Abstract
The conversion of nitrite to nitric oxide in the denitrification pathway is catalyzed by at least two structurally dissimilar nitrite reductases, NirS and NirK. Although they are functionally equivalent, a genome with genes encoding both reductases has yet to be found. This exclusivity raises questions about the ecological equivalency of denitrifiers with either nirS or nirK, and how different ecological and evolutionary factors influence community assembly of nirS and nirK denitrifiers. Using phylogeny-based methods for analyzing community structure, we analyzed nirS and nirK data sets compiled from sequence repositories. Global patterns of phylogenetic community structure were determined using Unifrac, whereas community assembly processes were inferred using different community relatedness metrics. Similarities between globally distributed communities for both genes corresponded to similarities in habitat salinity. The majority of communities for both genes were phylogenetically clustered; however, nirK marine communities were more phylogenetically overdispersed than nirK soil communities or nirS communities. A more in-depth analysis was performed using three case studies in which a comparison of nirS and nirK community relatedness within the sites could be examined along environmental gradients. From these studies we observed that nirS communities respond differently to environmental gradients than nirK communities. Although it is difficult to attribute nonrandom patterns of phylogenetic diversity to specific niche-based or neutral assembly processes, our results indicate that coexisting nirS and nirK denitrifier communities are not under the same community assembly rules in different environments.
Similar content being viewed by others
Introduction
Denitrification is a facultative respiratory pathway in which nitrate is reduced to nitrogen gas through the intermediates nitrite, nitric oxide and nitrous oxide, and this trait is found among a wide variety of bacterial and archaeal genera from different phyla. Nitrite reduction to nitric oxide is catalyzed by one of two functionally equivalent, yet structurally divergent, nitrite reductases (Zumft, 1997). One is a cytochrome cd1 heme type reductase (NirS) encoded by the nirS gene, whereas the other is a copper-oxidoreductase (NirK) encoded by nirK. Previous experiments have shown both reductases to be functionally redundant, as one nir gene in a denitrifying organism can be eliminated and replaced by the other type (Glockner et al., 1993). However, they are believed to be mutually exclusive among denitrifiers, as no genome from a denitrifier to date has been found with both types present, although several organisms have been found with two or three copies of one nir type (Jones et al., 2008). Both nirS and nirK have been used as marker genes for denitrifying bacteria (Braker et al., 1998; Hallin and Lindgren, 1999) and changes in denitrifier community structure are often observed in studies of denitrifiers in the environment, showing how the ecological processes that have shaped denitrifier community structure are reflected in the distribution of genotypes among communities (for example, Braker et al., 2000; Tuomainen et al., 2003; Hallin et al., 2009).
The exclusivity of Nir types in denitrifiers has led researchers to speculate whether denitrifiers with one Nir type are ecologically equivalent to those with the other type, despite the functional equivalence of the two reductases. Hallin et al. (2009) reported differences in nirS:nirK gene abundance ratios among treatments in a fertilization experiment, which suggest that habitat selective factors exert an effect differently on organisms with different Nir types. In addition, recent studies on changes in denitrifier community composition and diversity indicate a niche differentiation between bacteria carrying one or the other Nir type (Hallin et al., 2006; Oakley et al., 2007; Smith and Ogram, 2008). Other emerging questions are how different ecological and evolutionary factors influence community assembly of nirS and nirK denitrifiers in different environments, and whether it is possible to assign ecological niches to different lineages within each Nir type. Answering this question would help to sort out the relative influence of niche-related, neutral and historical processes on community ecology of denitrifiers.
Our aim was to compare lineages of nirS and nirK from a range of environments to draw a deeper understanding of the underlying processes leading to the community patterns of the extant nirS and nirK diversity. We analyzed the global pool of nirS and nirK communities from a variety of environments compiled from sequence repositories. First, we examined the difference in phylogenetic community structure among the environments obtained for each gene (phylogenetic β-diversity) using the unique fraction metric (Unifrac; Lozupone and Knight, 2005) to provide further insight into how the type of environment correlates with the clustering of nirS or nirK communities. Although pairwise comparison of phylogenetic community structure provides information about the extent of similarity between different communities, methods that examine the phylogenetic relatedness of a given community in comparison with the global pool of species may provide additional information about the underlying mechanisms of community assembly, such as niche-based (habitat filtering or competition) or neutral (dispersal) assembly processes (Cavender-Bares et al., 2009). Therefore, we also analyzed the significance of phylogenetic clustering within a local environment in relation to the global pool of sequences (γ-diversity) using metrics of phylogenetic distinctness (Net Relatedness and Nearest Taxa indices, defined as tree-wide and branch-tip clustering, respectively; Webb, 2000) in combination with analysis of diversification rates. Of the studies compiled within this work, three were used as case studies (Santoro et al., 2006; Oakley et al., 2007; Smith and Ogram, 2008) to directly compare underlying processes of nirS and nirK phylogenetic community structure along different environmental gradients.
Materials and methods
Sequence data and alignments
Databases of nirS and nirK nucleotide sequences from environmental studies, including the three case studies, were downloaded from the Functional Gene Pipeline/Repository (http://fungene.cme.msu.edu). Sequences from both published and unpublished studies were used, and metadata for unpublished studies was acquired from Genbank annotation. We also screened the metagenomic databases within the CAMERA (Community Cyberinfrastructure for Advanced Marine Microbial Ecology Research and Analysis) website (Seshadri et al., 2007) for NirS and NirK homologs. Full-length NirS and NirK amino acid sequences from Pseudomonas aeruginosa PAO1 (containing nirS), Rhodobacter sphaeroides and Neisseria gonorrhoeae (each containing a different structural variant of nirK) were queried against all metagenomic open reading frames using tBLASTn, and reads with an e-value <0.1 were retrieved along with all available metadata.
All retrieved nucleotide sequences were checked for quality using a Perl script that determined the number of base-calling errors in each sequence (percentage of N bases), and compared each translated frame to a seed alignment using HMMER (Eddy, 1998; http://hmmer.janelia.org/) to screen for sequences with possible frame shifts. Seed alignments were created using full-length amino acid sequences from genomic sources found in Functional Gene Pipeline/Repository, which were aligned using the MAFFT-homologs algorithm (Katoh et al., 2005; http://align.bmr.kyushu-u.ac.jp/mafft/software/). The full data sets were aligned by amino acid using HMMER with the respective seed alignment and adjusted manually using ARB (Ludwig et al., 2004; http://www.arb-home.de/). Sequences that contained >0.1% N positions, frame shifts or resulted in e-values >0.001 when compared with the hidden Markov models were excluded. The coding nucleotide sequences were aligned by amino acid position, and sequences with non-overlapping ends were excluded.
Phylogenetic analysis
Phylogenetic trees were generated from nucleotide and amino acid alignments using RAxML v7.04 (Stamatakis, 2006; http://icwww.epfl.ch/~stamatak/index-Dateien/Page443.htm), with identical sequences removed before analysis. Because of the prohibitively long computational times, selection of the appropriate amino acid and nucleotide substitution models was based on a subset of sequences from each data set consisting of full-length sequences from genomes. Nucleotide model selection was performed using jModelTest v0.1.1 (Posada, 2008; http://darwin.uvigo.es/software/jmodeltest.html), and amino acid models were chosen using ProtTest (Abascal et al., 2005; http://darwin.uvigo.es/software/prottest.html). For both data sets, amino acid alignments were analyzed using the Whelan and Goldman amino acid substitution model (Whelan and Goldman, 2001), with estimated invariant site frequency and rate distribution shape parameters (I+Γ). The nucleotide alignments were partitioned by codon position and analyzed using the general time-reversible model (Rodriguez et al., 1990), with estimated I+Γ for each partition. Node support for all trees was determined using the fast bootstrapping option in RAxML with 1000 bootstrap replicates, and trees with environmental associations were plotted using the interactive Tree of Life online program (Letunic and Bork, 2007; http://itol.embl.de/). After pruning of the phylogenies to exclude taxa from sites with <10 sequences to avoid spurious results in subsequent community analyses, we restricted our analysis to sequences from the Functional Gene Pipeline/Repository database, as the search within CAMERA returned no sequences that met the requirements.
Phylogenetic diversity among environments
The sequences were grouped into different habitats based on information obtained from published studies, or inferred from sampling location and description in Genbank annotations for unpublished data (Supplementary Table S1, Supplementary Information). Individual sites that were taken along an environmental gradient within the same study were treated as separate sites. Marine habitats were classified based on the Venice system of marine water classification by salinity (Anonymous, 1958) and soil habitats were divided into undisturbed (for example, forest, natural grassland and wetland) or disturbed soil ecosystems (for example, polluted sites and agricultural soil; Table 1). Environmental dissimilarities among sampling sites within each database were calculated from nucleotide phylogenies using unweighted Unifrac (Lozupone and Knight, 2005). Non-metric multidimensional scaling, cluster analysis and permutational multivariate analysis of variance of Unifrac environmental distance matrices were performed using the ‘vegan’ and ‘cluster’ packages for R (http://www.r-project.org/). The non-metric multidimensional scaling was performed using 100 random starting configurations, constrained to three-dimensional solutions. Final non-metric multidimensional scaling scores from runs with the lowest stress values were used for three-dimensional plots of sampling sites. Cluster analysis of the Unifrac distance matrices was performed using Ward's method, and its usage with non-Euclidian distances has been reported to be effective, provided the distance matrix has Euclidian properties (Legendre and Legendre, 1998). The Unifrac distances matrices were checked to ensure Euclidian properties before clustering, using the ‘ade4’ package for R that examines distance matrices according to Gower's theorem (Gower and Legendre, 1986). Permutational multivariate analysis of variance was performed using 9999 permutations.
Phylogenetic community structure from individual sites within the nirS and nirK data sets was compared using the net relatedness index (NRI), which examines the average taxonomic distance between species within a community, and the nearest taxon index (NTI), which examines the average taxonomic distance between each species and its closest relative in the tree (Webb, 2000). Significance of phylogenetic clustering and overdispersion was determined using the PHYLOCOM program (Webb et al., 2008; http://www.phylodiversity.net/phylocom/) with a null model of random community assembly (9999 permutations of randomly drawn communities) from all sequences within other environmental sites (null model 1 in the PHYLCOM manual). A two-tailed test was used to determine the significance of NRI and NTI values at P=0.05, in which observed ranks of >9750 or <250 indicated significant clustering or overdispersion, respectively.
Rates of diversification
The rate of diversification within sites for each gene was examined using the tree shape statistic-γ (Pybus and Harvey, 2000), calculated from ultrametric trees. Sub-trees for each site were generated from the original phylogeny by pruning branches from all other sites, and then transforming to ultrametric trees using non-parametric rate smoothing (Sanderson, 1997) implemented in the ‘ape’ package for R (Paradis et al., 2004). Data were tested for departure from a molecular clock using log-likelihood ratio tests calculated in PAUP* v4.0 (Swofford, 2003; http://paup.csit.fsu.edu/), and tree shape statistics were calculated using the ‘ape’ package. The γ-statistic was used to determine whether the rate of diversification in each environment significantly differed from a model of constant diversification. If γ>0, then a phylogeny's internal nodes are closer to the tips than expected under a constant rate of diversification, whereas γ<0 indicates that nodes are closer to the root than expected under a model of constant diversification. To test the significance of γ-values, the Monte Carlo Constant Rates test (Pybus et al., 2002) was used as implemented in the ‘laser’ package in R. As this method requires an estimate of the total number of possible species, we used the total number of sequences present in each data set as the total number of nirK or nirS genotypes.
Statistical analysis of clustering and diversification rates among habitat categories
The distribution of NRI, NTI and γ-values were analyzed using analysis of variance to determine the variation in phylogenetic clustering and diversification rates among the different habitat types. Tests of normality were performed using the Shapiro–Wilk W test, and the Kruskal–Wallis rank-sums test was used when values were not normally distributed. Regression analysis using Pearson's product-moment correlation coefficient was performed for comparing NRI, NTI and γ-values with environmental parameters retrieved from three case studies that compared nirS and nirK sequence diversity in aquatic (Santoro et al., 2006; Oakley et al., 2007) and soil habitats (Smith and Ogram, 2008). All analyses were performed using R.
Results
Distribution of data sets and phylogenetic analysis
The majority of nirK sequences were from terrestrial studies, whereas sequences from marine and estuary environments dominated the nirS databases (Supplementary Table S1, Supplementary Information). Phylogenetic analysis of both data sets resulted in nucleotide phylogenies with fewer polytomies and greater overall bootstrap support (58% and 55% median bootstrap support for nirS and nirK, respectively) than analysis of amino acids (37% and 32% median bootstrap support for nirS and nirK, respectively), and we therefore based our analysis on nucleotide phylogenies (Supplementary Figures S1 and S2, Supplementary material). Sequences from marine and soil samples were largely concentrated in a few different clades of the nirK phylogeny, whereas those from freshwater and activated sludge samples were more scattered throughout the tree. The same trend was observed in the nirS phylogeny; however, sequences from soil samples were more dispersed among sequences from marine or estuarine samples. Large branch lengths were observed in the nirK tree among a group of sequences from three studies on marine habitats, the validity of which was confirmed by inspection of amino acid substitutions at numerous sites in the alignment. Nevertheless, it has been shown that both Unifrac analysis and NRI and NTI are robust to minor variations in tree topology (Webb, 2000; Lozupone et al., 2007).
Comparison of β-diversity using Unifrac
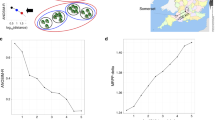
The non-metric multidimensional scaling of Unifrac dissimilarity matrices resulted in three-dimensional ordinations with moderate stress values (Figures 1a and b). Both data sets showed a pattern of separation between saline and non-saline environments, suggesting that differences in community relatedness occur along a salinity gradient. Freshwater lake and estuarine communities ranging from limnetic to oligohaline in the nirS data set differentiated from those found in habitats of higher salinity, which were more similar to marine environments. nirS communities from all mesohaline sampling sites in the Baltic sea grouped with those from a water column of approximately the same salinity in the Black sea, whereas communities from mixohaline sites in the Arabian sea grouped closer to those from a Pacific coastal aquifer. However, exceptions to groupings based on salinity were observed, as communities in oxygen-deficient sediments off the Pacific coast of Mexico were more similar to non-saline communities in both data sets. In addition, nirS communities in a mesohaline estuary and a Mediterranean biofilm did not group with other saline sites, but closer to soil or freshwater and activated sludge environments. Soil communities from disturbed soil environments did not separate from undisturbed sites, and activated sludge communities were similar to those from limnetic environments in both data sets.
Three-dimensional ordinations of Unifrac environmental distance matrices using non-metric multidimensional scaling. Habitat type is indicated by symbols listed in Table 1. (a) Ordination of nirK communities; final stress value=13.1. (b) Ordination of nirS communities; final stress value=11.9.
As the sequences were derived from PCR-based studies, we were concerned about the possibility of clustering because of differences in primer pairs used among the studies. Cluster analysis of Unifrac distance matrices showed varied clustering of sites from studies using the same primer sets for both nirS and nirK data sets (Supplementary Figure S3, Supplementary Information). Using permutational multivariate analysis of variance of Unifrac distance matrices, a significant effect of habitat type (P<0.001) was observed for both data sets, as well as a significant interaction between habitat type and primer set (P=0.023 and P=0.019 for nirS and nirK, respectively). However, the significant effect of habitat type, combined with the observation of several instances in which sites from studies using different primers formed distinct clusters and vice versa, suggests that any bias in environmental grouping due to primer usage was outweighed by the effect of the environment itself.
Phylogenetic clustering and diversification rates
Using the NTI metric resulted in the majority of the communities at the sites being significantly clustered in both data sets. Similar results were observed using NRI values; however, several sites were also significantly overdispersed (Supplementary Table S2, Supplementary Information). Soil communities in both data sets were for the most part significantly clustered; however, three nirS and four nirK soil communities from different studies were not significantly clustered using the NRI metric. Using NTI, only one of the nirK soil communities was not significantly clustered. Two of the five nirK freshwater communities showed significant clustering using NRI, but all were significantly clustered based on the NTI metric. All activated sludge and wastewater nirK communities were significantly clustered. In contrast, almost all of the nirK communities from saline environments showed no significant clustering using NRI, with three communities being significantly overdispersed. A majority of nirS marine and estuarine communities were significantly clustered using NRI, and all but five were clustered as determined by the NTI metric. Unlike the nirK data set, only half of the activated sludge/wastewater nirS communities showed significant clustering.
After grouping the sites into habitat categories according to Table 1, analysis of variance of NTI values showed no significant differences among environments in the nirK or nirS data set. However, significant effects of habitat type on the NRI values for nirK (Kruskal–Wallis; χ2=25.6, d.f.=5, P<0.001) and nirS (Kruskal–Wallis; χ2=12.73, d.f.=5, P<0.05) were observed. The nirK communities from mixohaline and mesohaline environments were less clustered than those from limnetic, soil and activated sludge and wastewater habitats, with the exception of an outlying mesohaline Black sea community (Figure 2a). The nirS communities in all habitat categories were largely clustered; however, those from mixohaline, limnetic and activated sludge/wastewater habitats resulted in lower NRI values than those observed in other habitat types (Figure 2b).
Distribution of net relatedness indices (NRI) for different habitat types among (a) nirK communities and (b) nirS communities. Values >0 indicate phylogenetically clustered communities, whereas negative values indicate overdispersed communities. Boxplots consist of habitat categories with n of ⩾3 sites, and categories with fewer sites are excluded from the figure. Circles indicate outlier data points (>1.5 times the interquartile range) within each habitat grouping. See Supplementary Table S2, Supplementary Information for complete list of values and significance tests for each site.
Analysis of tree shape among the different environments using the γ-statistic was performed for comparing the rates of diversification with the propensity for clustering among different communities, the logic being that communities that are more clustered yet have nodes closer to the root may be under different selection pressures than those that have nodes closer to the tips. Likelihood-ratio tests resulted in both data sets violating a molecular clock assumption (P<0.001); however, we proceeded with the analysis using ultrametric phylogenies determined using non-parametric rate smoothing of substitution rates. No communities were deemed to be significantly deviating from a constant rate of diversification, which was not surprising given the conservative nature of the Monte Carlo Constant Rates test (Pybus et al., 2002; Martin et al., 2004). Interestingly, a significant positive correlation was observed between γ-values and NTI values for both nirS (r=0.621, P<0.001) and nirK data sets (r=0.375, P=0.003), whereas only nirS NRI values were significantly correlated to γ-values (r=0.577, P<0.001). Analysis of variance showed no significant differences in γ-values among habitat types for either gene.
Case studies: direct comparisons of nirS and nirK communities
Three of the compiled studies allowed us to directly compare nirK and nirS phylogenetic community structure along environmental gradients (Table 2). The first is on a soil restoration chronosequence (Smith and Ogram, 2008) in which we observed a significant positive correlation between nitrate concentration and phylogenetic relatedness of nirS communities using NRI, NTI and γ. No significant correlations were observed between nitrate and nirK clustering; however, we did observe a significant negative correlation between nitrate and nirK diversification rates. In contrast, the relatedness and diversification rates of nirS communities were negatively correlated with ammonium, whereas a significant positive correlation was observed between the diversification rate of nirK communities and ammonium levels. There were no obvious trends when comparing nirS NRI or NTI values with soil moisture or organic matter. All nirK communities were significantly clustered based on both NRI and NTI values (Supplementary Table S2, Supplementary Information), and no trend was observed between level of clustering of these communities and any of the environmental parameters besides depth.
The second study, by Oakley et al. (2007), analyzed differences in denitrifying communities in a sub-oxic zone of the Black Sea, with information on nitrate, nitrite and ammonium concentrations at different depths. Changes in salinity among the sampling points was minimal, varying from 19.9 psu to 20.6 psu (B Oakley, personal communication); however, a significant negative correlation between salinity and nirS community relatedness (NRI) was found. Similar to the soil study, nirS NRI values were positively correlated with nitrate concentrations, whereas nirK communities showed a nonsignificant negative trend between NRI and nitrate. No significant correlations were observed when community relatedness or diversification rates were compared with nitrite; however, nirK NTI was positively correlated with ammonium concentrations.
The Pacific coastal aquifer study (Santoro et al., 2006) examined differences in nirK and nirS communities along a salinity and nitrate gradient, with additional information on ammonium levels. The salinity gradient ranged from 8.7 psu to 34.5 psu, and unlike the Black sea study we observed a significant positive relationship between salinity and nirS community relatedness. We also found a positive correlation between nirK community relatedness and salinity based on the NTI metric, as well as diversification rates. Interestingly, nirS communities responded differently to the nitrate gradient in comparison to the previous two studies, as a significant negative correlation was observed between nitrate concentrations and nirS community relatedness and diversification rates. Despite the large range of ammonium values among the sites (9.2–161.5 μM), no significant correlations were observed between ammonium and community relatedness or diversification rates for either gene.
Discussion
The objective of this study was to compare and contrast the structure of denitrifier communities, as determined by nirS and nirK phylogenies, to infer community assembly processes that may differ between denitrifiers with one nir gene or the other in different environments. Examination of phylogenetic β-diversity may provide a means to determine the relative importance of niche-based versus neutral community assembly, as pairwise differences in community structure may be compared with differences in geographical location or environmental parameters (Graham and Fine, 2008). A large proportion of the sequences were from studies in which detailed information on environmental parameters were not provided. This was especially true for the soil habitats and among these we found no separation of the nirS or nirK communities based on our habitat categories. Unfortunately, we did not have information on soil pH, which has been shown to affect denitrifier community composition (Enwall et al., 2005) and to be a global predictor of soil bacterial community structure (Fierer and Jackson, 2006). However, we observed a clear separation of nirS and nirK communities from saline and non-saline environments. In contrast to nirK, the nirS data set was dominated by marine studies, which could be a consequence of the limited number of studies that examine nirK communities in aquatic habitats. However, it is possible that denitrifying species with nirK that are endemic to marine environments may possess a different nirK structural type (Ellis et al., 2007), and are simply not detected using currently available primer sets. Nevertheless, for both genes, the denitrifier community composition changed in response to salinity rather than geographical location within the aquatic environments. Separation according to salinity was also observed in a study on global patterns of bacterial diversity using 16S rRNA sequences from a variety of environments (Lozupone and Knight, 2007). We found it interesting that functional gene lineages behave in a similar manner, despite being potentially more labile among prokaryotic genomes than 16S rRNA genes. If horizontal gene transfer is an important factor in the evolution of denitrification genes (Jones et al., 2008), it is working in concert with ecological mechanisms that constrain functional gene diversification within habitat types of different salinity. According to Graham and Fine (2008), communities in which neutral assembly processes are important should show a strong pattern of geographic structure. Thus, although not ruling out neutral processes, our observation of a significant effect of habitat type on Unifrac distances between sites suggests that niche-based processes, such as environmental filtering, have a role in the structuring of both nirS and nirK denitrifier communities.
Our analysis of phylogenetic community structure using NRI, NTI and diversification rates indicated niche-based assembly processes for the denitrifier communities with most being significantly clustered for both genes. It has been shown that the level of phylogenetic clustering among plant communities tends to increase with increasing spatial scale, which could be attributed to biogeographical processes (Cavender-Bares et al., 2006). However, the Unifrac analysis placed greater importance on habitat type rather than geographic location in our study as discussed earlier. Previous research examining measures of phylogenetic community structure have interpreted clustering as evidence of environmental filtering, or differential colonization and/or adaptive radiation events and overdispersed communities as the result of competition, although interactions between these niche-based processes may also occur (Webb, 2000; Horner-Devine and Bohannan, 2006). Moreover, Kembel (2009) used simulations to show that communities assembled under neutral processes might still show nonrandom phylogenetic structure. Regardless, even if the phylogenetic metrics are not convincingly teasing apart the different assembly processes, there was indication that nirK and nirS communities differ in phylogenetic structure among habitat types when analyzed at the global scale.
Differences in phylogenetic community structure between nir types were best explored by focusing on studies in which sequences for both genes were sampled along environmental gradients. In the soil study of Smith and Ogram (2008), the results based on ∫-LIBSHUFF and analysis of molecular variance indicated that not only did nirK and nirS denitrifiers respond differently to environmental gradients, but that nirK denitrifiers may show greater habitat selectivity, with differences in nirK communities strongly correlated with differences in soil moisture. Although the significant clustering of nirK communities corresponds with the results of their analysis of molecular variance, we did not observe any obvious trend between nirK NRI, NTI or γ-values and soil moisture. This may be attributed to differences in the type of analysis performed. Regardless, we did observe significant positive correlations of nirS NRI, NTI and γ-values with nitrate, whereas negative correlations were observed with ammonium concentrations. These results reflect the findings of Smith and Ogram (2008), in that communities of different nir-type denitrifiers responded differently to environmental gradients. In addition, we hypothesize that nitrate concentration was driving community assembly processes among nirS denitrifiers in this study, whereas nirK denitrifiers may be responding to a different environmental parameter. In the Pacific coastal aquifer study, Santoro et al. (2006) reported that nirK populations were specific to each station sampled along the gradient, whereas nirS populations tended to overlap to a greater degree, especially among sites with similar salinity. We observed nirS communities to be significantly clustered among all sites, whereas nirK communities were either not significantly clustered or overdispersed (NRI). However, attributing this pattern to either salinity or nitrate is difficult, given that both vary simultaneously. The Black Sea study by Oakley et al. (2007) showed a significant negative correlation between nirS community relatedness and salinity. However, the small difference in salinity among the sampling sites was most likely insignificant, thereby removing a potentially major driver of community differences. Oakley et al. (2007) state that sites with higher nitrate to nitrite ratios at suboxic conditions (CG157 and CG158) are likely to have the highest diversity of denitrifiers, as is shown in their analysis of nirK libraries. This pattern is also reflected in our results, as nirK communities in these sites showed random or overdispersed community structure, whereas nirS communities were significantly clustered (Supplementary Table S2, Supplementary Information). Interestingly, nirK communities at site CG159, with low nitrate, high nitrite and increasing ammonium levels, were significantly clustered, but showed decreasing rates of diversification (γ). Yet, the next sampling point, with high ammonium and nitrite but low nitrate, nirK communities were far less clustered but undergoing rapid diversification. In contrast, all nirS communities were significantly clustered and had increasing rates of diversification, suggesting differences in community assembly processes among nirK communities along the gradient of available nitrogen species. The different values of NRI and γ observed between sites CG157 and CG160 may be explained by CG157 being a border zone for oxic and anoxic conditions, and thus the variety of niches may be greater here than at the deeper zone.
Our work shows that the functional equivalence of nirS and nirK in denitrifying organisms is not reflected in their ecological distribution, and it is likely that different processes underlie the community assembly of each type of denitrifier. The scale at which this analysis is performed poses a challenge in trying to identify the relative importance of niche-based versus neutral assembly rules. We could extract a clear trend of community similarity by habitat type from both nirS and nirK phylogenies, with salinity emerging as an important factor in determining community relatedness at the global scale. The importance of other environmental parameters in determining the community structure of nirK or nirS denitrifiers was difficult to answer because of the limited amount of information about each site. However, these methods do allow for the detection of differences in phylogenetic community structure between the two types of denitrifier communities. Although this analysis may provide further insight into the possible mechanisms and important environmental parameters that govern denitrifier community assembly, additional information from manipulative experiments would aid in confirming (or falsifying) the inferences presented in this study.
References
Abascal F, Zardoya R, Posada D . (2005). ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21: 2104–2105.
Anonymous . (1958). The Venice system for the classification of marine waters according to salinity. Limnol Oceanogr 3: 346–347.
Braker G, Fesefeldt A, Witzel K . (1998). Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl Environ Microbiol 64: 3769–3775.
Braker G, Zhou JZ, Wu LY, Devol A, Tiedje JM . (2000). Nitrite reductase genes (nirK and nirS) as functional markers to investigate diversity of denitrifying bacteria in Pacific northwest marine sediment communities. Appl Environ Microbiol 66: 2096–2104.
Cavender-Bares J, Keen A, Miles B . (2006). Phylogenetic structure of Floridian plant communities depends on taxonomic and spatial scale. Ecology 87: S109–S122.
Cavender-Bares J, Kozak KH, Fine PVA, Kembel SW . (2009). The merging of community ecology and phylogenetic biology. Ecol Lett 12: 693–715.
Eddy SR . (1998). Profile hidden Markov models. Bioinformatics 14: 755–763.
Ellis M, Grossman JG, Eady RR, Hasnain SS . (2007). Genomic analysis reveals widespread occurrence of new classes of copper nitrite reductases. J Biol Inorg Chem 12: 1119–1127.
Enwall K, Philippot L, Hallin S . (2005). Activity and composition of the denitrifying bacterial community respond differently to long-term fertilization. Appl Environ Microbiol 71: 8335–8343.
Fierer N, Jackson RB . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci 103: 626–631.
Glockner AB, Jungst A, Zumft WG . (1993). Copper-containing nitrite reductase from Pseudomonas aureofaciens is functional in a mutationally cytochrome-cd1-free background (nirS-) of Pseudomonas stutzeri. Arch Microbiol 160: 18–26.
Gower JC, Legendre P . (1986). Metric and Euclidian properties of dissimilarity coefficients. J Classif 3: 5–48.
Graham CH, Fine PVA . (2008). Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time. Ecol Lett 11: 1265–1277.
Hallin S, Jones CM, Schloter M, Philippot L . (2009). Relationship between N-cycling communities and ecosystem functioning in a 50-year-old fertilization experiment. ISMEJ 1: 1–9.
Hallin S, Lindgren PE . (1999). PCR detection of genes encoding nitrile reductase in denitrifying bacteria. Appl Environ Microbiol 65: 1652–1657.
Hallin S, Throback IN, Dicksved J, Pell M . (2006). Metabolic profiles and genetic diversity of denitrifying communities in activated sludge after addition of methanol or ethanol. Appl Environ Microbiol 72: 5445–5452.
Horner-Devine MC, Bohannan BJM . (2006). Phylogenetic clustering and overdispersion in bacterial communities. Ecology 87: S100–S108.
Jones CM, Stres B, Rosenquist M, Hallin S . (2008). Phylogenetic analysis of nitrite, nitric oxide, and nitrous oxide respiratory enzymes reveal a complex evolutionary history for denitrification. Mol Bio Evol 25: 1955–1966.
Katoh K, Kuma K, Toh H, Miyata T . (2005). MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33: 511–518.
Kembel SW . (2009). Disentangling niche and neutral influences on community assembly: assessing the performance of community phylogenetic structure tests. Ecol Lett 12: 949–960.
Legendre P, Legendre L . (1998). Numerical Ecology. Elsevier: Amsterdam, The Netherlands.
Letunic I, Bork P . (2007). Interactive Tree of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23: 127–128.
Lozupone CA, Knight R . (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71: 8228–8235.
Lozupone CA, Hamady M, Scott KT, Knight R . (2007). Quantitative and qualitative beta diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73: 1576–1585.
Lozupone CA, Knight R . (2007). Global patterns in bacterial diversity. Proc Natl Acad Sci 104: 11436–11440.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Martin AP, Costello EK, Meyer AF, Nemergut DR, Schmidt SK . (2004). The rate and pattern of cladogenesis in microbes. Evolution 58: 946–955.
Oakley BB, Francis C, Roberts KJ, Fuchsman CA, Srinivasan S, Staley JT . (2007). Analysis of nitrite reductase (nirK and nirS) genes and cultivation reveal depauperate community of denitrifying bacteria in the Black Sea suboxic zone. Environ Microbiol 9: 118–130.
Paradis E, Claude J, Strimmer K . (2004). APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20: 289–290.
Posada D . (2008). jModelTest: phylogenetic model averaging. Mol Bio Evol 25: 1253–1256.
Pybus O, Rambaut A, Holmes E, Harvey PH . (2002). New inferences from tree shape: numbers of missing taxa and population growth rates. Sys Bio 51: 881–888.
Pybus OG, Harvey PH . (2000). Testing macro-evolutionary models using incomplete molecular phylogenies. Proc R Soc London B Biol Sci 267: 2267–2272.
Rodriguez F, Oliver JL, Marin A, Medina JR . (1990). The general stochastic-model of nucleotide substitution. J Theor Bio 142: 485–501.
Sanderson MJ . (1997). A nonparametric approach to estimating divergence times in the absence of rate constancy. Mol Bio Evol 14: 1218–1231.
Santoro AE, Boehm AB, Francis CA . (2006). Denitrifier community composition along a nitrate and salinity gradient in a coastal aquifer. Appl Environ Microbiol 72: 2102–2109.
Seshadri R, Kravitz SA, Smarr L, Gilna P, Frazier M . (2007). CAMERA: a community resource for metagenomics. PLoS Biol 5: e75.
Smith J, Ogram A . (2008). Genetic and functional variation in denitrifier populations along a short-term restoration chronosequence. Appl Environ Microbiol 74: 5615–5620.
Stamatakis A . (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Swofford DL . (2003). PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates: Sunderland, MA.
Tuomainen JM, Hietanen S, Kuparinen J, Martikainen PJ, Servomaa K . (2003). Baltic Sea cyanobacterial bloom contains denitrification and nitrification genes, but has negligible denitrification activity. FEMS Microbiol Ecol 45: 83–96.
Webb CO . (2000). Exploring the phylogenetic structure of ecological communities: An example for rain forest trees. Am Nat 156: 145–155.
Webb CO, Ackerly DD, Kembel SW . (2008). Phylocom: software for the analysis of phylogenetic community structure and trait evolution. Bioinformatics 24: 2098–2100.
Whelan S, Goldman N . (2001). A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Bio Evol 18: 691–699.
Zumft WG . (1997). Cell biology and molecular basis of denitrification. Microbiol Mol Bio Rev 61: 533–616.
Acknowledgements
We thank C Lozupone and S Kembel for their help with software and statistical advice, as well as B Oakley for the additional information on the Black Sea study. We also thank L Philippot for helpful discussions. All phylogenetic analyses were performed using the UPPMAX computational center (Uppsala University, Sweden). Funding by the Swedish Research Council and the Uppsala Microbiomics Center grant from Formas is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Jones, C., Hallin, S. Ecological and evolutionary factors underlying global and local assembly of denitrifier communities. ISME J 4, 633–641 (2010). https://doi.org/10.1038/ismej.2009.152
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.152
Keywords
This article is cited by
-
Soil denitrification response to increased urea concentration constrains nitrous oxide emissions in a simulated cattle urine patch
Plant and Soil (2023)
-
Temporal characteristics of algae-denitrifying bacteria co-occurrence patterns and denitrifier assembly in epiphytic biofilms on submerged macrophytes in Caohai Lake, SW China
Journal of Oceanology and Limnology (2023)
-
Reactive nitrogen restructures and weakens microbial controls of soil N2O emissions
Communications Biology (2022)
-
Impacts of graphitic nanofertilizers on nitrogen cycling in a sandy, agricultural soil
Journal of Nanoparticle Research (2022)
-
Saline and alkaline stresses alter soil properties and composition and structure of gene-based nitrifier and denitrifier communities in a calcareous desert soil
BMC Microbiology (2021)