- NEWS AND VIEWS
A stockpile of antiviral defences
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 556, 318-319 (2018)
doi: https://doi.org/10.1038/d41586-018-04367-y
References
Samson, J. E., Magadán, A. H., Sabri, M. & Moineau, S. Nature Rev. Microbiol. 11, 675–687 (2013).
Doron, S. et al. Science 359, eaar4120 (2018).
Labrie, S. J., Samson, J. E. & Moineau, S. Nature Rev. Microbiol. 8, 317-327 (2010).
Snyder, L. Mol. Microbiol. 15, 415–420 (1995).
Pingoud, A., Wilson, G. G. & Wende, W. Nucleic Acids Res. 42, 7489–7527 (2014).
Barrangou, R. et al. Science 315, 1709–1712 (2007).
Jinek, M. et al. Science 337, 816–822 (2012).
Paez-Espino, D. et al. Nature 536, 425–430 (2016).
Makarova, K. S., Wolf, Y. I., Snir, S. & Koonin, E. V. J. Bacteriol. 193, 6039–6056 (2011).
Kauffman, K. M. et al. Nature 554, 118–122 (2018).
Dupuis, M.-È., Villion, M., Magadán, A. H. & Moineau, S. Nature Commun. 4, 2087 (2013).

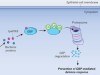 Bacteria disarm host-defence proteins
Bacteria disarm host-defence proteins
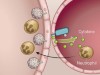 Melanin triggers antifungal defences
Melanin triggers antifungal defences