Abstract
Multiple questions relating to contributions of cultural and demographical factors in the process of human geographical dispersal remain largely unanswered. India, a land of early human settlement and the resulting diversity is a good place to look for some of the answers. In this study, we explored the genetic structure of India using a diverse panel of 78 males genotyped using the GenoChip. Their genome-wide single-nucleotide polymorphism (SNP) diversity was examined in the context of various covariates that influence Indian gene pool. Admixture analysis of genome-wide SNP data showed high proportion of the Southwest Asian component in all of the Indian samples. Hierarchical clustering based on admixture proportions revealed seven distinct clusters correlating to geographical and linguistic affiliations. Convex hull overlay of Y-chromosomal haplogroups on the genome-wide SNP principal component analysis brought out distinct non-overlapping polygons of F*-M89, H*-M69, L1-M27, O2a-M95 and O3a3c1-M117, suggesting a male-mediated migration and expansion of the Indian gene pool. Lack of similar correlation with mitochondrial DNA clades indicated a shared genetic ancestry of females. We suggest that ancient male-mediated migratory events and settlement in various regional niches led to the present day scenario and peopling of India.
Similar content being viewed by others
Introduction
India has been a hub of early human expansion since the first wave of out of Africa migration through the southern route.1, 2, 3 High phenotypic diversity and varied cultures of people of India along with recent genomic studies led to different views on how the peopling of India occurred. The population of India has been declared a Pleistocene gene pool,4, 5, 6, 7 living in a diverse landscape and practicing varying degrees of inbreeding and endogamy. In addition, various geographical niches and languages make their genetic history more complex. Our study on the people of Tamil Nadu, the southernmost state of India, highlighted the importance of considering the population structure along with cultural and subsistence elements in unraveling the peopling of India.8
Studies of genetic history of Indian population based on uni-parental markers have produced some conflicting results. For example, non-recombinant Y-chromosomal (NRY) markers analysis suggested the presence of many autochthonous and non-autochthonous lineages in the subcontinent. NRY lineages such as F*-M89, H-M69 and L-M20 may have had an autochthonous origin, whereas others such as J-M304, O2a-M95 and O3a3-M117 have an external origin, and the origin of certain other haplogroups such as R1a1-M17 are disputed.6, 9, 10 An external origin of NRY haplogroup R1a1-M17, the speculated marker of Indo-Aryan, was proposed based on the sharing of this haplogroup between Indo-European speakers of India and Eastern Europeans.2, 11 In contrast, a study by Sharma et al.,10 supported the Indian origin of the NRY haplogroup R1a1-M17, and, therefore, of the Indo-European speakers.10, 12 Interestingly, the Dravidian speakers are characterized by NRY haplogroups F*-M89 and L1-M27,6, 8 suggesting an in situ evolution, in contrast to the Proto Elamite theory of origin in the Fertile Crescent.13 The origin of NRY haplogroup O2a-M95, mostly represented in Austro-Asiatic speakers of Eastern India, is also controversial, and both indigenous and Southeast Asian origins have been proposed.14, 15 However, mitochondrial DNA (mtDNA) markers suggest a shared maternal gene pool of India,16, 17 with 60% of Indian mtDNA being composed of mtDNA haplogroup M and its derivatives,18 with a majority of the rest belonging to mtDNA haplogroup R and its derivatives.19, 20 Thus, studies of uni-parental markers point to a uniformity of a deep-rooted female gene pool of India. Interestingly, the male gene pool of India has had multiple origins (both inside and outside the subcontinent) and exhibit a strong correlation with the language family.
Recently, many genome-wide studies using microarray chips containing ancestry informative markers revolutionized human population genetics. A seminal study based on 132 Indian samples, encompassing 560 123 single-nucleotide polymorphisms (SNPs) suggested the existence of ancient North Indian and ancient South Indian gene pools, therefore making the present day Indo-European and Dravidian speakers of India a result of recent admixture between the two ancient gene pools.21, 22 Further, high similarity between North and South Indian populations have been found in whole-exome sequencing as well.23 These studies reiterate that the genetic variation of different linguistic groups may be a result of different migratory events.
The Genographic Consortium has developed the Geno 2.0 chip, a dedicated genotyping array containing only ancestry informative markers.24 This tool is useful for detection of accurate admixture proportions for world-wide populations. The ancestral information predicted by this tool has been validated using Genographic and HapMap reference samples.25 In the present study, 66 samples drawn from the entire geographical expanse of India, used previously in the GenoChip validation study, and 12 additional HapMap Gujarati samples were further analyzed. The study interrogated (i) how the Indian samples are stratified based on the ancestry informative markers studied in the GenoChip; (ii) does the genome-wide SNPs-based clustering correlate to covariates such as geography, language or other cultural traits?; and (iii) do the genome-wide signals correlate to variation of their uni-parental markers?
Materials and methods
Sampling
The 66 Indian samples used in this study were collected as part of the Genographic Project. Mouthwash was collected from volunteers and DNA was isolated using standard salting out method with ethanol precipitation.26 A signed informed consent was obtained before the sample collection. The study protocol was approved by the Institutional Ethical Committee of Madurai Kamaraj University, India and University of Pennsylvania, USA. To increase the representation of the West Indian samples, we added a random subset of 12 Gujarati samples from the HapMap data set to get a snapshot of the genome diversity of from West India.25
Genotyping
Genome-wide SNP genotyping was performed using the GenoChip array in an Illumina HD iSelect genotyping bead array.24 These results have formed a part of the validation study.25 We analyzed the data (Supplementary File 1) in the context of India and genotyped the Indian subset of samples for 23 Y-SNPs using custom-made Taqman assays (Applied Biosystems, Foster City, CA, USA) in 7900HT Fast Realtime System and a core set of 22 mtDNA SNPs using a custom-made SNaPSHOT assay.8, 27
Biogeography
We followed the protocol described in Elhaik et al.,25 the approach is briefly described below. We used the world-wide reference data set data on 127 000 autosomal SNPs of 615 individuals from 54 world-wide populations collected as part of the Genographic Project and genotyped on the GenoChip array.24 High data quality was achieved by applying two criteria to the SNP data: (i) low missingness rate (<5%), calculated as the average number of null genotypes over all samples in a population and (ii) individuals that exhibit exceptional admixture proportions (>Δ5% in any admixture coefficient) compared to the mean population proportions were considered outliers and omitted. We therefore omitted 2423 SNPs and five samples from the analysis.
Admixture framework
First, ADMIXTURE was used in an unsupervised manner to obtain nine allele frequencies for each SNP, corresponding to the allele frequencies of nine putative ancestral populations. We then used the allele frequencies to generate the genotypes of the putative ancestral populations, simulating 15 individuals of each putative ancestry. Second, we used these simulated samples in a supervised manner for all further analyses so that ADMIXTURE associates each SNP of each test sample with one of the putative ancestral populations based on their simulated genotypes. In this manner, the admixture inference is sample independent and depends solely on the putative ancestral populations, and not on the other tested samples as in alternative approaches.28 The relationship between admixture and geography was calculated using nine admixture coefficients and geographical position using the reference samples, as described in Elhaik et al.,.25 The bio-origin of a test sample of unknown origin was obtained using the GPS algorithm (Supplementary File 2).25, 29 f4 test was performed to estimate the admixture proportions in Indian populations in different combinations of clustering following approach described in Reich et al.21
Principal component analysis
Principal component analysis (PCA) of the SNP genotype data of the samples was performed using ‘Adegenet’ package implemented in R v3.0.1.30 Polygon overlay of ‘Language family’, ‘Y-haplogroup’ and ‘mtDNA haplogroup’ over the PCA plots were performed using the convex hull method implemented using grDevices package in R v3.0.1. Hierarchical clustering was conducted using the built-in function in R, hclust. In order to investigate hierarchical clustering of the sampled populations, we analyzed our data in TreeMix software.31 We used linkage disequilibrium (LD) pruned data set, and we set Kenyan population as a root of the tree. Mean Fst values32 were calculated in vcftools33 on data set pruned for linkage disequilibrium.
Results
The design of the GenoChip has been earlier validated on 615 reference samples from 54 world-wide populations collected as part of the Genographic Project, including that used in the present study, and tested in admixture analyses.24, 25 The results have revealed the existence of a clear substructure of the world populations essentially determined by geography. This is in agreement with the accumulated knowledge on global population history.34, 35, 36 In the present paper, we further carried out extensive analysis on the Indian data set in the context of other uni-parental data generated (Supplementary Table 1 and Supplementary File 1).
Three major components of Indian gene pool
Unsupervised admixture analysis37 identified three major components making the Indian gene pool, each contributing 30–50%, namely, Southwest Asian (pitch red), Southeast Asian (blue) and Northeast Asian components (brown) (Supplementary Figure 1).25 Here, we clustered the Indian samples into six groups: five geographical and one cultural (pan-Indian Brahmins). The Brahmins were grouped separately because of their unique and common cultural traits and history.38 The pan-Indian Brahmins and Western Indian samples (12 Gujarati [HapMap], 1 Parsee and 1 Katkari) showed a modest (~10% combined) Northern European and Mediterranean component that were not found in any other clusters (Supplementary Figure 1), thereby suggesting a limited shared genetic ancestry with these populations. The Southwest Asian component was ubiquitously present in all the Indian samples, whereas the Northeast Asian component was predominant among Northeast Indians.
Clustering analysis
To further investigate the genetic affinities of the studied Indian samples, we performed hierarchal clustering based on their admixture proportions (Figure 1 and Table 1). The resulting tree showed seven distinct clusters, each containing a major proportion of samples with common linguistic or geographical affiliation or both. Cluster 1 was made mostly of Tibeto-Burmese speakers from Northeast India; Cluster 2: Austro-Asiatic speakers from East India; Cluster 3: Tibeto-Burmese speakers from Northeast and North India (adjoining Himalayas); Cluster 4: Indo-European speaking Brahmins from various regions of India; Cluster 5: non-Brahmin Indo-European speakers dispersed across India; Cluster 6: South Indian Dravidian speakers; and Cluster 7: a single West Indian population, Parsee. This clustering analysis revealed a combined influence of geography and language in shaping the gene pool of India. PCA based on the admixture proportions also revealed similar distinct population clusters, thus adding strength to our proposition (Supplementary Figure 2).
Admixture proportion based hierarchical clustering of Indian samples. The Indian populations clustered into six groups, each reflecting their overall geographical or linguistic affiliations. The Brahmins across India, irrespective of their present domicile, clustered as a separate entity reflecting their unique shared ancestry. Cluster 1—Tibeto-Burmese speakers from Northeast India; Cluster 2—Austro-Asiatic speakers from East India; Cluster 3—Tibeto-Burmese speakers from Northeast and North India; Cluster 4—Indo-European speaking Brahmins from various regions of India; Cluster 5—non-Brahmin Indo-European speakers dispersed across India; Cluster 6—South Indian Dravidian speakers; and Cluster 7—Parsee population of Western India.
To further investigate the individual effect of various covariates of population classification, we performed the χ2-test between the population classification and the hierarchal cluster membership (Table 2). The most significant association (P-value of 2.95 × 10−33) was obtained for the classification based on regional clusters plus the Brahmin population. It is to be noted that we considered Brahmins as a unique entity irrespective of their present day geographical domicile owing to their historically common cultural traits. States of India (mostly geo-linguistic entities) and linguistic classification also showed significant P-values (7.0 × 10−24 and 1.57 × 10−17).
The proportion of various global components in the seven clusters, as with admixture analysis (Supplementary Figure 1), revealed that the majority of Indian samples contained three global components: Northeast Asian, Southwest Asian and Southeast Asian, ranging from 57.3 to 64.3%. Clusters 1 and 3 contained 64.3 and 41.9% of the Northeast Asian component. All seven clusters, except for the Cluster 1, showed a high proportion of the Southwest Asian component. This component was observed at high frequencies only in the Indian populations with >35% of Indian samples with a frequency of 65% (Supplementary Figure 3). Therefore, this effect can be attributed to the Indian origin of this component.
The South Indian Dravidian speakers (Cluster 6) had the maximum proportion of the Southwest Asian component (58%), suggesting a probable in situ origin or expansion of their gene pool. The accuracy of this analysis was supported by the distinct genetic makeup of Cluster 7 (Parsee samples), with 39.9% of Mediterranean component. Parsees are believed to have migrated from the Greater Iran region to India during the eighth to tenth centuries.39 Parsee is genetically closer to the Iranian population than any other Indian populations.
We also confirmed the results of the clustering analysis by using the Treemix algorithm (Supplementary Figure 4). All Indo-European speakers clustered together in one branch, with the exception of Indo-European speakers from North India. Indian Indo-European speaking groups clustered at the base of the branch, whereas European populations have a terminal position at the branch. Dravidian speakers separated according to geographical location while the southern Dravidian populations were placed in proximity of the Indo-European speakers’ branch; the Dravidian speakers from east/central India were located at the extremity of the tree, near the Austro-Asiatic speakers. Indian Tibeto-Burmese speakers formed an individual branch together with China, Japan and Vietnam.
In order to additionally validate the conclusions based on admixture proportions in the Indian samples, we calculated f4 statistics for two trees with two ancestral populations and one Indian population: (Yoruba, ancestral 1; ancestral 2, tested Indian) and (Yoruba, ancestral 2; ancestral 1, tested Indian). Ratio of statistics results gives the estimate of which tree better reflects the data. In the next step, we carried out linear regression and extrapolated line to obtain x and y intercepts. These intercepts represent respectively 0 and 100% of ancestry of ancestral population 1. For every data point, we found closest point on the regression line and we interpolated ancestry proportions from this point. Results of regression ancestry estimate corroborate clinal pattern observed in results of Admixture and TreeMix. South and West Indian populations are more close to Indo-European speaking populations (Italian or Brahmin), whereas North and North East Indian populations exhibit greater affinity to North-East Asian populations (Chinese) (Supplementary Figure 5).
Diversity of uni-parental markers in India
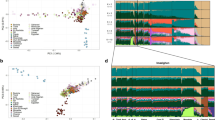
Having found the relationship between geography and spoken language with genomic variation based on admixture components, we investigated whether the observed variation could be attributed to the diversity of uni-parental markers observed in India, considering the regional differences in NRY composition in India.6 In the global PCA based on genotype data of GenoChip SNPs, Indian samples were placed in the midst of Asians and Europeans, whereas African samples clustered away from this main cluster (Figure 2a). The same was demonstrated using the smartPCA algorithm (Supplementary Figure 6). smartPCA is a Bayesian extension of a traditional PCA approach allowing incorporation of external knowledge. Inclusion of the ‘reference populations’ was necessary for the separation of Indian samples along the x, and particularly the y, coordinates in the PCA. Plot of the PCA3 vs PCA1 showed: (i) separation between Indian samples and the reference groups (American Indian, Asian, African and European); (ii) closer affinity of Indian samples to European and Asian samples than to African and American Indian ones; and, (iii) existence of clusters within Indian samples. The PCA using the whole-genome SNPs of Indian samples alone without the ‘reference populations’ revealed distinct geographical clusters of the samples (Figure 2b). East, South and Northeast Indians formed distinct clusters while the Brahmin and West Indian samples clustered together. The North Indian samples did not form a distinct cluster.
Principal component analysis (PCA) of Global and Indian populations based on the GenoChip single-nucleotide polymorphism (SNPs). Legend: (a) PCA of global samples, and (b) PCA of Indian samples. Each point represents samples and the symbols represent geographical origin of samples. Cross—Brahmin; square—North India; diamond—South India; star—East India; circle—Northeast India; circle cross—West India; bullet (small grey circle)—European; grey-filled diamond—African-American; grey-filled solid circle—African; grey colour star '*'—South America; grey colour '+'—East Asian.
Polygons representing linguistic family, NRY haplogroups and mtDNA haplogroups of Indian samples overlaid on the PCA clusters revealed a varied picture. The linguistic polygon overlay on the global PCA plot (Supplementary Figure 7a) showed a clear distinction between Dravidian, Austro-Asiatic and Tibeto-Burmese speakers similar to previous observations.21 The Dravidian and Indo-European speakers showed a larger overlap, whereas the Dravidian and Austro-Asiatic speakers showed a minimal overlap. The picture became clearer when PCA was restricted to Indian samples (Supplementary Figure 7b).
The NRY haplogroup polygon overlay on the global and Indian PCA brought out five distinct, non-overlapping polygons relevant to five different NRY haplogroups, namely: F*-M89, H*-M69, L1-M27, O2a-M95 and O3a3c-M117 (Supplementary Figure 8). The correlation between the NRY haplogroup variation and the whole-genome SNP variation was confirmed by a significant Mantel test P-value of 0.0003. On the other hand, the less common mtDNA haplogroups B, R8 and M36 were the only ones showing non-overlapping polygons (Supplementary Figure 9). The absence of non-overlapping mtDNA haplogroup polygons in genome-wide SNP PCAs (Mantel P–value=0.1275) supports the observation that almost 80% of Indian mtDNA lineages are descendants of only two mtDNA haplogroups, M and R, and that the female genepool of India is highly fluid, which may be attributed to patriarchal social practices.7, 16, 18, 19, 40
Discussion
The present study is a snapshot of Indian genetic variation based on ancestry informative markers spanning the entire genome. The results highlighted a correlation of genetic variation with geographical and linguistic affiliation of the samples. In addition, a correlation to Y-chromosomal variation suggested a male-dominated genetic structure.
Genetic structure of India
Previous studies have proposed two major genetic elements that have shaped the Indian gene pool. One, termed Ancestral North Indian, is closer to European, Middle Eastern and Central Asian, and the other is a distinct Ancient South Indian.21, 22 The present study described three major components in the Indian gene pool, namely, Southwest Asian, Southeast Asian and Northeast Asian. The Southwest Asian component was present in highest proportions in the Indian samples only (Supplementary Figure 3), thus postulating it to be a native Indian component. On the other hand, the Southeast Asian component was present in moderate levels in Chinese and other East Asian populations.
A unique identity of Brahmin populations surpassing geographic differences was brought out as evidenced from the unique cluster in the hierarchal clustering analysis based on admixture proportions (Figure 1). The Brahmin samples from all over India contained 11.4 and 10.6% of Northern Eurasian and Mediterranean components, thereby suggesting a shared ancestry with the Europeans (Table 1; Cluster 4). This was also reflected in the genetic relatedness based on Fst values between Brahmins and Europeans (Supplementary Table 2). Note that our Fst values agree with those previously reported.21 Earlier studies describing Ancestral North Indian genetic variance and NRY haplogroup R1a1-M17 suggested similar shared ancestries with Europeans and Mediterraneans.10, 21
We compared consistency of Brahmin and Western Indian samples using the GPS algorithm.25 We conducted the leave-one-out analysis, removing one sample at a time and re-computed mean admixture vector for the remaining individuals in the population, and predicted the origin of the removed sample using GPS. In this analysis, all 13 Brahmin samples were mapped to Brahmins; however, among the 12 Gujarati samples, only 25% were mapped to Gujarat. This indicates that the Western India samples studied were not homogeneous, or that the Gujarati samples were admixed while the pan Indian Brahmins might have had a common genetic ancestry. Other Indian populations were too small for the leave-one-out analysis.
The Northeast Asian component was predominant among the Northeast Indian Tibeto-Burmese speakers who share genetic elements with East Asians.9 The presence of varying degrees of Northeast Asian component (Supplementary Figure 1, Brown) in Northern Indian sample studies was attributed to the inclusion of Tibeto-Burmese speakers of the Himalayan ranges in this group (Changspa, Rajput Kinnura), whose genetic history supports a migration from the Northeast Asian region.40, 41
Patterns observed in result of TreeMix (Supplementary Figure 4) reflect documented migration events and gene flow. The separation of Dravidian into two (South and East) is expected. The Dravidian speakers of East/Central India speak a subgroup of Dravidian language called Central Dravidian languages. These people live in close proximity with Austro-Asiatic speakers of East India (many within the same village). Gene flow between these two groups (central Dravidian and Austro Asiatic) is documented.15, 42 The Indo-European speakers of North India seem to have diverse ancestral elements. Six out of the 11 Indo-European-North samples are living in close proximity with Tibeto-Burmese speakers of the Himalayan ranges (Tibeto Burmese-North). Hence, there is a small genetic affinity with them that pulls the Indo-European North away from other Indo-European speakers.
Y-chromosomal signatures of Indian gene pool
Studies based on uni-parental marker have shown diverse Y-chromosomal haplogroups making up the Indian gene pool. Many of these Y-chromosomal markers show a strong correlation to the linguistic affiliation of the population.6, 8 The genome-wide variation of the Indian samples in the present study correlated with the linguistic affiliation of the sample (Supplementary Figure 7). However, Dravidian speakers showed genetic relatedness with Austro-Asiatic and Indo-European speakers. This could be a result of a shared Southeast Asian genetic component (Table 1). This relatedness has been earlier shown with Y-markers as well. The central Dravidian speakers (a subfamily of Dravidian languages), living amidst the Austro-Asiatic speakers in East India, share uni-parental markers (Y-haplogroups H1a-M52 and O2a-M95).15, 42 Similarly, Indo-European and Dravidian speakers show genetic relatedness described by sharing of NRY markers such as R1a1-M17 and H1a-M52.8 This scenario supports the historical evidences of Indo-European speakers’ entry into South India.38, 43
The disparate Y-chromosomal polygons overlaid on genome-wide PCA highlighted the effect of male-dominated genetic elements shaping the Indian gene pool (Supplementary Figure 7). Interestingly, these NRY markers have earlier been correlated to various languages. For example, the NRY haplogroup F*-M89 and L1-M27 are present in higher proportions among Dravidian speakers, and their origin was proposed to be autochthonous.6, 8 The NRY haplogroup O2a-M95 and O3a3c-M117 are characteristics of Austro-Asiatic and Tibeto-Burmese speakers of India, respectively.9, 14, 16, 44 The unique genome-wide signatures of individuals with deep-rooted NRY haplogroups F*-M89 and H*-M69, dating back to mid Pleistocene, suggest an ancient settlement in the subcontinent.6, 8, 45 On the other hand, lack of correlation of mtDNA haplogroups may suggest the fluidity of female gene pools when in a patriarchal and patrilocal society, such as that of India.
Limitations of the study
We note that the present study involved a limited sample size, but the disparate genetic clustering and information obtained provides a snapshot of the large genetic diversity present in India. This pilot study will aid formulating hypotheses to be tested in larger sample-sized studies from this region in the future.
In conclusion, the present observations suggest the following four likely scenarios: (i) the Indian gene pool is primarily composed of native Indian components; (ii) the genome-wide SNP variation correlates to both geographical and linguistic variation among the study populations; (iii) the correlation of non-overlapping deep-rooted NRY haplogroup polygons to genome-wide SNPs suggests that male-mediated migratory events primarily shaped the Indian gene pool; and (iv) the absence of non-overlapping mtDNA haplogroup polygons over genome-wide SNP variation suggests a shared female genetic ancestry of India surpassing the geographical and linguistic barriers. We suggest that ancient male-mediated migratory events and settlement in various regional niches led to the present day scenario and peopling of India.
References
Cavalli-Sforza, L. L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes, (Princeton University Press, Princeton, NJ, USA, 1994).
Wells, R. S., Yuldasheva, N., Ruzibakiev, R., Underhill, P. A., Evseeva, I., Blue-Smith, J. et al. The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc. Natl Acad. Sci. USA 98, 10244–10249 (2001).
Armitage, S. J., Jasim, S. A., Marks, A. E., Parker, A. G., Usik, V. I. & Uerpmann, H. P. The southern route "out of Africa": evidence for an early expansion of modern humans into Arabia. Science 331, 453–456 (2011).
Majumder, P. P. Ethnic populations of India as seen from an evolutionary perspective. J Biosci. 26, 533–545 (2001).
Majumder, P. P. The human genetic history of South Asia. Curr. Biol. 20, R184–R187 (2010).
Sengupta, S., Zhivotovsky, L. A., King, R., Mehdi, S. Q., Edmonds, C. A., Chow, C. E. et al. Polarity and temporality of high-resolution y-chromosome distributions in India identify both indigenous and exogenous expansions and reveal minor genetic influence of Central Asian pastoralists. Am. J. Hum. Genet. 78, 202–221 (2006).
Sun, C., Kong, Q.-P., Palanichamy, M. G., Agrawal, S., Bandelt, H.-J., Yao, Y.-G. et al. The dazzling array of basal branches in the mtDNA macrohaplogroup M from India as inferred from complete genomes. Mol. Biol. Evol. 23, 683–690 (2006).
ArunKumar, G., Soria-Hernanz, D. F., Kavitha, V. J., Arun, V. S., Syama, A., Ashokan, K. S. et al. Population differentiation of Southern Indian male lineages correlates with agricultural expansions predating the caste system. PLoS ONE 7, e50269 (2012).
Trivedi, R., Sahoo, S., Singh, A., Bindu, G. H., Banerjee, J., Tandon, M. et al. Genetic imprints of Pleistocene origin of Indian populations: a comprehensive phylogeographic sketch of Indian Y-chromosomes. Int. J. Hum. Genet. 8, 97–118 (2008).
Sharma, S., Rai, E., Sharma, P., Jena, M., Singh, S., Darvishi, K. et al. The Indian origin of paternal haplogroup R1a1(*) substantiates the autochthonous origin of Brahmins and the caste system. J. Hum. Genet. 54, 47–55 (2009).
Underhill, P. A., Myres, N. M., Rootsi, S., Metspalu, M., Zhivotovsky, L. A., King, R. J. et al. Separating the post-Glacial coancestry of European and Asian Y chromosomes within haplogroup R1a. Eur. J. Hum. Genet. 18, 479–484 (2010).
Sahoo, S., Singh, A., Himabindu, G., Banerjee, J., Sitalaximi, T., Gaikwad, S. et al. A prehistory of Indian Y chromosomes: evaluating demic diffusion scenarios. Proc. Natl Acad. Sci. USA 103, 843–848 (2006).
McAlpin, D. W. Proto-Elamo-Dravidian: the evidence and its implications. Trans. Am. Phil. Soc. 71, 3–155 (1981).
Kumar, V., Reddy, A. N., Babu, J. P., Rao, T. N., Langstieh, B. T., Thangaraj, K. et al. Y-chromosome evidence suggests a common paternal heritage of Austro-Asiatic populations. BMC Evol. Biol. 7, 47 (2007).
Chaubey, G., Metspalu, M., Choi, Y., Magi, R., Romero, I. G., Soares, P. et al. Population genetic structure in Indian Austroasiatic speakers: the role of landscape barriers and sex-specific admixture. Mol. Biol. Evol. 28, 1013–1024 (2011).
Basu, A., Mukherjee, N., Roy, S., Sengupta, S., Banerjee, S., Chakraborty, M. et al. Ethnic India: a genomic view, with special reference to peopling and structure. Genome Res. 13, 2277–2290 (2003).
Cordaux, R., Saha, N., Bentley, G. R., Aunger, R., Sirajuddin, S. M. & Stoneking, M. Mitochondrial DNA analysis reveals diverse histories of tribal populations from India. Eur. J. Hum. Genet. 11, 253–264 (2003).
Kivisild, T., Rootsi, S., Metspalu, M., Mastana, S., Kaldma, K., Parik, J. et al. The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations. Am. J. Hum. Genet. 72, 313–332 (2003).
Palanichamy, M. G., Sun, C., Agrawal, S., Bandelt, H. J., Kong, Q. P., Khan, F. et al. Phylogeny of mitochondrial DNA macrohaplogroup N in India, based on complete sequencing: implications for the peopling of South Asia. Am. J. Hum. Genet. 75, 966–978 (2004).
Chaubey, G., Karmin, M., Metspalu, E., Metspalu, M., Selvi-Rani, D., Singh, V. K. et al. Phylogeography of mtDNA haplogroup R7 in the Indian peninsula. BMC Evol. Biol. 8, 227 (2008).
Reich, D., Thangaraj, K., Patterson, N., Price, A. L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Moorjani, P., Thangaraj, K., Patterson, N., Lipson, M., Loh, P. R., Govindaraj, P. et al. Genetic evidence for recent population mixture in India. Am. J. Hum. Genet. 93, 422–438 (2013).
Juyal, G., Mondal, M., Luisi, P., Laayouni, H., Sood, A., Midha, V. et al. Population and genomic lessons from genetic analysis of two Indian populations. Hum. Genet. 133, 1273–1287 (2014).
Elhaik, E., Greenspan, E., Staats, S., Krahn, T., Tyler-Smith, C., Xue, Y. et al. The GenoChip: a new tool for genetic anthropology. Genome Biol. Evol. 5, 1021–1031 (2013).
Elhaik, E., Tatarinova, T., Chebotarev, D., Piras, I. S., Maria Calo, C., De Montis, A. et al. Geographic population structure analysis of worldwide human populations infers their biogeographical origins. Nat. Commun. 5, 3513 (2014).
Miller, S. A., Dykes, D. D. & Polesky, H. F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Haak, W., Balanovsky, O., Sanchez, J. J., Koshel, S., Zaporozhchenko, V., Adler, C. J. et al. Ancient DNA from European early neolithic farmers reveals their near eastern affinities. PLoS Biol. 8, e1000536 (2010).
Yang, W. Y., Novembre, J., Eskin, E. & Halperin, E. A model-based approach for analysis of spatial structure in genetic data. Nat. Genet. 44, 725–731 (2012).
Kozlov, K., Chebotarov, D., Hassan, M., Triska, M., Triska, P., Flegontov, P. et al. Differential evolution approach to detect recent admixture. BMC Genomics. VarI-Sig 2014 Special Issue (2015).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Danecek, P., Auton, A., Abecasis, G., Albers, C. A., Banks, E., DePristo, M. A. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Cavalli-Sforza, L. L. Human evolution and its relevance for genetic epidemiology. Annu. Rev. Genomics Hum. Genet. 8, 1–15 (2007).
Shi, W., Ayub, Q., Vermeulen, M., Shao, R. G., Zuniga, S., van der Gaag, K. et al. A worldwide survey of human male demographic history based on Y-SNP and Y-STR data from the HGDP-CEPH populations. Mol. Biol. Evol. 27, 385–393 (2010).
Novembre, J. & Ramachandran, S. Perspectives on human population structure at the cusp of the sequencing era. Annu. Rev. Genomics Hum. Genet. 12, 245–274 (2011).
Alexander, D. H. & Lange, K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics 12, 246 (2011).
Champaklakshmi, R. in Structure and Society in Early South India: Essays in Honour of Noboru Karashima (ed. Hall, K. R.) 59–84 (Oxford University Press, Delhi, India, 2001).
Mastana, S. S. & Papiha, S. S. Genetic variability of transferrin subtypes in the populations of India. Hum. Biol. 70, 729–744 (1998).
Metspalu, M., Kivisild, T., Metspalu, E., Parik, J., Hudjashov, G., Kaldma, K. et al. Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans. BMC Genet. 5, 26 (2004).
Gayden, T., Cadenas, A. M., Regueiro, M., Singh, N. B., Zhivotovsky, L. A., Underhill, P. A. et al. The Himalayas as a directional barrier to gene flow. Am. J. Hum. Genet. 80, 884–894 (2007).
Reddy, B. M., Langstieh, B. T., Kumar, V., Nagaraja, T., Reddy, A. N., Meka, A. et al. Austro-Asiatic tribes of Northeast India provide hitherto missing genetic link between South and Southeast Asia. PLoS ONE 2, e1141 (2007).
Shastri, K. A. N. A History of South India: from Prehistoric Times to the Fall of Vijayanagar 4th edn (Oxford University Press, Madras, India, 1976).
Cai, X., Qin, Z., Wen, B., Xu, S., Wang, Y., Lu, Y. et al. Human migration through bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum revealed by Y chromosomes. PLoS ONE 6, e24282 (2011).
Karafet, T. M., Mendez, F. L., Meilerman, M. B., Underhill, P. A., Zegura, S. L., Hammer, M. F. et al. New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res. 18, 830–838 (2008).
Acknowledgements
We gratefully acknowledge the volunteers from India and also thank the community leaders and fieldwork assistants who helped in sampling expeditions and sample identification. The Genographic Project was supported and funded by National Geographic Society, IBM and The Ted Wait Family Foundation, USA. TVT was supported by NIH, GM068968, and NIH-NICHD, HD070996.
Disclaimer
The funding agencies (National Geographic Society, IBM and The Ted Wait Family Foundation) had no role in design of the study, collection and analysis of data, and decision to publish.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
The participants of the Genographic Consortium are as follows: Christina J Adlera1, Elena Balanovska2, Oleg Balanovsky2, Jaume Bertranpetit3, Andrew C Clarke4, David Comas3, Alan Cooper1, Clio SI Der Sarkissian1, Matthew C Dulik5, Jill B Gaieski5, Wolfgang Haak1, Marc Haber3,6, Angela Hobbs7, Asif Javed8, Li Jin9, Matthew E Kaplan10, Shilin Li9, Begoña Martínez-Cruz3, Elizabeth A Matisoo-Smith4, Marta Melé3, Nirav C Merchant10, R John Mitchell11, Amanda C Owings5, Laxmi Parida8, Daniel E Platt8, Lluis Quintana-Murci12, Colin Renfrew13, Daniela R Lacerda14, Ajay K Royyuru8, Theodore G Schurr5, Fabrício R Santos14, Himla Soodyall7, David F Soria Hernanz15, Pandikumar Swamikrishnan16, Chris Tyler-Smith17, Pedro Paulo Vieira18, Miguel G Vilar5, R Spencer Wells15, Pierre A Zalloua6, and Janet S Ziegle19 1University of Adelaide, South Australia, Australia; 2Research Centre for Medical Genetics, Russian Academy of Medical Sciences, Moscow, Russia; 3Universitat Pompeu Fabra, Barcelona, Spain; 4University of Otago, Dunedin, New Zealand; 5University of Pennsylvania, Philadelphia, Pennsylvania, United States; 6Lebanese American University, Chouran, Beirut, Lebanon; 7National Health Laboratory Service, Johannesburg, South Africa; 8IBM, Yorktown Heights, NY, USA; 9Fudan University, Shanghai, China; 10University of Arizona, Tucson, AZ, USA; 11La Trobe University, Melbourne, Victoria, Australia; 12Institut Pasteur, Paris, France; 13University of Cambridge, Cambridge, United Kingdom; 14Universidade Federal de Minas Gerais, Belo Horizonte, Minas Gerais, Brazil; 15National Geographic Society, Washington, DC, USA; 16IBM, Somers, NY, USA; 17The Wellcome Trust Sanger Institute, Hinxton, United Kingdom; 18Universidade Federal do Rio de Janeiro, Rio de Janeiro, Brazil; and 19Applied Biosystems, Foster City, CA, USA.
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
ArunKumar, G., Tatarinova, T., Duty, J. et al. Genome-wide signatures of male-mediated migration shaping the Indian gene pool. J Hum Genet 60, 493–499 (2015). https://doi.org/10.1038/jhg.2015.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.51
This article is cited by
-
Contrasting maternal and paternal genetic histories among five ethnic groups from Khyber Pakhtunkhwa, Pakistan
Scientific Reports (2022)
-
Mitochondrial DNA diversity in the Khattak and Kheshgi of the Peshawar Valley, Pakistan
Genetica (2020)
-
WhoGEM: an admixture-based prediction machine accurately predicts quantitative functional traits in plants
Genome Biology (2019)
-
On Historic migration to South Asia in the last two millennia: A case of Jews and Parsi populations By Ajai Kumar Pathak, et al.
Journal of Biosciences (2019)
-
Ancient Human Migrations to and through Jammu Kashmir- India were not of Males Exclusively
Scientific Reports (2018)