Abstract
Formation of bacterial biofilm communities leads to profound physiological modifications and increased physical and metabolic exchanges between bacteria. It was previously shown that bioactive molecules produced within the biofilm environment contribute to bacterial interactions. Here we describe new pore-forming colicin R, specifically produced in biofilms formed by the natural isolate Escherichia coli ROAR029 but that cannot be detected under planktonic culture conditions. We demonstrate that an increased SOS stress response within mature biofilms induces SOS-dependent colicin R expression. We provide evidence that colicin R displays increased activity against E. coli strains that have a reduced lipopolysaccharide length, such as the pathogenic enteroaggregative E. coli LF82 clinical isolate, therefore pointing to lipopolysaccharide size as an important determinant for resistance to colicins. We show that colicin R toxicity toward E. coli LF82 is increased under biofilm conditions compared with planktonic susceptibility and that release of colicin R confers a strong competitive advantage in mixed biofilms by rapidly outcompeting sensitive neighboring bacteria. This work identifies the first biofilm-associated colicin that preferentially targets biofilm bacteria. Furthermore, it indicates that the study of antagonistic molecules produced in biofilm and multispecies contexts could reveal unsuspected, ecologically relevant bacterial interactions influencing population dynamics in natural environments.
Similar content being viewed by others
Introduction
Biofilms are widespread mixed-species bacterial communities in which high cell density, reduced diffusion and heterogeneous structure favor physical and metabolic contact between bacteria. Biofilms are therefore ideal environments for development of antagonistic and synergistic interactions (James et al., 1995; Hibbing et al., 2010; Rendueles and Ghigo, 2012). It was hypothesized early on that the particular conditions prevailing within biofilms could lead to the emergence of biofilm-specific competition mechanisms (Ghigo, 2003; Valle et al., 2008). Consistently, biofilm communities were shown to release molecules that affect bacterial access to space and resources. Examples of such compounds range from metabolites and amino acids (Valle et al., 2008; Kolodkin-Gal et al., 2010, 2012) to anti-adhesion molecules (Bendaoud et al., 2011; Rendueles et al., 2011, 2013) and bacteriocins (Yan et al., 2003; Kreth et al., 2004).
Production of antagonistic molecules is a widespread trait across the microbial world (Chater, 2006; Cascales et al., 2007; Duquesne et al., 2007; Wenzel and Muller, 2009; Hibbing et al., 2010; Lee and Kim, 2011). In the specific context of biofilm development, bacteriocin production has been researched at length in oral biofilms (Kreth et al., 2004; Bucci et al., 2011; Merritt and Qi, 2012) in which Gram-positive bacteria are predominant; however, little is known about the occurrence and efficacy of bacteriocins produced within Gram-negative biofilms.
It is estimated that at least 50% of all Escherichia coli strains synthesize colicins, plasmid-encoded bacteriocins secreted by E. coli and closely related enterobacteria, and microcins (chromosomally encoded low-molecular-weight bacteriocins). Secretion of colicins is considered to provide an ecological advantage to colicin-producing bacteria against their colicin-sensitive neighbors (Gillor et al., 2004; Cascales et al., 2007). Colicinogenic bacteria are resistant to colicin biocidal action via a constitutively expressed immunity gene typically coded downstream the colicin gene. Colicin release into the extracellular medium is mediated by cell lysis, presumably encoded by the kil or lysis gene carried in the plasmid. For lethal activity, colicin interactions with target cells are mediated upon binding to receptors at the surface and subsequent colicin internalization. Colicins have two distinct mechanisms of action; they can either form pores in the inner membrane (activity encoded in the C terminal region of the colicin) or display an enzymatic activity that will degrade DNA or RNA, both of which will result in cell death (for review on colicin biology, see Cascales et al., 2007; Riley and Wertz, 2002). Colicin expression is tightly regulated by the SOS response (Gillor et al., 2008). In neutral environments, the dimerized repressor LexA inactivates the SOS regulon by binding to a DNA sequence called the LexA box. Upon DNA damage, RecA activates LexA self-cleavage allowing gene expression of >80 genes under RecA/LexA control (Fernandez De Henestrosa et al., 2000; Courcelle et al., 2001).
Here we describe a new colicin detectable only within biofilms formed by natural E. coli ROAR029 and that was identified while screening for molecules inhibiting E. coli adhesion and growth (Rendueles et al., 2011). We demonstrate that ROAR029 colicin (hereafter referred to as colicin R) is a pore-forming, plasmid-encoded colicin the production of which is not detected under planktonic conditions. We show that the increased SOS stress response induced in mature biofilms leads to induction of colicin R expression and to local concentration in biofilms. We demonstrate that lipopolysaccharide (LPS) integrity is a determinant for resistance, as colicin R is mainly active toward bacteria producing short LPS. Finally, we show that release of colicin R confers a strong competitive advantage in mixed biofilms and that colicin R toxicity toward the enteroaggregative E. coli LF82 clinical isolate is increased under biofilm conditions compared with planktonic susceptibility. This study therefore identifies the first biofilm-associated colicin targeting biofilm bacteria.
Materials and methods
Bacterial strains, plasmids and liquid growth conditions
E. coli isolates used in this study originated from published collections (Escobar-Paramo et al., 2006; Skurnik et al., 2006) and were compiled in Rendueles et al. (2011). Bacterial strains and plasmids used are listed in Supplementary Tables S1 and S2. Bacteria were cultured in 0.4% glucose M63B1 minimal medium (M63B1Glc) or in LB lysogeny broth medium at 37 °C. Antibiotics were used as follows: ampicillin, 100 μg ml−1; apramycin, 30 μg ml−1; chloramphenicol, 25 μg ml−1; spectinomycin, 50 μg ml−1; zeocin, 50 μg ml−1; apramycin, 30 μg ml−1; and kanamycin, 50 μg ml−1.
Purification of planktonic and biofilm extracts
Planktonic supernatants
Bacteria were cultured for 24 h in M63B1Glc and adjusted to an optical density (OD600 nm) of 3. Cultures were centrifuged for 10 min at 7500 r.p.m. at 4 °C, and supernatants were filtered through 0.2-μm filters.
Biofilm extracts
Sixty-milliliter microfermentors containing a removable glass stand were configured as continuous-flow culture bioreactors with a 40-ml h−1 flow rate, which maximizes biofilm development and minimizes planktonic growth of the bacteria (see http://www.pasteur.fr/recherche/unites/Ggb/biofilmfermenter.html for more detailed information; Ghigo, 2001; Rendueles et al., 2011). Inoculation was performed by dipping the glass spatula for 5 min in a culture adjusted to an OD600 nm of 1 from overnight bacterial cultures. The spatula was then reintroduced into the microfermentor. After 72 h at 37 °C, 45 ml of microfermentor medium (OD600 nm<0.1) was carefully discarded, and biofilm biomass was rapidly resuspended in the remaining 15 ml medium by vortexing for 30 s. The final biofilm biomass was measured, adjusted to an OD600 nm of 3, centrifuged and filtered through 0.22-μm filters (this biofilm conditioned media is hereafter referred to as biofilm extract). Alternatively, microtiter plates were inoculated with overnight diluted cultures (1:100) and grown for 24–48 h at 37 °C under static conditions. E. coli ROAR029 microtiter plate biofilms were resuspended in their own supernatant and directly spotted onto agar plates adequately supplemented with antibiotics to inhibit further growth of ROAR029.
Biofilm-related assays
Biofilm inhibition assays
Overnight cultures were diluted in M63B1Glc, inoculated into 96-well polyvinyl chloride (PVC) plates and filter-sterilized biofilm extract or planktonic supernatant was added at a 1:1 ratio. Biofilms were grown for 24 h at 37 °C before quantification.
Biofilm dispersion assays
Static biofilms were grown in 96-well PVC microtiter plates for 48 h at 37 °C. Free cells and used media were removed by inversion of the microtiter plate, biofilm was washed using M63B1 and 100 μl of ROAR029 biofilm extract were added to each biofilm for 2 h at 37 °C before quantification.
Biofilm quantification
Unbound cells were removed by inversion of the microtiter plate; biofilms were washed with water and adhering cells were then stained with 1% crystal violet for 20 min. Excess stain was removed by washing with deionized water three times. Quantification of attached biofilms was performed by addition of acetone–ethanol (20:80), and the dissolved crystal violet was measured at an OD of 595 nm.
Colicin susceptibility assays on planktonic and biofilm cells
Planktonic
Bacterial cultures were grown around 6 or 16 h (exponential and stationary, respectively) in M63B1Glc. Bacteria were then harvested by centrifugation and resuspended in the same volume of M63B1Glc (control planktonic) or in a 1:1 mix of M63B1Glc and ROAR029 biofilm extract to obtain a final cell quantity of 1.108 (1.109 colony-forming unit (CFU) ml−1). One hundred microliters were then transferred to microtiter plate wells. After 2 h of incubation at 37 °C, serial dilution plating (CFU counts) was used to assess colicin susceptibility of the treated planktonic population versus a planktonic exponential and stationary untreated controls.
Biofilm
Biofilms were formed for 48 h in microtiter plates as described above. Unattached and planktonic bacteria were removed by inversion of the microtiter plates and gentle tapping. One hundred microliters of either a 1:1 mix of M63B1Glc and ROAR029 biofilm extract or M63B1Glc (control) were added to each well in which the preformed biofilm (ca. 1.108 CFU, 1.109 CFU ml−1) had developed. The treated biofilms were incubated for 2 h at 37 °C and then resuspended by vigorous pipetting. Cell survival was determined by CFU count.
Biofilm resuspended cells
Biofilms were formed for 48 h in microtiter plates as described above. Unattached and planktonic bacteria were removed, and biofilms were washed by adding (without resuspending) and removing M63B1. Then the biofilms were resuspended in 100 μl of either a 1:1 mix of M63B1Glc and ROAR029 biofilm extract or M63B1Glc (control). Cells were then incubated at 37 °C for 2 h, and bacterial survival was monitored as above.
Growth inhibition test
The inhibitory effect of filter-sterilized supernatants and biofilm extracts was evaluated by spotting 8 μl on an overlay of targeted cells on appropriate media. Plates were incubated overnight at 37 °C. Batch growth in microtiter plates was also monitored by diluting overnight cultures to a final OD600 nm of 0.1 and mixed at a 1:1 ratio with cell-free biofilm extract and grown in microtiter plates for 24 h at 37 °C.
Generation of deletion mutants
Gene deletions in the different strains of E. coli were generated by the λ-red linear DNA gene replacement system (Chaveroche et al., 2000; Derbise et al., 2003) or by P1vir transduction from the Keio collection to E. coli MG1655 when indicated (Supplementary Table S2; Baba et al., 2006). Primers used are listed in Supplementary Table S3. ROAR029 constructs were checked by PCR and sequencing. Construction of ROAR029_KmRExTET_LexA in which lexA is under the control of an anhydrotetracycline-inducible promoter was performed as described in Da Re et al. (2007).
Construction of an LPS reconstituted mutant in MG1655
Construction of O-antigen-reconstituted E. coli K-12 strains was performed using O-antigen trans-complementation between two related E. coli K-12 backgrounds, EMG2 and WG1 (Liu and Reeves, 1994). Both backgrounds are O-antigen deficient, either by IS (insertion sequence) insertion in wbbL for EMG2 (rfb-50) or by a deletion encompassing the end of rfbB for WG1 (rfb-51). The KmFRT selectable genetic cassette from the pKD4 plasmid (Datsenko and Wanner, 2000) was inserted between the gnd and intact wbbL genes of WG1 strain using long primers (wbbL.KmFRT.long.L-5 and wbbL.KmFRT.long.L-3, Supplementary Table S3) and λ-red recombination. Primers hybridizing 200 base pairs before and after wbbL (wbbL200.ext-5 and wbbL200.ext-3 Supplementary Table S3) were then used to amplify the KmFRT-wbbL region that was used to replace the wbbL-deficient region of EMG2 derivatives MG1655 and TG1 strains. Restoration of functional wbbL gene and O-antigen production was verified by sequencing and LPS visualization on gel (Figure 5a).
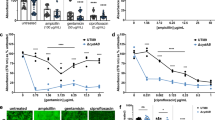
ROAR029 colicin activity is dependent on LPS length. (a) SDS-polyacrylamide (14%) gel showing LPS reconstruction of rough E. coli MG1655 and E. coli TG1 strains. (b) Effect of ROAR029 biofilm extract on bacterial lawns of rough (MG1655 and TG1) and smooth (MG1655 wbbL+ and TG1 wbbL+) strains. (c) SDS-polyacrylamide gel (14%) of E. coli 536 wild type and LPS (rfaH, manB, waaG and waaC), capsule (kps15) and colanic acid (cpsG) mutants. (d) Effect of ROAR029 biofilm extract on E. coli 536 wild type and LPS mutants lawns. (e) Growth curves showing the effect of ROAR029 biofilm extract addition to cultures of E. coli 536 and mutants with different LPS lengths. The graph shows means of eight independent wells from a representative experiment. ΔrfaHc, complemented rfaH mutant.
Random mutagenesis
The ROAR029 genome
Transposon mutagenesis of ROAR029 was performed using random insertion of the EZ-Tn5<KAN-2>Transposome (Epicentre Biotechnologies, Madison, WI, USA). A total of 5000 clones were analyzed.
ROAR029 plasmid
Plasmid pColR was mutagenized by EZ-Tn5<KAN-2>transposition. Pools of mutagenized plasmids were electroporated into E. coli TG1 electrocompetent cells. In EZ-Tn5<KAN-2>, the kanamycin gene is not followed by a transcription terminator, allowing possible transcriptional read-through.
Mixed biofilm assays
ROAR029 bearing pColR_Km, pColR− or pColR+ and MG1655F’ cells were grown overnight in M63B1Glc, washed twice in M63B1 to remove traces of colicin R (if any) and diluted to OD600 nm 0.1. Cells were mixed at different initial ratios of ROAR029 derivatives to MG1655F’ 1:1, 5.106 CFU: 5.106 CFU; 1:10, 5.105 CFU: 5.106 CFU; 1:100, 5.104 CFU: 5.106 CFU and 1:10 000, 5.102 CFU:5.106 CFU and inoculated into 96-well PVC plates for 24 h at 37 °C. We counted the proportion of each strain in the 24-h-grown biofilms after washing, resuspension and serial dilution plating in appropriate antibiotics.
Sodium dodecyl sulfate–polyacrylamide gel for LPS observation
Cell preparation
Cells were harvested by centrifugation, resuspended in deionized water and normalized to a final OD600 nm of 1. Samples were then heated for 10 min at 100 °C and treated with proteinase K (2 mg ml−1) for 1 h at 37 °C.
LPS gel and staining
Sodium dodecyl sulfate–polyacrylamide gels were made according to Lugtenberg et al. (1975). Gels were run at 75 mA for 60–90 min. LPS were stained with silver nitrate as described previously (Tsai and Frasch, 1982).
Sequence of ROAR029 colicin plasmid, colicin R sequence and structure analysis
The sequence of the pColR plasmid was deposited in Genbank (accession number KF137578).
Sequence alignments
Alignments were performed by MultiAlin software as described in Corpet (1988).
Phylogenetic dendrogram
Colicin sequences were directly obtained from GenBank (http://www.ncbi.nlm.nih.gov/gene). Sequences were aligned following ClustalW standards and further examined in order to create phylogenetic dendrograms with MEGA 5 software developed by Tamura et al. (2011).
Plasmid map
pColR plasmid map was constructed using the SnapGene viewer from GSL Biotech LLC (Chicago, IL, USA).
Colicin tertiary structure
Colicin R structure was simulated using the PHYRE software (Imperial College, London, UK; Bennett-Lovsey et al., 2008).
Statistical methods
Statistical analyses were performed using unpaired t-test with Prism v5.0 (GraphPad Software, San Diego, CA, USA).
Results
Commensal E. coli ROAR029 produces a biofilm-associated bacteriotoxic molecule
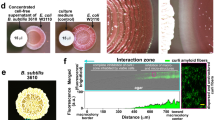
We had previously screened a collection of 122 E. coli natural isolates from different phylogenetic groups, hosts and origins for their ability to produce biofilm-specific molecules inhibiting biofilm formation of other competing strains (Rendueles et al., 2011). That study revealed that biofilm extracts from commensal E. coli ROAR029 isolated from an impala antelope (Aepyceros melampus) in Gabon (phylogenetic group D) inhibited biofilm formation and planktonic growth of the biofilm-forming MG1655F’tetΔtraD strain (referred to as MG1655F’; Valle et al., 2006; Rendueles et al., 2011; Figure 1a). Although filter-sterilized planktonic supernatant of E. coli ROAR029 did not show biocidal activity, a filter-sterilized extract from ROAR029 biofilms produced a growth inhibition halo when spotted on an MG1655F’ lawn (Figure 1b). Production of the bacteriotoxic compound was also detected in microtiter plates and colony biofilm models (Anderl et al., 2000; Figure 1c and data not shown). This biofilm-associated bacteriotoxic effect could not be attributed to the previously reported presence of valine in E. coli biofilms, as ROAR029 biofilm extract retained activity on the MG1655 ilvG+ strain in which the corrected ilvG frameshift confers resistance to valine (Figure 1d; Valle et al., 2006). Finally, determination of colony-forming units of a MG1655F’ planktonic culture upon exposure to ROAR029 biofilm extract in a 1:1 ratio for 2 h showed no surviving bacteria, indicating bacteriotoxic rather than bacteriostatic activity of the ROAR029 biofilm extract (data not shown).
Bacteriotoxic effect of ROAR029 extracts upon E. coli MG1655F’. (a) Quantification of the effects of ROAR029 planktonic (ROAR029 Pk) and biofilm (ROAR029 Bf) extracts on biofilm formation and growth of MG1655F’. M63B1, control. ***P<0.001. (b) Growth inhibition of ROAR029 planktonic (ROAR029 Pk) and biofilm extracts (ROAR029 Bf) on a lawn of E. coli MG1655. (c) Growth inhibition of ROAR029 colony on a lawn of E. coli MG1655. (d) Effects of ROAR029 biofilm extract (ROAR029 Bf) on a lawn of valine-resistant E. coli MG1655 ilvG+.
ROAR029 biofilm-associated bacteriotoxic molecule is a new pore-forming colicin
To identify the genetic determinants of ROAR029 bacteriotoxic activity, we performed random transposon mutagenesis and searched for ROAR029 mutants unable to produce active biofilm extract. We identified one transposon mutant inserted in pcnB, a gene coding for a poly-A polymerase involved in mRNA poly-A adenylation and plasmid copy number regulation (Liu and Parkinson, 1989). Introduction of a pcnB deletion into ROAR029 abolished the bacteriotoxic activity of microtiter-plate- or microfermentor-grown biofilm extracts (Figure 2a). As ROAR029 ΔpcnB did not show significant growth or biofilm defects (Supplementary Figure S1A), we hypothesized that the genetic determinants of the ROAR029 bacteriotoxin could be carried by a plasmid; indeed, we detected the presence of a ca. 5.8 kb plasmid in ROAR029 (Figure 2b). We subjected the ROAR029 plasmid to random in vitro mutagenesis and electroporated pools of mutagenized plasmids into E. coli K-12 strain TG1. We found that 67% of kanamycin-resistant transformants produced active bacteriotoxic compound in microtiter plate biofilm extracts, demonstrating that this plasmid carries genes involved in synthesis of ROAR029 bacteriotoxin. The remaining 33% kanamycin-resistant TG1 transformants did not produce active bacteriotoxic compounds. Sequence analysis showed that inactive mutagenized plasmids carried transposon insertions in DNA regions homologous to pore-forming colicins U- and Y-encoding genes, including genes potentially coding for a 69.3-kDa colicin, as well as lysis and immunity proteins (Table 1, Figures 2c and d, Supplementary Figures S2, S3 and S4; Smajs et al., 1997; Riley et al., 2000). We further tested whether the immunity encoded by ROAR029 plasmid conferred resistance to other colicins, and indeed, it diminished the killing of colicin U and completely blocked growth inhibition of colicin Y (Supplementary Figure S4A) but had no effect against unrelated colicin E2. Sequence and phylogenetic analyses further showed that while immunity genes of pore-forming colicins U, Y and ROAR029 cluster together, key aminoacids for interaction between the immunity protein and its colicin partner (Smajs et al., 2006) appeared more similar between ROAR029 and colicin Y than with colicin U (Supplementary Figure S4BC).
ROAR029 bacteriotoxic activity is due to plasmid-encoded colicin regulated by PcnB. (a) Addition of an 8-μl drop of ROAR029 wild type, ROAR029ΔpcnB or ROAR029 bearing a mutagenized plasmid unable to synthesize the bacteriotoxic molecule (ROAR029 pColR−) biofilm extract to a lawn of MG1655. (b) DNA gel of ROAR029 pColR plasmid linearized by the EcoRV enzyme. (c) ROAR029 pColR plasmid map. Arrows indicate transcription polarities of colicin genes. Unique-site restriction enzymes are indicated. (d) Dendrogram of sequences alignment of the 200 last amino acids from colicin-encoding genes of the C-terminal region. Phylogenetic analysis was carried out using the UPGMA method. Numbers represent bootstrap values >75.
We then replaced the original marker-less ROAR029 colicin-encoding plasmid (pColR), hereafter referred to as colicin R for ROAR029, with an inactive kanamycin-resistant plasmid by transformation (pColR−). Plasmid maintenance upon antibiotic selection led to exclusion of the native plasmid. Biofilm extracts of ROAR029 strains with inactive kanamycin-resistant plasmid did not display bacteriotoxic activity (Figure 2a). Additionally, one mutagenized plasmid seemed to confer an increased killing effect, referred to as pColR+ (Supplementary Figure S5).
To further confirm the colicinogenic nature of the antagonistic molecule, we checked that TG1 transformed with pColR was resistant to biofilm extracts produced by pColR and pColR+ strains by virtue of the immunity protein carried in the plasmid (Supplementary Figure S6).
Taken together, our results demonstrated that ROAR029 biofilm extracts’ bacteriotoxic activity was associated with production of colicin R encoded by the 5.8-kb plasmid pColR.
Colicin R is a group A Tol-dependent colicin
Colicins are classified into groups A and B based on the use of either Tol or Ton translocation systems, respectively (Cascales et al., 2007). Although E. coli MG1655 Ton system mutants ΔtonB, ΔexbB and ΔexbD were sensitive to colicin R, a ΔtolA mutant displayed full resistance to colicin R as it did for colicin U (Figure 3). Consistently, we identified a conserved TolA box consensus motif (DGTGW), as well as several repeated TolA motifs in the colicin R protein sequence, involved in colicin internalization (data not shown). Other Tol mutants (ΔtolQ, ΔtolR, ΔtolC and ΔtolB) showed various levels of tolerance to colicin R (Supplementary Figure S7). Residual activity of colicin R in other Tol mutants might be explained by partial phenotype complementation of Ton proteins. Indeed, E. coli mutant strains carrying double ton and tol deletions displayed full resistance to colicin R (data not shown). Finally, we showed that like for colicin U, an ompA mutation in E. coli MG1655 led to colicin R resistance (Figure 3 and Supplementary Figure S7). Interestingly, closely related colicins U and Y both used OmpF as a translocator (data not shown). However, colicin R activity remained unaffected in an ompF mutant (Supplementary Figure S7) as well as in an ompC mutant (data not shown). Hence, we cannot exclude the possibility that OmpA acts as a colicin R receptor, translocator or both. Accordingly, E. coli TG1 transformed with pColR inhibited E. coli MG1655 but not ompA and tolA mutants (data not shown).
Biofilm specificity of colicin R might be due to SOS regulatory activity
Colicins have been shown to be tightly regulated by the SOS system (Gillor et al., 2008). Consistently, we identified two overlapping LexA boxes in the promoter region located upstream of the ROAR029 colicin gene (Figure 4a). LexA is a repressor of SOS-regulated genes cleaved by RecA upon induction of the SOS response. We observed that overexpression of LexA correlated with decreased levels of colicin in E. coli ROAR029 biofilm extracts (Figure 4b), and addition of mitomycin C to either biofilm extracts or inactive E. coli ROAR029 planktonic cultures led to derepression of colicin R production (Figure 4c and data not shown, respectively).
ROAR029 biofilm-specific production of colicin R is due to an SOS regulatory effect. (a) Sequence analysis revealed two overlapping SOS boxes upstream of the ribosome binding sequence (rbs) and start codon ATG of ROAR029. Insertion site of transposon in pColR+ plasmid is indicated. Transcriptional read-through from kanamycin gene and disruption of LexA box1 binding site may explain constitutive expression of colicin R encoding gene. (b) Effect of LexA induction upon ROAR029 bacteriotoxic activity was assessed using ROAR029 with LexA under the control of an anhydrotetracycline-inducible (aTc) promoter. An 8-μl drop of ROAR029_KmRExTET_LexA treated with increasing concentrations of aTc ranging from 0 to 1000 ng ml−1 was added to a lawn of MG1655. (c) Inhibition halo of E. coli MG1655 when treated with an 8-μl drop of ROAR029 biofilm extract (ROAR029 Bf) with and without induction of the SOS system by 0.1 μg ml−1 of mitomycin C added after inoculation. (d) Growth inhibition of E. coli MG1655 by biofilm extracts produced by E. coli TG1 (black) and TG1 lexAind3 (gray) transformed by different ROAR029 plasmids; active kanamycin-resistant pColR_Km, defective colicin mutant (pColR-, dashed) and constitutively expressed colicin (pColR+, shaded).
We had previously shown that the SOS system is upregulated within E. coli biofilms (Beloin et al., 2004; Bernier et al., 2013). Thus, colicin R biofilm-specific production could be directly linked to an increased SOS response. To investigate this, we used three different plasmids derived from wild-type pColR: (i) pColR−, unable to produce colicin R due to transposon-based interruption of the colicin gene, (ii) pColR+ in which colicin gene transcription is constitutive due to a probable combination of read-through from the promoter of the kanamycin gene by the EZ-Tn5<KAN-2>transposon and dirsruption of LexA binding to LexA box 1 caused by transposon insertion (Figure 4a), and (iii) pColR_Km, active kanamycin-resistant pColR (Supplementary Figure S5). These plasmids were transformed into E. coli TG1 and TG1lexAind3, the latter unable to induce an SOS response due to a non-cleavable version of the LexA protein. As expected, biofilm extract from strains carrying pColR− did not show growth-inhibiting activity, whereas biofilm extracts from strains bearing pColR+ displayed strong SOS-independent growth-inhibiting activity (Figure 4d). In contrast, the growth-inhibiting activity of strains carrying the pColR_Km plasmid was totally dependent on a functional LexA protein (Figure 4d). Taken together, these results suggest that increased release of colicin R is a consequence of induction of the SOS response within biofilms.
Colicin R is only active toward E. coli strains with short LPS
To investigate the activity spectrum of colicin R, we tested a panel of pathogenic and commensal E. coli as well as other Gram-negative strains for their sensitivity to ROAR029 biofilm extract (Supplementary Table S1). We observed that all tested strains were resistant to colicin R except for E. coli K-12 EMG2 and B derivatives and adherent-invasive E. coli LF82. E. coli K-12 EMG2 derivatives are characterized by an insH-7 insertion element in wbbL resulting in rough LPS (Liu and Reeves, 1994). Reintroduction of a wild-type wbbL allele in E. coli K-12 MG1655 (MG1655 wbbL+) and E. coli TG1 (TG1 wbbL+) restored O-antigen production (Figure 5a) and led to total resistance to colicin R in bacterial lawns, batch cultures and microtiter plate biofilms (Figure 5b and Supplementary Figure S8). Furthermore, E. coli LF82 naturally lacks O-antigen due to a deletion of wbbLKIJH_glf_rfbX_rfbC genes (Miquel et al., 2010; Supplementary Figure S9), which probably explains its colicin R sensitivity. We also tested whether colicins closely related to colicin R, that is, colicins U and Y, or another colicin, colicin E2, had bacteriotoxic effects against E. coli K-12 MG1655 and E. coli MG1655 wbbL+; we observed that these three colicins were also inactive against the O-antigen LPS-restored MG1655 wbbL+strain (data not shown). Colicins U, Y and E2 as well as colicin R produced in TG1_pColR were also active against LF82 strain (data not shown).
We further investigated the influence of LPS length on colicin R activity using E. coli 536, a uropathogenic strain resistant to colicin R and its ΔrfaH, ΔmanB, ΔwaaG and ΔwaaC mutants displaying various LPS sizes (Beloin et al., 2006; Figure 5c). Addition of ROAR029 biofilm extract to E. coli 536 mutants revealed that resistance increased with LPS size (Figures 5d and e) and that restoration of whole LPS in a complemented rfaH mutant (rfaHc) led to total resistance to colicin R (Figures 5d and e). Interestingly, uropathogenic E. coli CFT073 and 536 mutants unable to synthesize other extracellular polymeric surface structures like colanic acid and capsules remained fully resistant to colicin R (Figures 5d and e and data not shown). Taken together, these results showed that a reduction in LPS size correlated with increased sensitivity to colicin R.
ROAR029 colicin excludes other E. coli from mixed biofilms
To determine whether colicin R production confers a competitive advantage to ROAR029 within structured communities, we compared the ability of ROAR029 carrying pColR_Km, pColR− or pColR+ plasmids to grow in mixed biofilms in competition with the biofilm-forming colicin-R-sensitive E. coli MG1655F’ strain (Ghigo, 2001). It was verified that ROAR029 strains expressing differing levels of colicin R did not display significantly different growth rates in liquid- and biofilm-forming abilities (Supplementary Figure S10). ROAR029 derivatives and MG1655F’ cultures were adjusted to different initial ratios and grown as mixed biofilms for 24 h. Although ROAR029 pColR− was naturally able to outcompete MG1655F’ due to a higher growth rate of ROAR029 derivatives (see Supplementary Figure S10), bacterial survival showed that ROAR029 pColR_Km and ROAR029 pColR+, colicin-producing strains, more efficiently outcompeted MG1655F’ (Figure 6). In order to further confirm the prominent role of colicin R in the outcompetition within biofilm, transformed TG1 with pColR plasmids was competed against a streptomycin-resistant TG1 at different initial ratios. Accordingly, TG1 pColR outcompeted TG1. This effect was increased when TG1 carried pColR+ plasmid and abolished when pColR− was used (Supplementary Figure S11).
We further characterized temporal dynamics of ROAR029/MG1655F’ interactions by inoculating the two strains at a 1:1 ratio, frequencies at which bacteriocin production provides the greatest advantage (Inglis et al., 2009). To avoid potential accumulation of colicin R in the medium, we used continuous flow microfermentors and observed a rapid reduction in MG1655F’ and complete outcompetition after 8 h (Supplementary Figure S12). This strong outcompetition was not due to higher ability of ROAR029 pColR_Km to initially adhere to glass spatula since, on the contrary, the initial adhesion of MG1655F’, probably by virtue of the production of the strong F pilus adhesion factor (Ghigo, 2001), was ∼20-fold higher than the one of ROAR029 pColR_Km (adhesion of ROAR029_pColR_Km and MG1655F’ are, respectively, 0.10±0.037% and 2.22±0.76%, P=0.00838, unpaired t-test). These results demonstrated that colicin R production and higher growth rate provide a strong competitive advantage to ROAR029 in mixed biofilms. This suggests that production of colicin R could have a role in population dynamics in natural situations.
Colicin R is more active against biofilms than planktonic cultures
The specific release of colicin R within E. coli ROAR029 biofilms led us to hypothesize that its production could correlate with increased sensitivity to colicin R in E. coli biofilms. To test this, we compared sensitivity to colicin R of planktonic and biofilm bacteria for two different biofilm-forming pathogens with smooth (full length) and rough (lacking O-antigen) LPS, E. coli 55989 and E. coli LF82, respectively (Supplementary Figure S9A). Whereas E. coli 55989 in planktonic and biofilm conditions remained unaffected by colicin R, LF82 biofilms displayed increased sensitivity compared to LF82 planktonic cultures (Figure 7). This increased sensitivity was not due to further reduction of LPS length during biofilm growth, as LF82 planktonic and biofilm LPS profiles were identical (Supplementary Figure S9A). In addition, we tested whether resistance to colicin R could be enhanced by the characteristic 3D structure and architecture displayed by biofilm cells by assessing the sensitivity of biofilm bacteria resuspended in liquid medium, maintaining the physiological specificities of biofilm bacteria but not their characteristic structured environment. Resuspended LF82 biofilm bacteria were more sensitive to colicin R planktonic bacteria and marginally more sensitive than biofilms (Figure 7), indicating that colicin R preferentially targets biofilm cells of susceptible bacteria.
Colicin R has enhanced activity towards biofilms of LF82. Differential effect of ROAR029 biofilm extract on planktonic (exponential and stationary phase) and biofilm cells (resuspended and non-resuspended) of two pathogenic E. coli strains, 55989 with full LPS and LF82 with no LPS. Survival was determined by CFU counts after 2-h exposure to equal quantities of ROAR029 biofilm extract containing colicin R. ***P<0.001.
Discussion
In the present study, we characterized an unknown antagonistic antimicrobial molecule released within biofilms formed by E. coli natural isolate ROAR029. We demonstrated that this biofilm-associated molecule corresponds to a new 5.8-kb group I plasmid-encoded colicin (colicin R) exhibiting group A colicin characteristics (Cascales et al., 2007). Phylogenetic analysis of colicin-associated genes of ROAR029 showed that they cluster with two other pore-forming colicins, U and Y, isolated from Shigella boydii, and a natural E. coli isolate from the Amazonian region, respectively (Smajs et al., 1997; Riley et al., 2000). Additionally, colicin R immunity protein confers full cross-immunity to colicin Y and partial immunity to colicin U but no immunity to unrelated colicin E2.
Biofilm-specificity of colicin R production
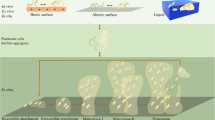
Here we demonstrate that induction of the SOS response, previously shown to be upregulated in E. coli biofilms (Beloin et al., 2004; Bernier et al., 2013), is directly correlated with increased production of colicin R in biofilms. Other reports have highlighted the fact that colicins are also upregulated in late stationary phase independently of the SOS system and are triggered by nutrient depletion in the medium (Eraso et al., 1996; Kuhar and Zgur-Bertok, 1999), a common situation in the deep layers of biofilms (Boles and Singh, 2008; Stewart and Franklin, 2008). Recently, Cornforth and Foster (2013) proposed that bacteriocin production is correlated with the stress response as a potential reaction to the presence of ecological competitors and probably has an important role in ecological interference within structured bacterial populations. In addition, several studies postulate that optimal efficiency of bacteriocins is achieved within structured environments, such as biofilms (Chao and Levin, 1981; Durrett and Levin, 1997; Gardner et al., 2004). In agreement with previous studies, we show that in mixed biofilm conditions colicin R confers a competitive advantage against closely related direct-niche competitors consuming similar potentially limited resources.
In addition, enhanced colicin R production within biofilms could correspond to a specific stress-induced defense mechanism, as release of group A colicins is a lethal event for the cell (Cascales et al., 2007). Induction of the SOS system in biofilms upon stresses such as waste product accumulation, microaerobic areas or increased oxidative stress could lead to colicin R release and subsequent cell death. Hence, biofilm-associated colicin production could have several benefits, including eliminating the most stressed bacteria from biofilms without negatively impacting growth or subsequent biofilm development of the overall population (Supplementary Figure S10). Other suicidal mechanisms have been reported in structured populations, including phage release (Carrolo et al., 2010), nitric oxide production (Barraud et al., 2009) and cannibalism toward Bacillus subtilis cells (Lopez et al., 2009).
Moreover, biofilm-specific colicin production could contribute to enabling kind discrimination mechanisms (Strassmann et al., 2011). Biofilms rely on cooperative acts such as matrix production and communication signals that can be exploited by other non-cooperative individuals (‘cheaters’) who benefit from them but do not share the associated cost. When uncontrolled, these cheaters can increase in frequency, undermine cooperation and even cause population crashes in situations where cooperation is important for survival (Hamilton, 1964). In this context, biofilm-specific colicin production could ensure close relatedness between interacting bacteria and promote cooperation by excluding possible cheaters and reducing social exploitation (kin selection) (Hamilton, 1964; Nadell et al., 2009).
Colicin R targets LPS-deficient and preferentially biofilm cells
Like previously described colicins, ROAR029 colicin has a narrow spectrum of action (Cascales et al., 2007). Here we demonstrate that in addition to receptor and translocator proteins on the surface of target bacteria, LPS may have an important role in determining sensitivity towards colicins. We show that E. coli strains with full-length LPS are resistant to colicin R, whereas strains without O-antigen display increased susceptibility. Interestingly, restoration of O-antigen led to resistance to two pore-forming colicins (U and Y) and to RNAse colicin E2 (data not shown), suggesting that intact O-antigen might be a general determinant of resistance to colicins. Although the presence of O-antigen has been previously suspected to alter the effect of temperature or EDTA on colicin susceptibility of target cells by means of the change of receptor activity (Gado et al., 1995), we postulate that colicin R resistance of full-length LPS strains could be due to charge or steric repulsion by O-antigen, therefore preventing colicin R interaction with its receptor at the cell surface. This phenomenon is consistent with the observation that purified LPS fractions do not seem to interact directly with colicins (Weltzien and Jesaitis, 1971; Law et al., 2003). In the case of colicin R, this potential steric or charge repulsion seems specific to O-antigen, as deletion of other large surface molecules such as colanic acid and capsules did not enhance susceptibility to colicin R of uropathogenic E. coli strains 536 and CFT073.
Additionally, we show that a reduction in the LPS lipid A core size was correlated with E. coli increased sensitivity to colicin R, probably because of an increased accessibility of colicin R to its receptor. This is in contrast to increased resistance to colicin U displayed by E. coli mutants with a shorter lipid A LPS core (Smajs et al., 1997) or cellular resistance of deep rough mutants to colicin N (Sharma et al., 2009), suggesting that lipid A core could have an important role in early interaction with certain colicins as shown for colicin E2 (Osada and Beppu, 1985). Hence, colicins U, E2 and R are all inactive when O-antigen is present. However, colicins U and E2 need an intact lipid A outer core for binding to the outer membrane and subsequent insertion, whereas colicin R efficacy is reduced by presence of lipid A core.
Furthermore, we provide evidence that strains with naturally reduced LPS length display increased sensitivity to colicin R when grown as biofilms. In the clinical enteroaggregative E. coli LF82 strain, colicin R indeed killed biofilm cells more efficiently than planktonic cells. Despite this increased activity toward biofilm cells, colicin R was unable to dissolve biofilms (data not shown), suggesting that it can penetrate biofilms, but that the 3D structure of biofilms is conserved after the colicin R effect. However, biofilm 3D architecture and the presence of extracellular matrix slightly contributed to protecting E. coli LF82 biofilm from the effects of colicin R (Figure 7). Although the specific mechanism by which LF82 biofilm bacteria display a decreased survival ratio compared with planktonic cells remains to be elucidated, it does not seem to be linked to further LPS modifications. Indeed, LPS extracted from biofilms and planktonic E. coli LF82 appeared to be identical. Alternatively, the particular metabolic and physiological properties displayed by biofilm bacteria may account for their increased sensitivity. For instance, a reduced growth rate resulting in decreased outer membrane component turnover or increased stress in biofilm layers or niches could lead to enhanced sensitivity to colicin R. As biofilm bacteria typically display higher tolerance to antimicrobial agents (Hoiby et al., 2010), this is, to our knowledge, the first example of biofilm bacteria being more sensitive to a bacteriotoxic compound than their planktonic counterparts.
Counterselection of LPS rough E. coli cells: ecological implications in species diversity
Production of antagonistic molecules has been proposed to have an important role in selection and evolution of strains (Riley and Gordon, 1999; Kirkup and Riley, 2004; Majeed et al., 2011). Indeed, colicin production promotes microbial diversity, both theoretically (Czaran et al., 2002) and empirically (Walker et al., 2004). Here we demonstrated that colicin R can preferentially target strains with reduced LPS (absence of O-antigen and reduced lipid A core) probably by an increased ability in these strains to access its receptor. This suggests that in mixed biofilm communities selective pressure exerted by colicin R producers may favor bacteria with long LPS structures. In addition, it has been recently been shown that electrostatic interactions between LPS and other polysaccharides enhances and stabilizes biofilms (Amini et al., 2009) and that LPS have a key role in mixed biofilm dynamics (Bandara et al., 2010). Taken together, colicin R could therefore directly impact population evolution. Interestingly, in P. aeruginosa and P. fluorescens biofilms reduction of LPS O-antigen has been reported (Giwercman et al., 1992; Spiers and Rainey, 2005; Ciornei et al., 2010), suggesting that such LPS modifications could also occur in E. coli biofilms, increasing relevance of producing a colicin against strains with short LPS.
In conclusion, our study contributes to the role played by colicins in representative multispecies contexts. It further establishes biofilms as a source of bioactive antagonistic molecules, as well as being a crucial site for ecological interactions that influence E. coli evolution and population dynamics in complex and relevant multispecies environments.
Accession codes
References
Amini S, Goodarzi H, Tavazoie S . (2009). Genetic dissection of an exogenously induced biofilm in laboratory and clinical isolates of E. coli. PLoS Pathog 5: e1000432.
Anderl JN, Franklin MJ, Stewart PS . (2000). Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob Agents Chemother 44: 1818–1824.
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M et al. (2006). Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2: 2006.0008.
Bandara HM, Lam OL, Watt RM, Jin LJ, Samaranayake LP . (2010). Bacterial lipopolysaccharides variably modulate in vitro biofilm formation of Candida species. J Med Microbiol 59: 1225–1234.
Barraud N, Schleheck D, Klebensberger J, Webb JS, Hassett DJ, Rice SA et al. (2009). Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J Bacteriol 191: 7333–7342.
Beloin C, Valle J, Latour-Lambert P, Faure P, Kzreminski M, Balestrino D et al. (2004). Global impact of mature biofilm lifestyle on Escherichia coli K-12 gene expression. Mol Microbiol 51: 659–674.
Beloin C, Michaelis K, Lindner K, Landini P, Hacker J, Ghigo JM et al. (2006). The transcriptional antiterminator RfaH represses biofilm formation in Escherichia coli. J Bacteriol 188: 1316–1331.
Bendaoud M, Vinogradov E, Balashova NV, Kadouri DE, Kachlany SC, Kaplan JB . (2011). Broad spectrum biofilm inhibition by Kingella kingae exopolysaccharide. J Bacteriol 193: 3879–3886.
Bennett-Lovsey RM, Herbert AD, Sternberg MJ, Kelley LA . (2008). Exploring the extremes of sequence/structure space with ensemble fold recognition in the program Phyre. Proteins 70: 611–625.
Bernier SP, Lebeaux D, DeFrancesco AS, Valomon A, Soubigou G, Coppee JY et al. (2013). Starvation, together with the SOS response, mediates high biofilm-specific tolerance to the fluoroquinolone ofloxacin. PLoS Genet 9: e1003144.
Boles BR, Singh PK . (2008). Endogenous oxidative stress produces diversity and adaptability in biofilm communities. Proc Natl Acad Sci USA 105: 12503–12508.
Bucci V, Nadell CD, Xavier JB . (2011). The evolution of bacteriocin production in bacterial biofilms. Am Nat 178: E162–E173.
Carrolo M, Frias MJ, Pinto FR, Melo-Cristino J, Ramirez M . (2010). Prophage spontaneous activation promotes DNA release enhancing biofilm formation in Streptococcus pneumoniae. PLoS One 5: e15678.
Cascales E, Buchanan SK, Duche D, Kleanthous C, Lloubes R, Postle K et al. (2007). Colicin biology. Microbiol Mol Biol Rev 71: 158–229.
Chao L, Levin BR . (1981). Structured habitats and the evolution of anticompetitor toxins in bacteria. Proc Natl Acad Sci USA 78: 6324–6328.
Chater KF . (2006). Streptomyces inside-out: a new perspective on the bacteria that provide us with antibiotics. Philos Trans R Soc Lond B Biol Sci 361: 761–768.
Chaveroche MK, Ghigo JM, d'Enfert C . (2000). A rapid method for efficient gene replacement in the filamentous fungus Aspergillus nidulans. Nucleic Acids Res 28: E97.
Ciornei CD, Novikov A, Beloin C, Fitting C, Caroff M, Ghigo JM et al. (2010). Biofilm-forming Pseudomonas aeruginosa bacteria undergo lipopolysaccharide structural modifications and induce enhanced inflammatory cytokine response in human monocytes. Innate Immun 16: 288–301.
Cornforth DM, Foster KR . (2013). Competition sensing: the social side of bacterial stress responses. Nat Rev Microbiol 11: 285–293.
Corpet F . (1988). Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16: 10881–10890.
Courcelle J, Khodursky A, Peter B, Brown PO, Hanawalt PC . (2001). Comparative gene expression profiles following UV exposure in wild-type and SOS-deficient Escherichia coli. Genetics 158: 41–64.
Czaran TL, Hoekstra RF, Pagie L . (2002). Chemical warfare between microbes promotes biodiversity. Proc Natl Acad Sci USA 99: 786–790.
Da Re S, Le Quere B, Ghigo JM, Beloin C . (2007). Tight modulation of Escherichia coli bacterial biofilm formation through controlled expression of adhesion factors. Appl Environ Microbiol 73: 3391–3403.
Datsenko K, Wanner B . (2000). One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci 97: 6640–6645.
Derbise A, Lesic B, Dacheux D, Ghigo JM, Carniel E . (2003). A rapid and simple method for inactivating chromosomal genes in Yersinia. FEMS Immunol Med Microbiol 38: 113–116.
Duquesne S, Petit V, Peduzzi J, Rebuffat S . (2007). Structural and functional diversity of microcins, gene-encoded antibacterial peptides from enterobacteria. J Mol Microbiol Biotechnol 13: 200–209.
Durrett R, Levin S . (1997). Allelopathy in spatially distributed populations. J Theor Biol 185: 165–171.
Eraso JM, Chidambaram M, Weinstock GM . (1996). Increased production of colicin E1 in stationary phase. J Bacteriol 178: 1928–1935.
Escobar-Paramo P, Le Menac'h A, Le Gall T, Amorin C, Gouriou S, Picard B et al. (2006). Identification of forces shaping the commensal Escherichia coli genetic structure by comparing animal and human isolates. Environ Microbiol 8: 1975–1984.
Fernandez De Henestrosa AR, Ogi T, Aoyagi S, Chafin D, Hayes JJ, Ohmori H et al. (2000). Identification of additional genes belonging to the LexA regulon in Escherichia coli. Mol Microbiol 35: 1560–1572.
Gado I, Paszti J, Laszlo VG . (1995). Effect of precultivation conditions on colicin susceptibility in Escherichia coli. J Chemother 7: 100–105.
Gardner A, West SA, Buckling A . (2004). Bacteriocins, spite and virulence. Proc Biol Sci 271: 1529–1535.
Ghigo JM . (2001). Natural conjugative plasmids induce bacterial biofilm development. Nature 412: 442–445.
Ghigo JM . (2003). Are there biofilm-specific physiological pathways beyond a reasonable doubt? Res Microbiol 154: 1–8.
Gillor O, Kirkup BC, Riley MA . (2004). Colicins and microcins: the next generation antimicrobials. Adv Appl Microbiol 54: 129–146.
Gillor O, Vriezen JA, Riley MA . (2008). The role of SOS boxes in enteric bacteriocin regulation. Microbiology 154: 1783–1792.
Giwercman B, Fomsgaard A, Mansa B, Hoiby N . (1992). Polyacrylamide gel electrophoresis analysis of lipopolysaccharide from Pseudomonas aeruginosa growing planktonically and as biofilm. FEMS Microbiol Immunol 4: 225–229.
Hamilton WD . (1964). The genetical evolution of social behaviour. I & II. J Theor Biol 7: 1–52.
Hibbing ME, Fuqua C, Parsek MR, Peterson SB . (2010). Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol 8: 15–25.
Hoiby N, Bjarnsholt T, Givskov M, Molin S, Ciofu O . (2010). Antibiotic resistance of bacterial biofilms. Int J Antimicrob Agents 35: 322–332.
Inglis RF, Gardner A, Cornelis P, Buckling A . (2009). Spite and virulence in the bacterium Pseudomonas aeruginosa. Proc Natl Acad Sci USA 106: 5703–5707.
James G, Beaudette L, JW. C . (1995). Interspecies bacterial interactions in biofilms. J Ind Microbiol Biotechnol 15: 257–262.
Kirkup BC, Riley MA . (2004). Antibiotic-mediated antagonism leads to a bacterial game of rock-paper-scissors in vivo. Nature 428: 412–414.
Kolodkin-Gal I, Romero D, Cao S, Clardy J, Kolter R, Losick R . (2010). D-amino acids trigger biofilm disassembly. Science 328: 627–629.
Kolodkin-Gal I, Cao S, Chai L, Bottcher T, Kolter R, Clardy J et al. (2012). A self-produced trigger for biofilm disassembly that targets exopolysaccharide. Cell 149: 684–692.
Kreth J, Merritt J, Bordador C, Shi W, Qi F . (2004). Transcriptional analysis of mutacin I (mutA) gene expression in planktonic and biofilm cells of Streptococcus mutans using fluorescent protein and glucuronidase reporters. Oral Microbiol Immunol 19: 252–256.
Kuhar I, Zgur-Bertok D . (1999). Transcription regulation of the colicin K cka gene reveals induction of colicin synthesis by differential responses to environmental signals. J Bacteriol 181: 7373–7380.
Law CJ, Penfold CN, Walker DC, Moore GR, James R, Kleanthous C . (2003). OmpF enhances the ability of BtuB to protect susceptible Escherichia coli cells from colicin E9 cytotoxicity. FEBS Lett 545: 127–132.
Lee H, Kim HY . (2011). Lantibiotics, class I bacteriocins from the genus Bacillus. J Microbiol Biotechnol 21: 229–235.
Liu D, Reeves PR . (1994). Escherichia coli K12 regains its O antigen. Microbiology 140 (Pt 1): 49–57.
Liu JD, Parkinson JS . (1989). Genetics and sequence analysis of the pcnB locus, an Escherichia coli gene involved in plasmid copy number control. J Bacteriol 171: 1254–1261.
Lopez D, Vlamakis H, Losick R, Kolter R . (2009). Cannibalism enhances biofilm development in Bacillus subtilis. Mol Microbiol 74: 609–618.
Lugtenberg B, Meijers J, Peters R, van der Hoek P, van Alphen L . (1975). Electrophoretic resolution of the "major outer membrane protein" of Escherichia coli K12 into four bands. FEBS Lett 58: 254–258.
Majeed H, Gillor O, Kerr B, Riley MA . (2011). Competitive interactions in Escherichia coli populations: the role of bacteriocins. ISME J 5: 71–81.
Merritt J, Qi F . (2012). The mutacins of Streptococcus mutans: regulation and ecology. Mol Oral Microbiol 27: 57–69.
Miquel S, Peyretaillade E, Claret L, de Vallee A, Dossat C, Vacherie B et al. (2010). Complete genome sequence of Crohn's disease-associated adherent-invasive E. coli strain LF82. PLoS One 5: e12714f.
Nadell CD, Xavier JB, Foster KR . (2009). The sociobiology of biofilms. FEMS Microbiol Rev 33: 206–224.
Osada H, Beppu T . (1985). Lipopolysaccharide defective mutant of Escherichia coli with decreased sensitivity to Colicin E2. Aric Biol Chem 49: 1813–1819.
Rendueles O, Travier L, Latour-Lambert P, Fontaine T, Magnus J, Denamur E et al. (2011). Screening of Escherichia coli species biodiversity reveals new biofilm-associated antiadhesion polysaccharides. MBio 2: e00043–00011.
Rendueles O, Ghigo JM . (2012). Multi-species biofilms: how to avoid unfriendly neighbors. FEMS Microbiol Rev 36: 972–989.
Rendueles O, Kaplan JB, Ghigo JM . (2013). Antibiofilm polysaccharides. Environ Microbiol 15: 334–346.
Riley MA, Gordon DM . (1999). The ecological role of bacteriocins in bacterial competition. Trends Microbiol 7: 129–133.
Riley MA, Cadavid L, Collett MS, Neely MN, Adams MD, Phillips CM et al. (2000). The newly characterized colicin Y provides evidence of positive selection in pore-former colicin diversification. Microbiology 146 (Pt 7): 1671–1677.
Riley MA, Wertz JE . (2002). Bacteriocins: evolution, ecology, and application. Annu Rev Microbiol 56: 117–137.
Sharma O, Datsenko KA, Ess SC, Zhalnina MV, Wanner BL, Cramer WA . (2009). Genome-wide screens: novel mechanisms in colicin import and cytotoxicity. Mol Microbiol 73: 571–585.
Skurnik D, Ruimy R, Andremont A, Amorin C, Rouquet P, Picard B et al. (2006). Effect of human vicinity on antimicrobial resistance and integrons in animal faecal Escherichia coli. J Antimicrob Chemother 57: 1215–1219.
Smajs D, Pilsl H, Braun V . (1997). Colicin U, a novel colicin produced by Shigella boydii. J Bacteriol 179: 4919–4928.
Smajs D, Matejkova P, Weinstock GM . (2006). Recognition of pore-forming colicin Y by its cognate immunity protein. FEMS Microbiol Lett 258: 108–113.
Spiers AJ, Rainey PB . (2005). The Pseudomonas fluorescens SBW25 wrinkly spreader biofilm requires attachment factor, cellulose fibre and LPS interactions to maintain strength and integrity. Microbiology 151: 2829–2839.
Stewart PS, Franklin MJ . (2008). Physiological heterogeneity in biofilms. Nat Rev Microbiol 6: 199–210.
Strassmann JE, Gilbert OM, Queller DC . (2011). Kin discrimination and cooperation in microbes. Annu Rev Microbiol 65: 349–367.
Tamura K, Peterson N, Stecher G, Nei M, Kumar S . (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739.
Tsai CM, Frasch CE . (1982). A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal Biochem 119: 115–119.
Valle J, Da Re S, Henry N, Fontaine T, Balestrino D, Latour-Lambert P et al. (2006). Broad-spectrum biofilm inhibition by a secreted bacterial polysaccharide. Proc Natl Acad Sci USA 103: 12558–12563.
Valle J, Da Re S, Schmid S, Skurnik D, D'Ari R, Ghigo JM . (2008). The amino acid valine is secreted in continuous-flow bacterial biofilms. J Bacteriol 190: 264–274.
Walker D, Rolfe M, Thompson A, Moore GR, James R, Hinton JC et al. (2004). Transcriptional profiling of colicin-induced cell death of Escherichia coli MG1655 identifies potential mechanisms by which bacteriocins promote bacterial diversity. J Bacteriol 186: 866–869.
Weltzien HU, Jesaitis MA . (1971). The nature of the colicin K receptor of Escherichia coli Cullen. J Exp Med 133: 534–553.
Wenzel SC, Muller R . (2009). Myxobacteria—'microbial factories' for the production of bioactive secondary metabolites. Mol Biosyst 5: 567–574.
Yan L, Boyd KG, Adams DR, Burgess JG . (2003). Biofilm-specific cross-species induction of antimicrobial compounds in bacilli. Appl Environ Microbiol 69: 3719–3727.
Acknowledgements
We are grateful to Juraj Bosák and David Šmajs for providing colicin U- and colicin Y-producing strains. We thank Erick Denamur for the strain ROAR029. We thank Franziska Stressmann for critical reading of the manuscript. OR was supported by a fellowship from the Network of Excellence EuroPathogenomics; European Community Grant LSHB-CT-2005-512061. This work was supported by a grant from the Région Ile-de-France (DIM Malinf) and from the French Government’s Investissement d’Avenir program, Laboratoire d'Excellence ‘Integrative Biology of Emerging Infectious Diseases’ (Grant no. ANR-10-LABX-62-IBEID).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Rendueles, O., Beloin, C., Latour-Lambert, P. et al. A new biofilm-associated colicin with increased efficiency against biofilm bacteria. ISME J 8, 1275–1288 (2014). https://doi.org/10.1038/ismej.2013.238
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.238
Keywords
This article is cited by
-
Engineering Escherichia coli for increased Und-P availability leads to material improvements in glycan expression technology
Microbial Cell Factories (2024)
-
Genomic and proteomic characterization of two strains of Shigella flexneri 2 isolated from infants’ stool samples in Argentina
BMC Genomics (2022)
-
Structural and functional insights into colicin: a new paradigm in drug discovery
Archives of Microbiology (2022)
-
Colicin Z, a structurally and functionally novel colicin type that selectively kills enteroinvasive Escherichia coli and Shigella strains
Scientific Reports (2019)