Abstract
Although there are a number of different allopolyploids in the plant kingdom, the exact ancestral parents of some allopolyploids have not been well characterized. We propose a strategy in which virtual allopolyploid lines derived from different types of parental species are used to investigate the progenitors of an allopolyploid. The genotypes of the parental lines and the natural allopolyploid were established using a set of DNA molecular markers. The genotypes of the virtual lines were then derived from those of the parental lines, and compared extensively with that of the natural allopolyploid. We applied this strategy to investigate the progenitors of the C subgenome of Brassica napus (rapeseed, AACC). A total of 39 accessions from 10 wild and 7 cultivated types of the B. oleracea cytodeme (CC), and 4 accessions of B. rapa (AA) were used to construct 156 virtual rapeseed lines. Genetic structure was compared among natural rapeseed, virtual rapeseed lines, and their parental lines by principal component analysis and analysis of ancestry. Our data showed that the C subgenome of natural rapeseed was related closely to the genome of cultivated B. oleracea and its related wild types, such as B. incana, B. bourgeaui, B. montana, B. oleracea ssp. oleracea and B. cretica. This finding indicated that these types or their progeny might be ancestral donors of the C subgenome of rapeseed. The successful application of the strategy of virtual allopolyploidy in rapeseed demonstrates that it can possibly be used to identify the progenitors of an allopolyploid species.
Similar content being viewed by others
Introduction
Polyploidy has an important role in the evolution of plants. Polyploid plants have a number of advantages over diploids, such as an improved ability to survive in harsh environments, increased resistance to pathogens and permanent fixation of hybrid vigor (Stebbins, 1971; Soltis and Soltis, 2000; Otto, 2007). The frequency of polyploidy in angiosperms has been estimated to be 15% (Wood et al., 2009). Moreover, many ‘diploid’ plants are in fact ancient polyploids that have undergone at least one round of genome duplication during their evolution; hence, it can be estimated that over 70% of angiosperms are polyploid (Masterson, 1994; Wolfe, 2001).
Winge (1917) proposed that allopolyploids arose from interspecific hybridization followed by chromosome doubling. Subsequently, a number of important crops, such as wheat (Triticum aestivum), cotton (Gossypium hirsutum) and rapeseed (Brassica napus) have been confirmed to be allopolyploids in which two or more divergent genomes combined (Chen, 2007). Some natural allopolyploids show less genetic diversity than the parental species (Becker et al., 1995; Tanksley and McCouch, 1997; Abdalla et al., 2001; Reif et al., 2005), which leads us to speculate that a small number of variants of the parental species contributed to these allopolyploids. However, little is known about their exact progenitors in some cases.
B. napus (rapeseed, AACC) is thought to have originated from a spontaneous hybridization between B. rapa (AA) and B. oleracea (CC) (UN, 1935), but its exact progenitors have not been identified for the following reasons: (1) Although rapeseed has been domesticated as a crop for only 300–400 years, wild rapeseed has not been found in nature (Gómez-Campo and Prakash, 1999). Hence, no information about the progenitors of rapeseed can be gained from wild rapeseed. (2) The frequent introgression of B. rapa into B. napus in breeding programs has led to substantial amounts of variation in the A subgenome of modern rapeseed (Qian et al., 2006). Hence, it is difficult to distinguish the genetic contributions of the B. rapa progenitor from genetic material that has been introduced more recently. (3) There are 10 wild relatives of B. oleracea in nature (Snogerup et al., 1990), and various cultivated forms. The progenies between wild and cultivated types of B. oleracea exhibit partial fertility in some cases (Snogerup et al., 1990; Kianian and Quiros, 1992; Von Bothmer et al., 1995). Harberd (1976) referred to these types as the B. oleracea cytodeme. However, the C subgenome of rapeseed shows much less variation than the genome of B. oleracea (Becker et al., 1995; Seyis et al., 2003), which raises the question of which types of the B. oleracea cytodeme contributed to the formation of rapeseed.
It has been proposed previously that rapeseed originated in the Mediterranean region of Southwest Europe, where the distributions of the wild types of B. rapa and B. oleracea overlap (Sinskaia, 1928; Schiemann, 1932). Owing to obvious differences between European and Japanese rapeseed, it has been suggested that Japanese rapeseed was derived from natural hybridization between Asian B. rapa and cultivated European B. oleracea (Olsson, 1954; Naughton, 1976). Gómez-Campo and Prakash (1999) have suggested that rapeseed originated from crosses between cultivated types of the parental species, because it would be relatively easy for B. oleracea and B. rapa to mate reciprocally in an agricultural environment. An alternative hypothesis was suggested by Song and Osborn (1992), who found that the majority of accessions of B. napus contain the same chloroplast haplotype as B. montana (a wild taxon of B. oleracea), and thus assumed that B. montana was involved in the origination of rapeseed. However, the ancestral donor(s) of the C subgenome in rapeseed has not been clarified in these reports. One possible reason is that only a few types of the B. oleracea cytodeme were considered in these studies.
Genetic changes that have occurred in resynthesized allopolyploids relative to parental lines have been documented (Song et al., 1995; Prakash et al., 1999; Comai et al., 2000; Ozkan et al., 2001; Kashkush et al., 2002; Gaeta et al., 2007). However, these changes are much fewer than the genetic differences between the genomes of the parental species (Comai et al., 2000; Ozkan et al., 2001; Kashkush et al., 2002; Parkin et al., 2003; Rana et al., 2004; Lukens et al., 2006; Rousseau-Gueutin et al., 2008). Given that there are high degrees of macrosynteny and colinearity between an allopolyploid and its progenitors, the majority of genetic characteristics of the progenitor species are maintained in the allopolyploid during its generation and evolution (Rana et al., 2004; Rousseau-Gueutin et al., 2008). Thus, it should be possible to identify the progenitors of an allopolyploid by comparing the genetic structure of synthetic lines derived from all natural variants of the parental species with that of the natural allopolyploid: it is most likely that the progenitors are the parental variants that give rise to the synthetic lines that are most similar to the natural allopolyploid.
In this study, we have proposed a strategy to investigate the progenitors of an allopolyploid. The parental lines and the natural allopolyploid are genotyped using DNA molecular markers, and the genotype of a virtual line is then derived from that of its parents, that is, it is assumed that markers that are present in the parental lines are also present in the virtual allopolyploid. The genotypes of the virtual lines are then compared extensively with that of the natural allopolyploid. We have performed extensive comparisons of genetic structure among natural rapeseed and virtual rapeseed lines derived from different variants of the B. oleracea cytodeme. The results of the study suggest that the C subgenome of natural rapeseed is related closely to that of cultivated B. oleracea and its related wild types, such as B. incana, B. bourgeaui, B. montana, B. oleracea ssp. oleracea and B. cretica. Therefore, these types might be the ancestral donors of the C subgenome of rapeseed. This study demonstrates that it is possible to use virtual allopolyploid lines to investigate the progenitors of an allopolyploid species.
Materials and methods
Plant materials
We used a panel of accessions of Brassica to investigate the ancestral donor(s) of the C subgenome of natural rapeseed (Figure 1). A total of 25 accessions from 10 wild types and 14 accessions from 7 cultivated types of the B. oleracea cytodeme, on the basis of the classification proposed by Snogerup et al. (1990), were collected from the Centre for Genetic Resources (CGN; Wageningen, The Netherlands), the Institute of Plant Genetics and Crop Plant Research (IPK; Gatersleben, Germany), the Universidad Politécnica de Madrid (UPM; Madrid, Spain), the University of California (UC; Davis, CA, USA) and Southwest University (SWU; China). Four accessions of B. rapa that represented genetic variants from two centres of origin of B. rapa (East Asia and Europe) and six accessions of B. napus that represented natural variants of rapeseed (two European winter, two Chinese semi-winter and two European spring rapeseed) were selected on the basis of studies of the evolution of B. rapa (Gómez-Campo and Prakash, 1999; Qian et al., 2003; Zhao et al., 2005) and analyses of genetic diversity in natural rapeseed (Diers and Osborn, 1994; Becker et al., 1995; Qian et al., 2006). A total of 156 virtual lines of rapeseed were developed from the 39 accessions of the B. oleracea cytodeme and from the 4 accessions of the B. rapa as described below.
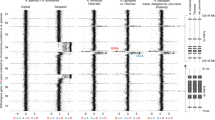
Cluster analysis of Nei's matrix distances among 49 accessions of Brassica, as revealed with AFLP and SSR markers. Bootstrap values above 50%, which were calculated with 1000 replications, are shown on the branches. At a genetic distance of 0.51, the accessions of the B. oleracea cytodeme were classified into three subgroups, whereas the accessions from B. napus and B. rapa were clustered separately into individual groups.
DNA isolation and development of molecular markers
Young leaves from the 49 accessions of the B. oleracea cytodeme, B. rapa, and B. napus were collected and pooled from at least five individuals of each accession at the seedling stage. Genomic DNA was isolated using the CTAB method (Saghai-Maroof et al., 1984). Amplified fragment length polymorphisms (AFLPs) were amplified using 11 primer pairs and the resulting PCR products were separated on a Li-Cor 4300 Sequencer (LI-COR Biosciences; Lincoln, NE, USA). Distinct bands with a fragment size of 100–600 bp were scored. In all, 83 primer pairs for simple sequence repeats (SSRs) in the A and C genomes of Brassica were selected randomly from the website www.brassica.info. The PCR products that corresponded to the SSR markers were separated on a 10% polyacrylamide gel and stained with silver nitrate.
Development of virtual allopolyploid lines
The accessions of the parental species and natural rapeseed were genotyped with the AFLP and SSR markers as described above. The genotypes of different accessions of B. oleracea and B. rapa were then combined in the following manner to give those of the virtual lines. For each locus, a score of ‘1’ was given to the virtual line if the AFLP or SSR marker was present in at least one of the parental lines, whereas if the marker was absent in both parents, the virtual line was assigned a score of ‘0’. In this way, the genotypes of the virtual allopolyploid lines could be derived from those of the corresponding parental lines without the need for interspecific hybridization. In total, 156 virtual rapeseed lines were constructed from 39 accessions of the wild and cultivated forms of B. oleracea and from the 4 accessions of B. rapa. The genotypes of the virtual lines were then compared with those of the natural rapeseed lines.
Data analysis
Several methods were used to analyze the genetic structure of the accessions of interest. The matrix of genetic distance among the 49 accessions of Brassica was calculated using the formula of Nei and Li (1979), and subjected to cluster analysis using the unweighted pair group method with arithmetic averages from the NTSYS-PC program (http://www.exetersoftware.com/cat/ntsyspc/ntsyspc.html) (Rohlf, 1997). The confidence values of the resulting dendrogram were tested by bootstrap analysis with 1000 replications, using the software package WINBOOT (http://www.riceworld.org/science/software/winboot.asp) (Yap and Nelson, 1996). To explore the relationships of ancestry among the 49 accessions of Brassica, the STRUCTURE ver. 2.2 software was used for 10 000 iterations after a burn-in of 1000 iterations (Falush et al., 2007). To compare genetic structures among natural rapeseed, the virtual rapeseed lines, and the parental species, the genetic distance was evaluated by principal component analysis in NTSYS-PC (Rohlf, 1997).
Statistical analyses such as Pearson's simple correlation, F- and t-tests were performed using SAS ver. 6.07 (http://www.sas.com/) (SAS Institute, 1992).
Results
Genetic structure within the genus Brassica
Two types of molecular markers, SSR and AFLP, were used to analyze the genetic structure at the whole genome level of 49 accessions, which represented natural variants of B. napus, B. rapa and the B. oleracea cytodeme. In total, 355 polymorphic bands from 11 AFLP primer pairs and 464 polymorphic bands from 83 SSR primer pairs were scored. On average, 32 polymorphic bands were detected for each AFLP primer pair among the 49 accessions, whereas an average of 5.2, 2.0 and 2.8 alleles were detected for each SSR primer pair in the B. oleracea cytodeme, B. rapa and B. napus, respectively. Among the three species of Brassica tested, the highest genetic diversity was found in the B. oleracea cytodeme, with an average genetic distance of 0.59 obtained with the SSR markers and 0.32 with the AFLP markers, followed by B. rapa (0.33 with SSR and 0.25 with AFLP markers) and B. napus (0.24 with SSR and 0.13 with AFLP markers).
Although the genetic variation among the 49 accessions of Brassica could be detected more sensitively with the SSR than with the AFLP markers, a high degree of correlation (r=0.963, P<0.01) was found between the genetic distances obtained with the SSR and AFLP markers. Therefore, the polymorphic SSR and AFLP bands were combined to analyze the genetic structure. Clustering analysis was performed among the 49 accessions using NTSYS software and the results are shown in Figure 1. Bootstrap values above 50%, which were calculated with 1000 replications, are shown on the branches. At a genetic distance of 0.51, the accessions of the B. oleracea cytodeme were classified into three subgroups, whereas the accessions of B. napus and B. rapa were clustered separately into individual groups with high-bootstrap values. All the cultivated types of B. oleracea were clustered into group C-I together with the wild-types B. incana, B. bourgeaui, B. montana, B. oleracea ssp. oleracea and B. cretica. Group C-II was comprised of the accessions of B. rupestris, B. macrocarpa, B. villosa and B. insularis, whereas B. hilarionis was categorized as group C-III (Figure 1).
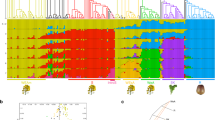
The findings of the cluster analysis were supported by the analysis of ancestry among the 49 accessions of Brassica (Figure 2). Owing to the maximal log likelihood obtained, the number of populations (K) was fixed at three by running the STRUCTURE software for 10 000 iterations after a burn-in of 1000 iterations. It was found that all accessions of B. rapa had a common ancestry, and that the accessions of the B. oleracea cytodeme could be classified into two different populations, with the exception of B. hilarionis, which shared a few components with the ancestry of B. rapa (Figure 2). The cultivated types of B. oleracea had the same ancestry as the wild types in group C-I, whereas the wild types in group C-II shared a different ancestry, although some wild accessions apparently had mixed ancestry. It was interesting that the ancestry of natural rapeseed corresponded to that of B. rapa and group C-I, which supports the view that rapeseed originated from B. rapa and B. oleracea.
Ancestry of 49 accessions of Brassica estimated with the STRUCTURE ver. 2.2 software. The number of ancestries of Brassica was fixed at three because of the maximal log likelihood obtained by running STRUCTURE for 10 000 iterations after a burn in of 1000 iterations. The ancestries of the Brassica species are indicated by bars with three different colors. The code and classification of the accessions are shown in Figure 1, and the roman numerals I, II, III, IV and V correspond to groups C-I, C-II, C-III, A and AC, respectively.
Comparison of the C subgenome in natural B. napus with the B. oleracea cytodeme
Given that an allopolyploid contains the entire set of chromosomes from each parental species and shows stable disomic chromosome pairing behavior (Chen 2007), the progenitors of an allopolyploid can be deduced by comparing the genotype of the natural allopolyploid with that of virtual allopolyploid lines synthesized from all variants of the parental species. The genotype of a virtual line can be derived from those of its parental lines, which are determined using DNA molecular markers.
We investigated the genetic relationships between the C subgenome of natural rapeseed and the genomes of the B. oleracea cytodeme. A set of 156 virtual allopolyploid lines was synthesized from 4 B. rapa accessions and 39 accessions of the B. oleracea cytodeme, which represented natural variants of the parental species. The genetic structure of all virtual lines and their parents was analyzed by principal component analysis, together with that of 6 accessions of rapeseed that represented natural variants of rapeseed, and the results are shown in Figure 3. The total variation explained by the first and second principal components was 48.3 and 27.0%, respectively. The virtual and natural B. napus lines were grouped in the middle of the figure, whereas the accessions of B. rapa and the B. oleracea cytodeme were clustered separately in the upper left and lower right corners, respectively. With respect to genetic diversity within the parental species of the B. oleracea cytodeme, the accessions from group C-I (indicated in Figure 1) were clustered closely together, whereas the other wild types were more separated. With respect to the genetic diversity of rapeseed, all six natural accessions from the three distinct gene pools of rapeseed (European winter, Chinese semi-winter and European spring rapeseed) were clustered closely with the virtual lines derived from the accessions in group C-I.
Associations among natural rapeseed, virtual rapeseed and parental lines as revealed by principal component analysis. The classification of the accessions is shown in Figure 1. Open triangles, closed triangles in light grey and closed triangles in dark grey represent the B. oleracea accessions from groups C-I, C-II and C-III, respectively; open circles, closed circles in light grey and closed circles in dark grey represent the virtual rapeseed lines derived from accessions of groups C-I, C-II and C-III, respectively; open squares represents accessions of B. rapa; and open stars represent accessions of natural B. napus.
The substantial contribution of the B. oleracea cytodeme to the genetic diversity of the virtual rapeseed lines was supported by the results of the F-test. When the virtual lines were categorized according to the four B. rapa parental lines, no significant differences were found among them (F3,932=1.10, P=0.35), whereas significant differences were detected among the virtual lines when they were categorized according to the subgroups of the B. oleracea parent (F2,933=567.8, P<0.001). This indicates that there is great potential for broadening the genetic diversity of rapeseed using the B. oleracea cytodeme.
Table 1 shows the average genetic distances between natural rapeseed and the B. oleracea cytodeme or virtual rapeseed lines. A high and significant correlation was found for the genetic distance between natural rapeseed and the B. oleracea cytodeme or the virtual rapeseed lines (r=0.96, P<0.001). It was obvious that natural rapeseed was closer to the virtual rapeseed lines, especially the lines derived from group C-I, than to the B. oleracea cytodeme. These results indicate that the different types in group C-I or their progeny may be the ancestral donors of the C subgenome in rapeseed.
Discussion
Virtual allopolyploid lines
In the present study, the strategy of virtual allopolyploidy was used successfully to investigate the ancestral donors of the C subgenome in rapeseed. Our findings support the previous findings that the origin of rapeseed occurs on more than one occasion (Song and Osborn, 1992; Allender and King, 2010), and that the C subgenome of rapeseed is related to cultivated B. oleracea (Olsson, 1954; Naughton, 1976; Song and Osborn, 1992; Gómez-Campo and Prakash, 1999) or B. montana (Song and Osborn, 1992). In addition, our data suggest that the potential progenitors of the C subgenome of rapeseed can be extended to include other wild types, such as B. incana, B. bourgeaui, B. oleracea ssp. oleracea and B. cretica. Our findings with respect to rapeseed support the suitability of this strategy to identify the progenitors of an allopolyploid.
The success of the strategy of using virtual lines to investigate the progenitors of allopolyploids is due to the following two factors: (1) in theory, any natural variants of parental species can be used to construct a virtual allopolyploid line without performing interspecific crossing; and (2) the majority of genotypes can be deduced reciprocally between the natural allopolyploid and the progenitors, and between the virtual allopolyploid and its parental lines. Compared with other strategies to explore the evolution of allopolyploidy, for example, DNA sequence analyses (Petersen et al., 2006; Albach, 2007) and the use of cytoplasmic molecular markers (Song and Osborn, 1992; Allender and King, 2010), the strategy used in this study has several advantages. For example, it saves both time and money, and all variants of the parental species can be compared extensively with the natural allopolyploid at the DNA level without the need for interspecific hybridization.
None of the virtual allopolyploid lines generated in this study were found to match perfectly with natural B. napus. Similar findings have been reported for natural B. napus and synthetic allopolyploids (Becker et al., 1995; Seyis et al., 2003). A possible explanation is that the allopolyploid has evolved post-polyploidization. Synthesized allopolyploids of Brassica have been found to show rapid genetic alteration and phenotypic instability relative to their parental species, especially during early generations (Song et al., 1995; Prakash et al., 1999; Gaeta et al., 2007), and natural and artificial selection might increase the genetic differences between an allopolyploid and its parental species (Song et al., 1988). Moreover, we only used a small number of B. rapa lines from two gene pools (Qian et al., 2003; Zhao et al., 2005), because we were focusing on the C subgenome of B. napus. This resulted in a limited variation in the A subgenome of the virtual allopolyploids, and therefore might have prevented the finding of a perfect match between natural B. napus and a synthetic line.
Widening the genetic variance of allopolyploid crops
Currently, there is a tension between the great demands on agricultural productivity to meet the increase in the human population and the fact that agricultural inputs, such as fertilizers, need to be reduced in order to preserve the environment. However, traditional plant-breeding processes threaten the genetic base on which breeding depends, because new varieties are usually derived from crosses among genetically-related modern varieties, and primitive ancestors, which are genetically more variable, but less productive, are excluded (Tanksley and McCouch, 1997). These ancestors are important sources for crop improvement because they can provide beneficial alleles or genes (Xiao et al., 1996; Fridman et al., 2004; He et al., 2006), and widen the genetic base (Seyis et al., 2003; Reif et al., 2005; Qian et al., 2006).
In this study, natural accessions of rapeseed from three divergent gene pools (Diers and Osborn, 1994; Becker et al., 1995; Qian et al., 2006) clustered together when compared with virtual rapeseed lines and parental species in principal component analysis. This indicates that the amount of variation in natural rapeseed is small compared with that in the parental species, and that there is great potential to widen the genetic base of natural rapeseed by using the parental species. The B. oleracea cytodeme could be of particular importance because it was shown to contribute strongly to the genetic diversity of the virtual rapeseed lines. Our data suggest that it might be more effective to broaden the diversity of the C subgenome of rapeseed using wild types of B. oleracea, such as B. macrocarpa, B. rupestris, B. villosa, B. insularis and B. hilarionis, which are genetically distant from the C subgenome of natural rapeseed, rather than by using members of group C-I.
References
Abdalla A, Reddy O, El-Zik K, Pepper A (2001). Genetic diversity and relationships of diploid and tetraploid cottons revealed using AFLP. Theor Appl Genet 102: 222–229.
Albach D (2007). Amplified fragment length polymorphisms and sequence data in the phylogenetic analysis of polyploids: multiple origins of Veronica cymbalaria (Plantaginaceae). New Phytol 176: 481–498.
Allender C, King G (2010). Origins of the amphiploid species Brassica napus L investigated by chloroplast and unclear molecular markers. BMC Plant Biol 10: 54.
Becker H, Engqvist G, Karlsson B (1995). Comparison of rapeseed cultivars and resynthesized lines based on allozyme and RFLP markers. Theor Appl Genet 91: 62–67.
Chen Z (2007). Genetic and epigenetic mechanisms for gene expression and phenotypic variation in plant polyploids. Annu Rev Plant Biol 58: 377–406.
Comai L, Tyagi A, Winter K, Holmes-Davis R, Reynolds S, Stevens Y et al. (2000). Phenotypic instability and rapid gene silencing in newly formed arabidopsis allotetraploids. Plant Cell 12: 1551–1568.
Diers B, Osborn T (1994). Genetic diversity of oilseed Brassica napus germ plasm based on restriction fragment length polymorphisms. Theor Appl Genet 88: 662–668.
Falush D, Stephens M, Pritchard J (2007). Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7: 574–578.
Fridman E, Carrari F, Liu Y, Fernie A, Zamir D (2004). Zooming in on a quantitative trait for tomato yield using interspecific introgressions. Science 305: 1786–1789.
Gaeta R, Pires J, Iniguez-Luy F, Leon E, Osborn T (2007). Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. Plant Cell 19: 3403–3417.
Gómez-Campo C, Prakash S (1999). Origin and domestication. In:Gómez-Campo C (eds) Biology of Brassica Coenospecies. Elsevier: The Netherlands, pp 33–58.
Harberd D (1976). Cytotaxonomic studies of Brassica and related genera. In:Vaughan J, Macleod A, Jones B (eds) The Biology and Chemistry of the Cruciferae. Academic Press: London, pp 47–68.
He G, Luo X, Tian F, Li K, Zhu Z, Su W et al. (2006). Haplotype variation in structure and expression of a gene cluster associated with a quantitative trait locus for improved yield in rice. Genome Res 16: 618–626.
Kashkush K, Feldman M, Levy A (2002). Gene loss, silencing and activation in a newly synthesized wheat allotetraploid. Genetics 160: 1651–1659.
Kianian S, Quiros C (1992). Trait inheritance, fertility, and genomic relationships of some n=9 Brasscia species. Genet Resour Crop Ev 39: 165–175.
Lukens L, Pires J, Leon E, Vogelzang R, Oslach L, Osborn T (2006). Patterns of sequence loss and cytosine methylation within a population of newly resynthesized Brassica napus allopolyploids. Plant Physiol 140: 336–348.
Masterson J (1994). Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264: 421–424.
Naughton M (1976). Swedes and Japanese Brassica napus (Cruciferae). In: Simmonds N (ed). Evolution of Crop Plants. Lougman Inc: NY.
Nei M, Li W (1979). Mathematical model for studying genetic variation in terms of restriction endouncleases. Proc Natl Acad Sci USA 76: 5269–5273.
Olsson G (1954). Crosses between Brassica napus and Japanese Brassica napella. Hereditas 40: 243–252.
Otto S (2007). The evolutionary consequences of polyploidy. Cell 131: 452–462.
Ozkan H, Levy A, Feldman M (2001). Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops-Triticum) group. Plant Cell 13: 1735–1747.
Parkin I, Sharpe A, Lydiate D (2003). Patterns of genome duplication within the Brassica napus genome. Genome 46: 291–303.
Petersen G, Seberg O, Yde M, Berthelsen K (2006). Phylogenetic relationships of Triticum and Aegilops and evidence for the origin of the A, B, and D genomes of common wheat (Triticum aestivum). Mol Phylogenet Evol 39: 70–82.
Prakash S, Takahata Y, Kirti P, Chopra V (1999). Cytogenetics. In:Gómez-Campo C (ed) Biology of Brassica Coenospecies. Elsevier: The Netherlands, pp 59–106.
Qian W, Liu R, Meng J (2003). Genetic effects on biomass yield in interspecific hybrids between Brassica napus and B. rapa. Euphytica 134: 9–15.
Qian W, Meng J, Li M, Frauen M, Sass O, Noack J et al. (2006). Introgression of genomic components from Chinese Brassica rapa contributes to widening the genetic diversity in rapeseed (B napus L), with emphasis on the evolution of Chinese rapeseed. Theor Appl Genet 113: 49–54.
Rana D, Van Den Boogaart T, O'neill C, Hynes L, Bent E, Macpherson L et al. (2004). Conservation of the microstructure of genome segments in Brassica napus and its diploid relatives. Plant J 40: 725–733.
Reif J, Zhang P, Dreisigacker S, Warburton M, Van Ginkel M, Hoisington D et al. (2005). Wheat genetic diversity trends during domestication and breeding. Theor Appl Genet 110: 859–864.
Rohlf F (1997). NTSYS-PC 2.1. Numerical taxonomy and multivariate analysis system. In: Setauket, editor. Exeter Software: NY.
Rousseau-Gueutin M, Lerceteau-Kohler E, Barrot L, Sargent D, Monfort A, Simpson D et al. (2008). Comparative genetic mapping between octoploid and diploid Fragaria species reveals a high level of colinearity between their genomes and the essentially disomic behavior of the cultivated octoploid strawberry. Genetics 179: 2045–2060.
Saghai-Maroof M, Soliman K, Jorgensen R, Allard R (1984). Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81: 8014–8018.
SAS Institute (1992). SAS technical report. SAS statistics software: changes and enhancements. Release 6.07 ed. Cary.
Schiemann E (1932). Entstehung der Kulturplanzen. Handb Vereb Wis 3: 271–288.
Seyis F, Snowdon R, Luhs W, Friedt W (2003). Molecular characterization of novel resynthesized rapeseed (Brassica napus) lines and analysis of their genetic diversity in comparison with spring rapeseed cultivars. Plant Breed 122: 473–478.
Sinskaia E (1928). The oleiferous plants and root crops of the family Cruciferae. Bull Appl Bot Genet Plant Breed 9: 1–648.
Snogerup S, Gustafsson M, Von Bothmer R (1990). Brassica sect. Brassica (Brassicaceae)I Taxonomy and variation. Willdenowia 19: 271–365.
Soltis P, Soltis D (2000). The role of genetic and genomic attributes in the success of polyploids. Proc Natl Acad Sci USA 97: 7051–7057.
Song K, Lu P, Tang K, Osborn T (1995). Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92: 7719–7723.
Song K, Osborn T (1992). Polyphyletic origins of Brasscia napus: new evidence based on organelle and nuclear RFLP analyses. Genome 35: 992–1001.
Song K, Osborn T, Williams P (1988). Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs) 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75: 784–794.
Stebbins G (1971). Chromosomal Evolution in Higher Plants. Edward Arnold: London.
Tanksley S, Mccouch S (1997). Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277: 1063–1066.
UN (1935). Genomic analysis in Brassica with special reference to the experimental formation of B napus and peculiar bode of fertilization. Jpn J Bot 7: 389–452.
Von Bothmer R, Gustafsson M, Snogerup S (1995). Brassica sect. Brassica (Brassicaceae) II. Inter- and intraspecific crosses with cultivars of B. oleracea. Genet Resour Crop Ev 42: 165–178.
Winge O (1917). The chromosomes: their number and general importance. CR Trav Lab Carlsberg 13: 131–275.
Wolfe K (2001). Yesterday's polyploidization and the mystery of diploidization. Nat Rev Genet 2: 333–341.
Wood T, Takebayashi N, Barker M, Mayrose I, Greenspoon P, Rieseberg L (2009). The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106: 13875–13879.
Xiao J, Grandillo S, Ahn S, Mccouch S, Tanksley S, Li J et al. (1996). Genes from wild rice improve yield. Nature 384: 223–224.
Yap I, Nelson R (1996). WINBOOT, a program for performing bootstrap analysis of binary data to determine the confidence limits of UPGMA-based dendrograms. International Rice Research Institute: Manila, Philippines.
Zhao J, Wang X, Deng B, Lou P, Wu J, Sun R et al. (2005). Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor Appl Genet 110: 1301–1314.
Acknowledgements
We thank the CGN in The Netherlands, IPK in Germany, UPM in Spain, Prof S Snogerup from UL in Sweden and Prof C Quiros from UC Davis for kind support in the collection of materials. We thank Mr JO Disi for reading the manuscript critically. This work was supported financially by grants from NPZ Company, 863 Project (2006AA100106), by the Natural Science Foundation Project of Chongqing (2007BB1362), by the Ministry of Education of China, by the Ministry of Personnel of China to W Qian and by Southwest University Scientific and Technological Innovation Fund and Graduates (kb2009006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mei, J., Li, Q., Qian, L. et al. Genetic investigation of the origination of allopolyploid with virtually synthesized lines: Application to the C subgenome of Brassica napus. Heredity 106, 955–961 (2011). https://doi.org/10.1038/hdy.2010.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2010.143
Keywords
This article is cited by
-
Investigating genetic relationship of Brassica juncea with B. nigra via virtual allopolyploidy and hexaploidy strategy
Molecular Breeding (2021)
-
Co-linearity and divergence of the A subgenome of Brassica juncea compared with other Brassica species carrying different A subgenomes
BMC Genomics (2016)
-
Effective accumulated temperature is associated with the efficiency of hybrid ovary culture between Brassica napus and B. oleracea
Acta Physiologiae Plantarum (2015)
-
Sub-genomic selection patterns as a signature of breeding in the allopolyploid Brassica napus genome
BMC Genomics (2014)
-
Improvement of Brassica napus via interspecific hybridization between B. napus and B. oleracea
Molecular Breeding (2014)