Abstract
Jacobsen syndrome (JBS) is a haploinsufficiency syndrome caused by partial deletion of the long arm of chromosome 11. It is characterized by developmental delay (DD)/mental retardation (MR), physical growth retardation, facial dysmorphism, visceral malformations and thrombocytopenia. We report two JBS patients from China out of a total of 451 patients with unexplained DD/MR. The genotypes of these patients were compared with earlier reported patients in North America and Europe. Both patients presented with severe DD, microcephaly and facial dysmorphism; one patient had a low birth weight, congenital heart disease and structural brain abnormalities. Neither patient was thrombocytopenic at the time of diagnosis. The two deletions were 4.1 and 12.8 Mb. The 4.1 Mb deletion is the smallest of all pathogenic regions earlier reported in JBS. Therefore, the critical region underlying DD/MR might be located in the distal portion of the chromosomal segment within 4.1 Mb of the telomere. Candidate genes for DD/MR in this region include SNX19, THYN1, OPCML, NCAPD3 and NTM. One of the critical regions for craniofacial abnormalities may be within 130.3–134.4 Mb in chromosome 11q. Further analysis of Chinese JBS patients would elucidate the relation of phenotype to genotype further.
Similar content being viewed by others
Introduction
Jacobsen syndrome (JBS: OMIM 147791) is a contiguous gene syndrome resulting from the partial deletion of chromosome 11q. This syndrome was first described by Jacobsen in 1973.1 The deletion size reported earlier ranges from 7 to 20 Mb.2, 3 The deletions usually extend to the telomere. The breakpoints occur within or distal to 11q23.3. Approximately 200 cases of JBS have been reported to date.3, 4 The incidence of distal 11q deletions in the population is difficult to estimate, but the occurrence of JBS is ∼1 in 100 000 births,2, 3, 4 and the female:male ratio is 2:1.
The typical features of JBS include developmental delay (DD) and mental retardation (MR), failure to thrive, macrocephaly, trigonocephaly, facial dysmorphism, visceral malformations and Paris–Trousseau thrombocytopenia. But none of these features are invariably present.
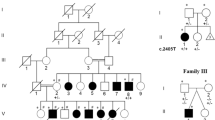
In JBS, most chromosomal aberrations (84.6% of cases) are usually de novo deletions.4 The remaining cases result from an unbalanced segregation of a familial balanced translocation, as was the case in the original report by Jacobsen,1 unbalanced translocations occurring de novo, or other chromosomal rearrangements, such as a ring chromosome 11.5, 6
In a proportion of individuals with JBS, the breakpoints cluster in 11q23.3, which is close to a rare folate-sensitive fragile site, FRA11B.5, 7 Folate-sensitive fragile sites are caused by the expansion of CGG trinucleotide repeats and hypermethylation of adjacent CpG islands. However, only ∼10% cases of JBS are related to amplification of the CCG triplet repeat at the FRA11B fragile site, and individuals with the FRA11B fragile site rarely have children with JBS. Therefore, the presence of the FRA11B fragile site may raise the risk of the offspring of affected individuals having an 11q deletion, but this does not necessarily lead to the children developing JBS. Alternatively, some smaller deletions are likely to occur through different mechanisms; for example, chromosome recombinations can be mediated by palindromic AT-rich repeats, low copy repeats or olfactory receptor gene clusters.8, 9, 10 Consequently, there are likely to be multiple mechanisms that result in the generation of deletions in long arm of chromosome 11.
The aims of this study were to report the first patients with JBS in China, to establish the fine location of the deletion region and to analyze the phenotypes and genotype characteristics of the individuals.
Materials and methods
Patients
From December 2006 to July 2009, 451 Chinese patients with unexplained DD/MR who met the inclusion criteria were examined in the pediatric neurological clinics of Peking University First Hospital.
Inclusion criteria
The inclusion criteria include (1) moderate to severe DD/MR; (2) no known perinatal brain injury; (3) no history of toxication, hypoxia, infection of central nervous system and cranial trauma; (4) normal routine karyotyping; (5) no evidence of typical inherited metabolic disorder or specific neurodegenerative disorders from cranial neuro-imaging and blood/urinary metabolic diseases screening; (6) no mutation of FMR1 in male patients; and (7) no typical clinical manifestation of Rett syndrome in female patients.
Multiplex ligation-dependent probe amplification for screening of subtelomeric rearrangements
Multiplex ligation-dependent probe amplification (MLPA) was used with a new set of probes that were designed for the detection of aberrations in a larger sample. It is a robust and easy technique that can detect both deletions and duplications.11 The approximate resolution of MLPA is high enough to detect several bases, but it is limited to specific regions of the chromosomes.
Screening of subtelomeric rearrangements was carried out in all 451 patients. A set of probes was specifically designed to test for subtelomeric imbalances using the SALSA P070 human telomere test kit (MRC-Holland, Amsterdam, Netherlands; http://www.mrc-holland.com), and these were used for first-stage screening. For each patient with a positive result, a subsequent second-stage test was performed with a P036B kit (MRC-Holland, Amsterdam, Netherlands). The parents of patients who had positive results confirmed by the second kit were tested to assess whether the genetic aberrations were de novo or inherited.
Affymetrix human single nucleotide polymorphism array 6.0 assay
If the subtelomeric aberrations detected by MLPA of patients were de novo, then we used the affymetrix genome-wide human single nucleotide polymorphism (SNP) array 6.0 to confirm and accurately define the exact size of each subtelomeric aberrant region. This new array features 1.8 million genetic markers, including >906 600 SNPs and >946 000 probes for the detection of copy number variation (CNV) with a ∼1.4 kb resolution. The high density of the probes across the genome, as well as the combination of SNP array with array comparative genomic hybridization in a single chip, makes it possible to define the size of each aberrant region accurately. Partek software (version 6.3) was used for the analysis of CNVs. The determination of duplicated or deleted regions in each sample was by the hidden Markov model (HMM) calculation. The involvement of at least three contiguous probe sets was required, and when the copy number calculated by HMM was <1.5, it was considered to be a deletion; if the copy number was >2.5, this was considered to be a duplication.
Results
Phenotype of patients with JBS
The clinical presentations of the two JBS patients are presented in Table 1. Their presenting signs and symptoms are summarized in Table 2 and compared with those earlier reported.12 Both patients presented with severe DD, microcephaly and facial dysmorphism; one patient had a low birth weight, congenital heart disease and structural brain abnormalities. Neither patient was thrombocytopenic at the time of diagnosis.
Genotype of patients with JBS
Of the 451 children that were recruited with clinically unexplained DD/MR, screening with MLPA revealed an 11q deletion in two patients. The subsequent MLPA assay for their parents showed that all of these rearrangements had occurred de novo.
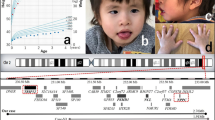
To confirm and accurately define the exact size of each subtelomeric aberrant region that was detected by MLPA, a subsequent assay with the affymetrix genome-wide human SNP array 6.0 was performed in these two patients. The starting points, ending points, lengths and exact locations of each 11q deletion are presented in Table 3. The total length of the deletions were 4.1 and 12.8 Mb. The 4.1 Mb deletion is smaller than any other deletion that has been earlier reported in a patient with JBS. The location of the deletions for both patients was between 11q23.3 and 11q25.
Discussion
In this study, we report the clinical and molecular cytogenetic features of two individuals with DD, microcephaly and facial dysmorphism who were found to have de novo submicroscopic rearrangements in the long arm of chromosome 11. In Patient 1, the chromosomal rearrangement consisted of a 4.1 Mb deletion (130.33–134.4 Mb) and this region is smaller than any that has been reported earlier. In Patient 2, a 12.8 Mb deletion was found (119.98–132.81 Mb). Their deletions lie within the region that is commonly deleted in patients with JBS.13 It seems that the patients with larger deletions in 11q show a more severe phenotype, but this may vary among patients. For example, most patients with JBS have Paris–Trousseau syndrome (PTS), but only 56% of them have congenital heart disease, and this phenotype is independent of the size of their 11q deletion. Both of our patients had DD and facial dysmorphism, and one patient also had cardiac defects and skeletal abnormalities. Neither of our patients exhibited thrombocytopenia or pancytopenia at the time of diagnosis, although this is an important feature of JBS. However, we neither have the record of the platelet count at birth, nor the results of platelet function studies and the analysis of peripheral blood for the presence of giant platelets until the time of this study. Both thrombocytopenia in this syndrome and PTS are characterized by thrombocytopenia at birth, which usually improves during childhood. JBS is also commonly characterized by platelet dysfunction and the presence of giant platelets in the blood, although the platelet count might be normal. Therefore, we could not completely exclude the presence of PTS in these two patients, although no clinical presentation of platelet dysfunction and thrombocytopenia was found. It would be beneficial to collect these data in future to establish whether this is a typical feature of JBS in Chinese patients.
Developmental delay
It was reported that psychomotor development is normal in ∼3% of JBS cases, and mild to severe MR is observed in ∼97% of cases.12 The severity of MR is strongly linked to the size of the deletion.3, 14 Coldren et al. reported that JBS patients with deletion sizes ⩾12.1 Mb have severe cognitive impairment, whereas patients with deletion sizes ⩽11.8 Mb have mild cognitive impairment. Furthermore, their study revealed that patients with deletion sizes ⩽11.8 Mb, but ⩾9.1 Mb, may have a selective impairment in freedom from distractability compared with the patients with smaller deletions that are ⩽9.1 Mb.14 Behavioral problems such as attention deficit hyperactivity disorder were reported. However, more severe psychiatric disorders, such as schizophrenia, were reported infrequently15, 16 and seizures were rarely reported.
The location of 11q23-qter is a gene-rich region. Grossfeld et al. have defined some critical regions for 14 individual phenotypes. The smallest critical region (6.8 Mb) that they found extended from D11S1351 (127.5 Mb) to the telomere, and this region was associated with MR.3 B3GAT1 is expressed in the brain, and this gene is located in the 11q25 at 133.77 Mb. Yamamoto et al.17 reported that when this gene was knocked out, the mice showed impaired synaptic plasticity and learning disorders. Coldren et al. have reported two critical regions containing candidate genes for global and selective deficits in neurocognitive function ‘BSX for global cognitive function and Neurogranin, a gene essential for synapse plasticity and long-term potentiation, for auditory attention’.14
Patient 1 has a 4.1 Mb deletion in 11q25, which spans a region between 130.3 and 134.5 Mb in size. Furthermore, the deleted region in this patient is smaller than any other regions reported earlier. The phenotype of this patient includes DD, microcephaly and some facial dysmorphisms. Therefore, the critical region underlying the phenotype of DD/MR in 11q terminal deletion disorder might be narrowed by the results of our study to the very distal part of the long arm of chromosome 11 to within 4.1 Mb of the telomere. Twenty-one genes reside within this deleted region, and 14 of these are expressed in the human brain. SNX19, THYN1, OPCML, NCAPD3 and NTM might be candidate genes because of their important function in the central nervous system, such as phosphoinositide binding, mitotic chromosome assembly/segregation, neurite outgrowth or apoptosis. The severe DD that was found in Patient 2, who had an interstitial deletion, was a result that was consistent with the findings of Coldren et al.14
Alternatively, the DD/MR in one or both of these patients may be due to other genetic factors. This is suggested by the presence of microcephaly in both of these patients, which is an unusual finding in JBS.
Craniofacial abnormalities
Typical features of JBS include skull deformities, such as a high prominent forehead, macrocrania and trigonocephaly, and facial dysmorphisms, such as ocular hypertelorism, hypertelorism, ptosis, down-slanting palpebral fissures, a broad nasal bridge with a short nose, thin upper lips and low-set ears. A region that influences craniofacial development was believed to be mapped distal to D11S1351(127.6 Mb).2 This region contains BARX2. This gene contains four exons and three introns and encodes for a homeobox protein. Krasner et al.18 found that its murine homolog barx2 was expressed in the developmental nerves and cephalofacial structures. This region was deleted in Patient 2 in our study, and she displays some of the facial dysmorphisms that can be seen in a significant proportion of individuals with JBS. However, this region is not in the deleted region of Patient 1, who also displays typical craniofacial abnormalities. Furthermore, Krasner et al.18 reported no mutation in the BARX2 gene in nine patients with isolated tigonocephaly. Therefore, it can be presumed that there may be other genes or regions that account for the craniofacial abnormalities in individuals with JBS. According to our results, genes or regions that map to the distal part of the long arm of chromosome 11, within 4.1 Mb of the telomere, should be considered.
Congenital heart malformations
Congenital heart diseases occur in 56% of patients with JBS. Many kinds of serious cardiac abnormalities are observed in these individuals. The most common heart defects are ventricular septal defects and left heart obstructive malformations. Critical genes or regions that influence cardiac development are believed to map to a locus that is distal to D11S707 (125.8 Mb).3 In our study, the deletion of 11q in Patient 1 only consisted of a 4.1 Mb region (from 130.3 to 134.4 Mb) and he had no congenital heart malformations. Moreover, the deletion in Patient 2, who had a ventricular septum defect, spanned the entire critical region that was mentioned above. These results are consistent with earlier reports. Deletion of ETS-1 in mice, one gene in this region, has recently been shown to cause ventricular septal defects.19
Conclusions
The characteristics of two patients with JBS from China appear to be consistent with those reported earlier. Genotype–phenotype analysis in two individuals with submicroscopic 11q23.3–25 rearrangements facilitated the refinement of the JBS critical region or genes. We suggest that the critical region underlying DD/MR might be in the distal 11q within 4.1 Mb of the telomere, and, therefore, SNX19, THYN1, OPCML, NCAPD3 and NTM might be the candidate genes. One of the critical regions for craniofacial abnormalities may be within 130.3–134.4 Mb. Genotype–phenotype analysis is notable for the variable penetrance of specific phenotypes despite the deletion of proposed critical regions. We also realize that some smaller CNVs may not have been detected by our assay, and these might contribute to the phenotype of JBS in some way. It is hoped that additional cases of JBS can be studied in the future and more finely analyzed in the future, so that a more accurate relationship between phenotype and genotype in Chinese patients with JBS may be defined.
References
Jacobsen, P., Hauge, M., Henningsen, K., Hobolth, N., Mikkelsen, M. & Philip, J. An (11; 21) translocation in four generations with chromosome 11 abnormalities in the offspring. A clinical, cytogenetical, and gene marker study. Hum. Hered. 23, 568–585 (1973).
Penny, L. A., Dell’Aquila, M., Jones, M. C., Bergoffen, J., Cunnif, C., Fryns, J. P. et al. Clinical and molecular characterization of patients with distal 11q deletion. Am. J. Hum. Genet. 56, 676–683 (1995).
Grossfeld, P. D., Mattina, T., Lai, Z., Favier, R., Jones, K. L., Cotter, F. et al. The 11q terminal deletion disorder: a prospective study of 110 cases. Am. J. Med. Genet. 129A, 51–61 (2004).
Pivnick, E. K., Velagaleti, G. V., Wilroy, R. S., Smith, M. E., Rose, M. E., Tipton, R. E. et al. Jacobsen syndrome: report of a patient with severe eye anomalies, growth hormone deficiency, and hypotiroidism associated with deletion 11 (q23q25) and review of 52 cases. J. Med. Genet. 33, 772–778 (1996).
Niikawa, N., Jinno, Y., Tomiyasu, T., Kukushima, Y. & Kudo, K. Ring chromosome 11 [(46,XX,r(11)(p15q25)] associated with clinical features of the 1 1q- syndrome. Ann. Genet. 24, 172–175 (1981).
Cousineau, A. J., Higgins, J. V., Scott-Emaukpor, A. B. & Mody, G. Brief clinical report: ring-11 chromosome: phenotype-karyotype correlation with deletions of llq. Am. J. Med. Genet. 14, 29–35 (1983).
Voullaire, L. E., Webb, G. C. & Leversha, M. A. Chromosome deletion at 11q23 in an abnormal child from a family with inherited fragility at 11q23. Hum. Gene. 76, 202–204 (1987).
Edelmann, L., Spiteri, E., McCain, N., Goldberg, R., Pandita, R. K., Duong, S. et al. A common breakpoint on 11q23 in carriers of the constitutional t(11,22) translocation. Am. J. Hum. Genet. 65, 1608–1616 (1999).
Kurahashi, H., Shaikh, T. H., Zackai, E. H., Celle, L., Driscoll, D. A., Budarf, M. L. et al. Tightly clustered 11q23 and 22q11 breakpoints permit PCR-based detection of the recurrent constitutional t(11;22). Am. J. Hum. Genet. 67, 763–768 (2000).
Giglio, S., Broman, K. W., Matsumoto, N., Calvari, V., Gimelli, G., Neumann, T. et al. Olfactory receptor gene clusters, genomicinversion polymorphism, and common chromosome rearrangements. Am. J. Hum. Genet. 68, 874–883 (2001).
Shen, Y. & Wu, B L. Designing a simple multiplex ligation-dependent probe amplification (MLPA) assay for rapid detection of copy number variants in the genome. J. Genet. Genomics. 36, 257–265 (2009).
Mattina, T., Perrotta, C S. & Grossfeld, P. Jacobsen syndrome. Orphanet. J. Rare. Dis. 4, 9 (2009).
Fryns, J P., Kleczkowska, A., Buttiens, M., Marien, P. & vanden Berghe, H. Distal 11q monosomy. The typical 11q monosomy syndrome is due to deletion of subband 11q24.1. Clinical. Genetics. 30, 255–260 (1986).
Coldren, C. D., Lai, Z., Shragg, P., Rossi, E., Glidewell, S. C., Zuffardi, O. et al. Chromosomal microarray mapping suggests a role for BSX and Neurogranin in neurocognitive and behavioural defects in the 11q terminal deletion disorder (Jacobsen syndrome). Neurogenetics. 10, 89–95 (2009).
Neavel, C. B. & Soukup, S. Deletion of (11)(q24.2) in a mother and daughter with similar phenotypes. Am. J. Med. Genet. 53, 321–324 (1994).
Böhm, D., Hoffmann, K., Laccone, F., Wilken, B., Dechent, P., Frahm, J. et al. Association of Jacobsen syndrome and bipolar affective disorder in a patient with a de novo 11q terminal deletion. Am. J. Med. Genet. 140, 378–382 (2006).
Yamamoto, S., Oka, S., Inoue, M., Shimuta, M., Manabe, T., Takahashi, H. et al. Mice deficient in nervous system-specific carbohydrate epitope HNK-1 exhibit impaired synaptic plasticity and spatial learning. J. Biol. Chem. 277, 27227–27231 (2002).
Krasner, A., Wallace, L., Thiagalingam, A., Jones, C., Lengauer, C., Minahan, L. et al. Cloning and chromosomal localization of the human BARX2 homeobox protein gene. Gene. 250, 171–180 (2000).
Ye, M., Coldren, C., Liang, X., Mattina, T., Goldmuntz, E., Benson, D. W. et al. Deletion of ETS-1, a gene in the Jacobsen syndrome critical region, causes ventricular septal defects and abnormal ventricular morphology in mice. Hum. Mol. Genet. 19, 648–656 (2010).
Acknowledgements
We thank the patients and their families involved in this study for their cooperation. This work was funded by the National Key Research Project-973 of China (2007CB5119004), and Beijing Municipal Natural Science Key Project (7081004).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ji, T., Wu, Y., Wang, H. et al. Diagnosis and fine mapping of a deletion in distal 11q in two Chinese patients with developmental delay. J Hum Genet 55, 486–489 (2010). https://doi.org/10.1038/jhg.2010.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2010.51
Keywords
This article is cited by
-
Patients with Chromosome 11q Deletions Are Characterized by Inborn Errors of Immunity Involving both B and T Lymphocytes
Journal of Clinical Immunology (2022)
-
Detecting TF-miRNA-gene network based modules for 5hmC and 5mC brain samples: a intra- and inter-species case-study between human and rhesus
BMC Genetics (2018)
-
de novo interstitial deletions at the 11q23.3-q24.2 region
Molecular Cytogenetics (2016)
-
Fragile X syndrome screening in Chinese children with unknown intellectual developmental disorder
BMC Pediatrics (2015)
-
Application of custom-designed oligonucleotide array CGH in 145 patients with autistic spectrum disorders
European Journal of Human Genetics (2013)