Abstract
Attempts at generating novel chemistries by genetically manipulating polyketide synthases (PKSs) usually result in no detectable or poor product yield. Understanding processes that drive the evolution of PKSs might provide a solution to this problem. The synonymous-to-non-synonymous nucleotide substitution ratios across alignments of well-characterized PKS modules were examined using a sliding windows approach. Not surprisingly, the overall substitution ratios showed that PKS modules are generally under strong purifying selection, confirming experimental observations that changes to the primary amino acid sequence, regardless of whether these changes are conservative or not, will most likely result in some loss in function. Despite the masking effect of negative selection, by judicious choice of window size, it was possible to recognize amino acid residues that appear to be under strong positive selection. The importance of these amino acids has not been recognized by other analysis methods before and we suggest that they may function to ‘fine tune’ modular PKSs. Future efforts will concentrate on understanding if this ‘fine tuning’ is at the level of protein expression, for example, transcription or translation, or at the level of protein function, for example, efficient selection and channeling of acyl intermediates between domains.
Similar content being viewed by others
Introduction
Polyketide secondary metabolites have diverse roles in the chemical ecology of the organisms that produce them, as well as being of economic importance as natural products to the pharmaceutical and agrochemical industries.1, 2 The biosynthesis of many polyketides is catalyzed through successive condensation of acyl-thioester units on modular polyketide synthases (PKSs), the best studied being obtained from Streptomyces and related actinomycete bacteria. These large enzymes consist of multi-domain polypeptides that pass the growing polyketide chain from the active site of one catalytic domain to the next, generating chemical diversity depending on the substrate specificity of each domain. Assembly lines of domains can be grouped into modules, with the core of each module consisting of a keto-synthase domain (KS), an acyltransferase domain (AT) and an acyl-carrier protein (ACP) domain. The KS-AT-ACP domains extend the growing polyketide chain by two carbon atoms, generating an ACP-bound β-ketoacyl intermediate. This β-keto group can be further reduced by optional accessory ketoreductase (KR), dehydratase (DH) and enoyl reductase (ER) domains. The co-linear arrangement of modules is also mirrored in the gene clusters that encode them, offering exciting possibilities for combinatorial biosynthesis.3 However, experiments to manipulate modular biosynthetic clusters to create novel chemistries often result in no detectable product or product yield is extremely low.4, 5
Many mechanisms have been evoked to explain the evolution of modular PKSs, including gene duplication, deletion, recombination and horizontal gene transfer.6, 7, 8, 9, 10 However, although these processes offer mechanisms that introduce genetic variation into a gene cluster, they do not take into account natural selection acting on individual nucleotide loci that ultimately influence the phenotype whereby PKSs evolve. The appearance and maintenance of protein function can be explained in terms of positive (adaptive) and negative (purifying) natural selection with many residues, understandably, being under strong negative selection.11 A common way to measure natural selection in orthologous protein-coding nucleotide sequences between species (that is, inter-specific polymorphism) is by estimating the non-synonymous (causing amino acid replacement, Ka) to synonymous (silent, Ks) nucleotide substitution rate (ω=Ka/Ks). As a general rule, values of ω>1, ω∼1 and ω<1 are taken as indicating positive, neutral and negative selection, respectively.12, 13, 14 This can be taken as an average over all codons in a gene, but often, different regions of a gene that encodes a multi-domain protein will be influenced by different selective pressures depending, for example, on the structural or catalytic functions of each domain. This is especially prominent between homologous sequences within a population of the same species (intra-specific polymorphism). In such cases, calculating Ka/Ks as an average over the entire length of the gene will not provide a detailed picture of the selective constraints acting at different positions in the sequence and evolutionary hotspots will be missed. An equivalent measure to ω is to estimate the nucleotide diversity (π) at each position in a multiple alignment of intra-specific sequences.15, 16 The ratio between non-synonymous (πa) to synonymous nucleotide polymorphisms (πs) can then be calculated (πa/πs) and plotted in a sliding window against nucleotide position.17, 18 In this paper, we examine natural selection that acts on successful examples of PKSs with the aim of identifying critical amino acid residues to gain a better overview of the evolutionary constraints that govern functionality, which might, in the future, be exploited for more efficient synthesis of new compounds from hybrid PKSs.
Materials and Methods
Using MEGA 4 software (http://www.megasoftware.net/),19 the DNA sequences from 17 well-annotated modular PKSs (Table 1) extracted from ClustScan20, 21 were translated into proteins and aligned so that modules of approximately equal length and containing the same domain organization could be grouped together. These groups of modules were then back-translated to the corresponding DNA sequences so that polymorphism ratios (πa/πs) could be calculated and graphically displayed using a sliding window analysis in DNASP version 5.10.01, taking a window size of 50 and step size of 10 (http://www.ub.edu/dnasp/).22 The average πa/πs ratio and the s.d. were calculated from non-overlapping windows of length 12.
Results
The modules were extracted from the 17 PKSs selected for this study (Table 1) and grouped into types depending on their structure. Starter modules (loading domains) were not included in the analysis, because their diversity resulted in groups that were too small for reliable statistical analysis. There were only three non-reducing extension modules (KS-AT-ACP), which were also not analyzed further. The remaining extension modules were grouped into four types depending on the domains present (although not all domains are necessarily active):
-
i)
KS-AT-KR-ACP: this was the largest group with 73 members.
-
ii)
KS-AT-DH-KR-ACP: 62 modules with a full-length DH domain.
-
iii)
KS-AT-dhX-KR-ACP: this group had part of the DH domain, which could be recognized by HMMER searches, but usually not with BLAST searches.20, 21 There were 23 modules of this type.
-
iv)
KS-AT-DH-ER-KR-ACP: 22 members.
Protein sequence alignments were carried out for the members of each class and used to generate DNA alignments with codons correctly aligned. The ratio of the nucleotide diversity index for non-synonymous to synonymous changes (πa/πs) was calculated and averaged in 50-nt sliding windows along each group of modules.
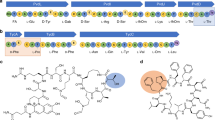
Figure 1 shows the πa/πs ratio along Group I modules. The lowest values of the ratio occur in the KS domain and the ratio remains low over much of this domain, suggesting that there is strong purifying selection that is, many of the residues in KS cannot be changed without losing function. The highest value of the ratio also occurs in the KS domain. This value is approximately 1, which initially suggested that this might be a region that is nearly neutral with respect to selection. A more detailed examination using a 12-nt sliding window showed that there is a double peak. The first peak corresponded to four residues (V153-F156), which were located using the 3-D crystal structure of the erythromycin KS3-AT3 didomain (accession number 2QO3). These residues are at the interaction interface for dimerization of the PKS subunits. The second peak (residues G159-Y171) corresponded to a surface loop poorly defined in the 3-D structure, with different modules having differing numbers of residues in this region that is indels. The double peak in the πa/πs ratio was also present in the KS domains of the other three module groups (Figures 2, 3 and 4).
Sliding window analysis of the synonymous-to-non-synonymous nucleotide substitution ratios across an alignment of 73 PKS modules with the domain architecture KS-AT-KR-ACP. Arrows show a double peak in the KS domain (regions under positive and neutral selection respectively) and the position of the substrate-determining F/S residue indicated by the symbol •. Lines show the mean value±s.d. of the ratio for the whole module.
Sliding window analysis of the synonymous-to-non-synonymous nucleotide substitution ratios across an alignment of 62 PKS modules with the domain architecture KS-AT-DH-KR-ACP, where the DH domain is predicted to be functional. Arrows show a double peak in the KS domain (regions under positive and neutral selection, respectively) and the position of the substrate-determining F/S residue indicated by the symbol •. Lines show the mean value±s.d. of the ratio for the whole module.
Sliding window analysis of the synonymous-to-non-synonymous nucleotide substitution ratios across an alignment of 23 PKS modules with the domain architecture KS-AT-dhX-KR-ACP, in which the DH domain is not predicted to be functional. Arrows show a double peak in the KS domain (regions under positive and neutral selection respectively) and the position of the substrate-determining F/S residue indicated by the symbol •. Lines show the mean value±s.d. of the ratio for the whole module.
Sliding window analysis of the synonymous-to-non-synonymous nucleotide substitution ratios across an alignment of 22 PKS modules with the domain architecture KS-AT-DH-ER-KR-ACP. Arrows show a double peak in the KS domain (regions under positive and neutral selection respectively) and the position of the substrate-determining F/S residue indicated by the symbol •. Lines show the mean value±s.d. of the ratio for the whole module.
In general, the πa/πs ratios were higher in other domains, as were the ratios for the linkers between these domains compared with the linker between the KS and AT. The AT domains show a similar pattern in the πa/πs ratio to the KS in all four module groups, with peaks in the sliding window corresponding to regions under positive or neutral selection, interdispersed with regions under purifying selection. The residues responsible for substrate specificity appear, on first inspection, to be under positive selection. In particular, there is an F/S choice corresponding to incorporation of C2/C3 units.23, 24, 25, 26, 27, 28 However, the ratios in windows containing this residue are low (Figures 1, 2, 3 and 4). A closer examination of the region using a smaller window size reveals that the higher ratio associated with the F/S choice is hidden by averaging with the low ratios associated with neighboring highly conserved residues. There is a peak close to the end of the AT domain and as with the KS, this peak corresponds to codons under positive selection specifying previously innocuous amino acids. The ACP and reductive domains also show similar patterns in all four module classes (Figures 1, 2, 3 and 4), which are yet to be analyzed in greater detail.
Discussion
The πa/πs ratios are low (much less than 1) for most of the regions of the modules (Figures 1, 2, 3 and 4). This is expected, as PKS sequences are highly conserved and likely to be subjected to strong purifying selection. Also, experiments manipulating PKS clusters show that most changes result in large drops in product yield, suggesting that residues cannot be easily changed without loss in PKS function. The KS domain shows the highest degree of sequence conservation and, not surprisingly, has low ratios for most of its length. However, there is a prominent double peak present in KS domains in all four groups of modules. This peak has a ratio of approximately 1, which initially suggests that it might correspond to a relatively unimportant region of the protein that is nearly neutral with respect to selection. Location of the residues in a 3-D crystal structure (accession number 2QO3) shows that one component of the peak (residues G159-Y171) corresponds to a surface loop that may well be nearly neutral for selection. However, the other component (residues V153-F156) lies on the interaction interface for dimerization of the PKS subunits. It seems unlikely that this sequence is selectively neutral. The residues differ markedly between modules of a single cluster and it is conceivable that they have a part in ensuring homodimerisation rather than heterodimerization.
Not all selected residues are revealed by this approach. Thus, a residue involved in substrate specificity of AT domains showed a low ratio. Although the amino acid choice F/S is important for the selection of C2/C3 extension, respectively, the residue is embedded in a highly conserved region so that the low ratios for neighboring amino acids masks the signal from the selected residue. Such problems can be partially solved by optimizing window sizes to achieve maximum sensitivity,29 but is unlikely to detect the specificity-determining residues in AT domains, as they are scattered through the protein primary sequence and occur in regions of sequence conservation.20, 21
The high degree of amino acid conservation in PKS modules implies that there is a strong purifying selection so that most nucleotides will show low ratios of πa/πs (≪1) and detection of positively selected residues is difficult. However, the example in KS of the four residues in the dimer interface region shows that careful analysis of peaks can identify interesting sequences. Likewise, peaks corresponding to codons specifying previously unsuspecting amino acids also under positive selection can be identified in all other domains used in this study and must now be examined in greater detail. The low productivity of many manipulated constructs suggests that a subsequent ‘fine-tuning’ of the structure of the module is necessary. It is likely that natural selection drives a similar process during evolution of clusters, and the analysis methods used in this paper may reveal critical residues necessary for this ‘fine-tuning’ in hybrid PKSs to achieve useful product yields.
References
Yim, G., Wang, H. H. & Davies, J. The truth about antibiotics. Int. J. Med. Microbiol. 296, 163–170 (2006).
Damain, A. L. & Sanchez, D. Microbial drug discovery: 80 years of progress. J. Antibiot. 62, 5–16 (2009).
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Ed. Engl. 48, 4688–4716 (2009).
Hranueli, D., Cullum, J., Basrak, B., Goldstein, P. & Long, P. F. Plasticity of the Streptomyces genome—evolution and engineering of new antibiotics. Curr. Med. Chem. 12, 763–771 (2005).
Weissman, K. J. & Leadlay, P. F. Combinatorial biosynthesis of reduced polyketides. Nat. Rev. Microbiol. 3, 925–936 (2005).
Callahan, B., Thattai, M. & Shraiman, B. I. Emergent gene order in a model of modular polyketide synthases. Proc. Natl Acad. Sci. USA. 106, 19410–19415 (2009).
Jenke-Kodama, H. & Dittmann, E. Evolution of metabolic diversity: insights from microbial polyketide synthases. Phytochemistry. 70, 1858–1866 (2009).
Fischbach, M. A., Walsh, C. T. & Clardy, J. The evolution of gene collectives: how natural selection drives chemical innovation. Proc. Natl Acad. Sci. USA 105, 4601–4608 (2008).
Ridley, C. P., Lee, H. Y. & Khosla, C. Evolution of polyketide synthases in bacteria. Proc. Natl Acad. Sci. USA. 105, 4595–4600 (2008).
Jenke-Kodama, H., Sandmann, A., Müller, R. & Dittmann, E. Evolutionary implications of bacterial polyketide synthases. Mol. Biol. Evol. 22, 2027–2039 (2005).
Ohta, T. An examination of the generation time effect on modular evolution. Proc. Natl Acad. Sci. USA. 90, 10676–10680 (1993).
Sharpe, P. M. In search of molecular Darwinism. Nature 385, 111–112 (1997).
Akashi, H. Within-and between-species DNA sequence variation and the ‘footprint’ of natural selection. Gene 238, 39–51 (1999).
Crandall, K. A., Kelsey, C. R., Imanichi, H., Lane, H. C. & Salzman, N. P. Parallel evolution of drug resistance in HIV: failure of non-synonymous/synonymous substitution rate ratio to detect selection. Mol. Biol. Evol. 16, 372–382 (1999).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl Acad. Sci. USA. 76, 5269–5273 (1979).
Kryazhimskiy, S. & Plotkin, J. B. The population genetics of dN/Ds. PLoS Genetics 12, e1000304 (2008).
Tajima, F. Determination of window size for analysing DNA sequences. J. Mol. Evol. 33, 470–473 (1991).
Hughes, A. L. & Nei, M. Nucleotide substitution at major histocompatibility complex class II loci: evidence for overdominant selection. Proc. Natl Acad. Sci. USA. 86, 958–962 (1989).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Starcevic, A. et al. ClustScan: an integrated program package for the semi-automatic annotation of modular biosynthetic gene clusters and in silico prediction of novel chemical structures. Nucleic Acids Res. 36, 6882–6892 (2008).
Zucko, J. et al. From DNA sequences to chemical structures—methods for mining microbial genomic and metagenomic datasets for new natural products. Food Technol. Biotechnol. 48, 234–242 (2010).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Kimura, M. Preponderance of synonymous changes as evidence for the neutral theory of molecular evolution. Nature 267, 275–276 (1977).
Kimura, M. Possibility of extensive neutral evolution under stabilizing selection with special reference to nonrandom usage of synonymous codons. Proc. Natl Acad. Sci. USA. 78, 5773–5777 (1981).
Nielsen, R. & Yang, Z. Likelihood models for detecting positively selected amino acid sites and application to the HIV envelope gene. Genetics 148, 929–936 (1998).
Suzuki, Y. & Gojobori, T. A method for detecting positive selection at single amino acid sites. Mol. Biol. Evol. 16, 1315–1328 (1999).
Yang, Z., Neilsen, R., Goldman, N. & Pedersen, A. M- K. Codon substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155, 431–449 (2000).
Yadav, G., Gokhale, R. S. & Mohanty, D. Computational approach for prediction of domain organization and substrate specificity of modular polyketide synthases. J. Mol. Biol. 328, 335–363 (2003).
Fares, M. A., Elena, S. F., Ortiz, J., Moya, A. & Barrio, E. A sliding-window based method to detect selective constraints in protein coding genes and its application to viruses. J. Mol. Evol. 55, 509–521 (2002).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to late Dr C Richard Hutchinson for his exceptional contributions to natural product biosynthesis, engineering and drug discovery.
Rights and permissions
About this article
Cite this article
Zucko, J., Cullum, J., Hranueli, D. et al. Evolutionary dynamics of modular polyketide synthases, with implications for protein design and engineering. J Antibiot 64, 89–92 (2011). https://doi.org/10.1038/ja.2010.141
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2010.141
Keywords
This article is cited by
-
Engineering of PKS Megaenzymes—A Promising Way to Biosynthesize High-Value Active Molecules
Topics in Catalysis (2022)
-
Evolutionary concepts in natural products discovery: what actinomycetes have taught us
Journal of Industrial Microbiology and Biotechnology (2014)
-
Databases of the thiotemplate modular systems (CSDB) and their in silico recombinants (r-CSDB)
Journal of Industrial Microbiology and Biotechnology (2013)
-
Horizontal gene transfer and gene conversion drive evolution of modular polyketide synthases
Journal of Industrial Microbiology and Biotechnology (2012)
-
Recombinatorial biosynthesis of polyketides
Journal of Industrial Microbiology and Biotechnology (2012)