)