Abstract
Par-4 is a unique proapoptotic protein with the ability to induce apoptosis selectively in cancer cells. The X-ray crystal structure of the C-terminal domain of Par-4 (Par-4CC), which regulates its apoptotic function, was obtained by MAD phasing. Par-4 homodimerizes by forming a parallel coiled-coil structure. The N-terminal half of Par-4CC contains the homodimerization subdomain. This structure includes a nuclear export signal (Par-4NES) sequence, which is masked upon dimerization indicating a potential mechanism for nuclear localization. The heteromeric-interaction models specifically showed that charge interaction is an important factor in the stability of heteromers of the C-terminal leucine zipper subdomain of Par-4 (Par-4LZ). These heteromer models also displayed NES masking capacity and therefore the ability to influence intracellular localization.
Similar content being viewed by others
Main
Par-4 is a 332 amino-acid proapoptotic protein with tumor suppressor activity. Par-4 had been predicted to be largely disordered.1 Its downregulation or non-functional state has been linked to various cancers.2 In contrast, elevated levels of Par-4 have been observed in neurons affected by various neurodegenerative diseases3, 4, 5 and, similarly, in the pathogenesis of HIV encephalitis.6 The most striking feature of Par-4 is that its ectopic expression induces apoptosis selectively in certain cancer cells but not in normal or immortalized cells. Instead, it sensitizes the latter to various apoptotic stimuli.7 Par-4 has been involved in both the extrinsic/death receptor-mediated and the intrinsic/mitochondrial pathways of apoptosis.8 Nevertheless, Par-4 is ubiquitously expressed with few exceptions and evolutionarily conserved in vertebrates,9 indicating that the endogenous Par-4 under normal conditions is inactive.
The N-terminal half of Par-4 contains the SAC domain (residues 137–195), which selectively induces apoptosis in cancer cells. It is sufficient for the activation of the Fas pathway, inhibition of NF-κB activity and apoptotic induction.10 However, the latter two functions require its nuclear translocation and phosphorylation at Thr1558 by protein kinase A (PKA)11 or death-associated protein-like kinase (Dlk/DAPK3).12 The SAC domain includes a nuclear localization sequence (residues 137–153), which is essential for the nuclear translocation of Par-4.10 Between the SAC domain and the C-terminal region of Par-4, a phosphorylation site for Akt1 is located, that is, Ser249. It acts as a general prosurvival switch and sequesters Par-4 to the cytoplasm upon phosphorylation.13
The C-terminal region of Par-4 (Par-4CC, residues 254–332) was predicted to form a coiled coil1 and was shown to sensitize cells to various apoptotic stimuli.14 It was suggested to contain a leucine zipper domain (Par-4LZ, residues 292–332) and a nuclear export sequence (NES).1, 8 This Par-4CC region was mainly shown to mediates the interaction with partner proteins such as PKCζ,15 WT1,16 Akt1,13 apoptosis antagonizing transcription factor (AATF)/Che-1,17 DAPK3,18 Amida,19 THAP120 and p62.21 Par-4 binds and inhibits the activity of PKCζ and thereby NF-κB activity, whereas p62 antagonizes this inhibition by forming a ternary complex.21 Par-4 regulates the processing of the β-amyloid precursor protein, which is blocked by the interaction with AATF.17 Par-4 interacts with DAPK3 and Amida, respectively, whereupon coexpression it initiates their relocation from the nucleus to colocalize at the cytoskeleton causing the cytoskeletal reorganization and apoptosis induction.18, 19 Par-4 interaction with WT1 transcriptionally downregulates BCL-2 and thereby induces apoptosis.8 Par-4 interacts with THAP1 (a promyelocytic leukemia nuclear bodies (PML-NBs)-associated protein), which contains a DNA-binding THAP domain.20 In spite of these interactions, some being exclusively nuclear, Par-4CC is not necessary for the direct induction of apoptosis.10 The aforementioned features of Par-4 point to the C-terminal region of Par-4 as a regulatory domain, which interacts with a multitude of proteins forming an obscure signaling network.
We have determined the X-ray crystal structure of Par-4CC to determine its subdomains. Their regulatory interactions with AATF, Amida, DAPK3 and THAP1 have been analyzed by modeling to discover the structural basis for the formation of stable heteromeric complexes and control of subcellular localization.
Results
Structure of Par-4CC domain confirms homodimerization
The structure of Par-4CC was determined by multiwavelength anomalous dispersion from a selenomethionine-labeled Par-4CC (SeMet-Par-4CC) at 3.0 Å resolution to an Rwork/Rfree of 24.54%/29.44% (Table 1). The asymmetric unit contains seven helical molecules (chains A, B, C, D, E, F and G) forming three homodimeric structures (dimers AB, CD and FG) (Figure 1). A fourth dimer is generated by a crystallographic symmetry (dimer EE′). One of the non-crystallographic homodimers (dimer FG) displayed electron density for their C-terminal halves lacking connectivity, which did not allow completion of the model. Our result confirms the yeast two-Hybrid assay showing the self-association of Par-4 via its Par-4CC region.15
Asymmetric unit of the Par-4CC crystal. Representation of the asymmetric unit of the SeMet-labeled Par-4CC crystal. The asymmetric unit contains seven molecules, which are shown with different colors. Each of them forms homodimeric coiled-coil structures. Chain E forms a crystallographic homodimer with a symmetry equivalent molecule
Par-4CC homodimerizes forming parallel coiled-coil structure
The structure of Par-4CC includes the C-terminal 79 residues, of which the residues 257–330 form a canonical coiled-coil structure with two right-handed helices running parallel to each other having a left-handed super helical twist. Also, the diffraction images of Par-4CC crystals displayed diffuse scattering characteristic of coiled-coil (Supplementary Figure S1) (see also Supplementary Results, and Supplementary Figure S2) formation of such coiled-coil structures are guided by heptad repeats in their sequence designated (abcdefg)n, with positions a and d being predominantly occupied by hydrophobic amino acids. Analyzing the structure by SOCKET,22 11 heptad repeats can be assigned to residues 257–330, with only four residues in the eleventh heptad (Figure 2a). The three asparagines of the homodimeric coiled-coil of Par-4CC (Asn278, Asn313 and Asn320 of non-crystallographic dimers), each in position a of their respective heptads, show two rotamer conformations breaking the twofold non-crystallographic symmetry locally (Figure 2b). The structure is in agreement with the observation that the presence of Asn in position a of a heptad favors dimerization with parallel packing of helices.23, 24 The Cα–Cβ bonds of the residues at positions a and d of each heptad are oriented parallel and perpendicular, respectively, to the peptide bonds facing each of them in the opposite helix, as shown previously for two-stranded parallel coiled coils.25 As expected, the structure shows the side chains at positions a and d facing the interhelical space of the coiled coil (Figure 2b). These residues being predominantly (73%) hydrophobic constitute the hydrophobic dimer interface.
Homodimeric interactions of Par-4CC. (a) Structure-based heptad register of the protein sequence of Par-4CC as assigned by the SOCKET program. Double-headed arrows depict salt bridges: intrahelical salt bridge (black) and interhelical salt bridge (magenta). The red double-headed arrow represents the interhelical charge repulsion. Sequence numbering is as per the UniProt id: Q62627. The region of Par-4CC corresponding to the homodimerization and leucine zippers domains are depicted. (b) Chain representation of the crystal structure of the two domains of the homodimeric coiled-coil of Par-4CC: homodimerization domain (left) and leucine zipper domain (right). Side chains of the residues occupying the coiled-coil interface are shown in crimson color for chain A (ice blue) and cyan for chain B (gold). The interhelical and intrahelical salt bridges are depicted. For clarity only one set of residues involved in salt-bridge formation are shown with their side chains in atom-type representation. Asparagines (blue) occupying the heptad positions a in both the domains, and the residues at every heptad positions d of the leucine zipper domain are labeled. Electron density (2mFo-DFc type) is contoured at 1.0σ. See also Supplementary Table S1
The coiled-coil structure of Par-4CC also includes Par-4LZ (Figure 2b). In the LZ motifs, the position d of their heptads is predominantly occupied by leucine.26 Such structures are known to dimerize through coiled-coil formation.27 The last six heptads of Par-4CC belong to Par-4LZ featuring four hydrophobic and two polar residues at position a (Ile292, Ile299, Leu306, Asn313, Asn320, Val327), and exclusively leucine residues at position d (Leu295, Leu302, Leu316, Leu323, Leu330), with the exception of the eighth heptad containing a methionine (Met309) (Figure 2b).
N-terminal half of Par-4CC displays stabilizing factors for the homodimer
Coiled-coil structures gain stability via interhelical salt bridges,28 while intrahelical salt bridges can stabilize monomeric α-helices.29, 30 The structure of Par-4CC shows both interhelical and intrahelical salt bridges of g–e′, d–e′ and g–c type (Figure 2b). The interactions Arg263-Glu268′, Glu270-Arg275′ and Glu312-Lys317′ belong to the g–e′ type, whereas Glu274-Arg275′ and Lys288-Glu289′ belong to the d–e′ type (Figure 2b). There are two intrahelical salt bridges of the g–c type, that is, Glu270-Arg273 and Glu277-Arg280 (Figure 2b).
The N-terminal halves (residues 257–291) of the helices display systematically lower disorder than the C-terminal halves (residues 292–330) (Supplementary Table S1), which resulted in relatively poor map quality for the C-terminal parts of the molecules. Although all the helices displayed disorder to some extent, chains A–E are relatively less disordered than the helices (chains F and G) with an average B-value of ≈69 and ≈107 Å2, respectively (Supplementary Table S1).
The N-terminal half of the structure with the aforementioned salt bridges displayed significantly more favorable interactions between charged residues than the C-terminal half (Figure 2b). In contrast, Par-4LZ with only one stabilizing salt bridge depended mostly on the hydrophobic interface for stability of the homodimeric coiled-coil structure (Figure 2b). Weakening of side-chain electron density and increasing temperature factors were observed starting around residue 290 (Figure 2b) (Supplementary Table S1). This clearly indicated that the N-terminal half of Par-4CC constitutes the homodimerization subdomain (Par-4HDD).
Masking of the NES by dimerization
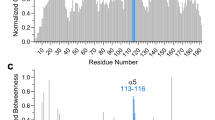
Par-4LZ was claimed to contain an NES8 and Par-4 displayed sensitivity towards selective inhibitors of nuclear export,31 indicating its NES-dependent nuclear export. The protein sequence analysis using NetNES32 and ValidNES33 suggests the residues 295–301 (LKEEIDL) as the possible NES (Par-4NES). The general NES consensus includes hydrophobic residues (shown in bold) that are essential for binding to the hydrophobic pockets of exportin-1/XPO1/CRM1.34 The homodimeric structure of Par-4CC shows the hydrophobic surface formed by Par-4NES being masked by dimerization (Figure 3), suggesting a mechanism to escape CRM1-mediated nuclear export by being inaccessible. It also displays relatively higher temperature factors for Par-4NES (Supplementary Table S1), which is also true for majority of the structures of NES.34
Structure of Par-4NES. Structure of predicted NES of Par-4CC. Electron density map of 2mFo-DFc type contoured at 0.8σ is shown. See also Supplementary Table S1
Modeling of Par-4CC regulatory interactions
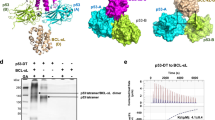
Docking analysis of Par-4CC using some of its binding partners, that is, AATF, Amida, DAPK3 and THAP1 was performed for parallel helical pairing to understand the regulatory mechanism of Par-4. Only the region of the sequence of the binding partners displaying high probability for the coiled-coil conformation was used for modeling and docking. The predicted coiled-coil regions overlap significantly with the reported binding regions of respective proteins with Par-4 (Supplementary Table S2). Three of the four interacting partners, that is, AATF, Amida and DAPK3 showed interaction in the Par-4LZ region, whereas THAP1 showed the interaction in the Par-4HDD region with six interhelical salt bridges (Figure 4). This strong interaction with THAP1 could explain the colocalization of Par-4 in the PML-NBs. The model of DAPK3 used for docking did not include the kinase's LZ region because Par-4 was shown to interact with the LZ deletion mutant of DAPK3.18 The interaction models of Par-4CC with AATF and DAPK3, which are known to interact with each other via their LZ motifs,35 displayed binding with Par-4LZ, respectively. Therefore, these models suggested a competition of AATF and DAPK3 for Par-4LZ. The interaction region of Par-4LZ in the Amida/Par-4LZ model included two leucine residues (Leu295 and Leu316) that were mutated in a leucine zipper triple mutant of Par-4 (Leu295Ala, Leu316Ala and Leu330Ala), which failed to interact with Amida.19
Par-4 coiled-coil interactions with binding partners. Left to right, surface representations of the crystal structure of homodimeric coiled-coil Par-4CC and its heterodimeric-interaction models with THAP1, AATF, Amida and DAPK3 (dashed box). Model surfaces are colored by electrostatic potential (ESP) on a sliding scale from blue (positive) to red (negative). For heterodimeric-interaction models, ESP surfaces are shown only for interacting regions. The black double-headed arrow in the Par-4CC homodimer represents the charge repulsion between Asp305 and Glu310′. Regions corresponding to the Par-4CC charge repulsion and Par-4NES are depicted with dashed lines in the heterodimeric-interaction models. Salt bridges are listed: underlined are the residues that show repulsion in Par-4CC homodimer. See also Supplementary Figure S3 and Supplementary Table S2
The interaction models of AATF, Amida and DAPK3 with Par-4CC covered the complete region of the Par-4NES, suggesting an NES masking in these heteromers, similar to that in the Par-4CC homodimer (Figure 4). The coiled-coil stability of Par-4LZ was found to depend strongly on pH, which was previously attributed to intermolecular charge repulsion between Asp305 and Glu310′ (g–e′ type) based on mutation analysis.36, 37 The region corresponding to this destabilizing g–e′ type charge repulsion of the Par-4CC homodimer in the former three interaction models did not show such repulsion (Figure 4). Moreover, the residues Asp305 and Glu310 of Par-4LZ were stabilized in DAPK3 containing model by forming salt bridges (Figure 4). Overall, the docking analysis suggested charge stabilization as an important factor for specific heteromerization of Par-4CC.
Discussion
Par-4HDD contains a likely trigger motif for homodimerization
Two-stranded parallel coiled-coil structures were shown to contain a trigger sequence, which represents an autonomous helical folding unit that is indispensable for coiled-coil formation.38 Based on the trigger sequences of different dimeric coiled-coil structures, a trigger motif consensus was derived, that is, xxLExc-hxcxccx, where x, h and c are any residue, hydrophobic residue and charged residue, respectively.38 These consensus sequences occur with little variations. Par-4HDD contains such consensus sequences twice (conserved residues in bold), that is, 258-STLEKR-IEDLEKE-270 and 265-EDLEKE-VLRERQE-277. They show less disorder than any other part of the Par-4CC structure (Supplementary Table S1). These features of Par-4HDD suggest that this region, which triggers the homodimeric coiled-coil interaction of Par-4, represents the most stable part of the structure. Moreover, the heteromeric-interaction model of THAP1 with Par-4CC shows also for THAP1 such a trigger consensus in the interaction region, that is, HQLEQQ-VEKLRKK. This region interacts directly with the second trigger consensus sequence of Par-4HDD (Supplementary Figure S3). In contrast, the Par-4- interacting proteins AATF, Amida and DAPK3 do not contain such a trigger sequence and exhibit dimerization only with Par-4LZ in the interaction models (Figure 4).
The presence of Asn at position a in two heptads of Par-4LZ (Asn313 and Asn320) should have a destabilizing effect besides giving specificity for the dimeric coiled-coil structure formation.24 Par-4HDD also contains such an Asn, that is, Asn278. Therefore, difference in the stability, respectively, apparent disorder, inferred from the structures of Par-4HDD and Par-4LZ (Supplementary Table S1) could be explained at least partially by the difference in the number of destabilizing Asn residues within these subdomains. Furthermore, Par-4HDD contains intrahelical salt bridges that stabilize monomeric α-helices and interhelical salt bridges that stabilize its homodimer (Figure 2b). Interestingly, the majority of these stabilizing salt bridges are located within the trigger sequences of Par-4HDD. The same is true for the trigger sequence in THAP1, which mimics in the heterodimer (Figure 4) the salt-bridge formation of the Par-4HDD homodimer (Figure 2a).
Preferential heterodimerization of Par-4LZ
Par-4LZ was shown to display an environment-dependent structural interconversion between a partially ordered monomer state and a predominantly coiled-coil dimer state.39 Despite the formation of a stable homodimer in the N-terminal segment of the Par-4CC, the presence of g–e′ type charge repulsion between Asp305-Glu310′ is likely causing instability in Par-4LZ of the homodimeric structure of Par-4CC. Clearly, the structure of Par-4LZ is only weakly stabilized by the hydrophobic interface in the homodimer. This suggests that more stable interactions involving the LZ domain may be formed with other proteins. This is supported by our analysis of heterodimeric-interaction models, where Par-4LZ showed heterodimeric coiled-coil interactions allowing 1–3 interhelical salt bridges and, importantly, no charge repulsions with AATF, Amida and DAPK3 (Figure 4). The interaction model of DAPK3 is an example of optimal heteromeric interaction, where the destabilizing repulsive g–e′ interaction of the Par-4CC homodimer is replaced by stabilizing interactions, that is, between Par-4Asp305-DAPK3Arg387 and Par-4Asp310-DAPK3Arg382 (Figure 4). Although the Par-4LZ homodimer has one interhelical salt bridge (Glu312-Lys317′), the g–e′ type repulsion and the presence Asn313 and Asn320 in the helical interface (Figure 2b) disfavor its homodimerization. In contrast, the heterodimeric models involving the Par-4LZ domain display more favorable interactions with regard to charge repulsion and salt bridges. Therefore, we conclude that Par-4LZ provides interactions with signaling proteins that are distinct from the binding of molecules by Par-4HDD. The detailed analysis of the molecular interaction of Par-4 from the structure of the homodimer and the models of the heterodimers shows that Par-4HDD contributes more important interactions than Par-4LZ. This conclusion is supported by the observation that the Par-4LZ domain itself is unfolded under physiological conditions, that is, inherently instable,36 but forms coiled-coil structure within Par-4.1
Regulation of complex formation by Par-4CC
The structure of Par-4CC along with the docking studies suggested a dual function of Par-4CC. One is homodimerization, which is mediated by Par-4HDD, and the second is heteromeric oligomerization, which is mainly mediated by Par-4LZ. Nevertheless, there is one case in our study where Par-4HDD is used for the heteromeric binding interaction with THAP1. This suggests that the THAP1/Par-4 complex is most likely to be heterodimeric, whereas the complexes with the other three proteins, that is, AATF, Amida and DAPK3, might be heterotetrameric assemblies that consist of a Par-4 homodimer formed by Par-4HDD, which binds interaction partners via Par-4LZ. For example, the interaction of Par-4 and DAPK3 is mediated via Par-4LZ and the predicted coiled-coil domain within the interacting region of DAPK3 (Supplementary Table S2). This interaction model is in accordance with the fact that DAPK3 does not need its LZ motif for interaction with Par-4 (Figure 4). In fact, the LZ domain of DAPK3 is needed for its homodimerization and kinase activation.40 Taken together, these considerations suggest that the putative complex of Par-4 and DAPK3 is a tetramer containing two intertwined homodimers (Figure 5). This model for the quaternary structure requires bending of the Par-4 helix. The appearance of high temperature factors after the Par-4HDD region starting around residue 290 for molecules A–E and the disorder for molecules F and G point at this region as the most likely area for bending. A closer inspection of the predicted coil-coil region of DAPK3 homodimer (residues 366–393) shows that there are repulsive g–e′ interaction between Arg382 and Arg387′, which are replaced by attractive interactions in the suggested heterotetramer (Figure 4). No structural information is available for the region of DAPK3 adjacent to its terminal LZ. Therefore, there is no clear picture of a possible interaction of DAPK3 with Par-4HDD and its preceding segment that connects the Par-4HDD region to the Par-4SAC domain (Figure 5).
Model of the Par-4-DAPK3 complex. Left, the homodimeric model of Par-4. Middle, homodimeric model of DAPK3 with kinase domain, predicted coiled-coil region (CC) and C-terminal LZ motif acting as dimerization domain. Right, suggested complex of Par-4 and DAPK3 where an active DAPK3 with its CC is bound to the LZ motif of Par-4 and its kinase domain is reaching out to the SAC domain of Par-4 to phosphorylate the Thr155 residue
Par-4 is known to show different localizations in different cell types,10 but the regulatory mechanism of its nuclear import/export is not described. Masking of Par-4NES was observed for the homodimeric structure of Par-4CC as well as the protein interaction models (Figures 3 and 4). The nuclear export of Par-4 by exportin-1 requires the Par-4NES being accessible. A masking of NES by oligomerization to regulate nuclear localization was already suggested for p53.41 Similarly, the masking of Par-4NES could also be such a regulatory mechanism to retain Par-4 in the nucleus to induce apoptosis by inhibiting NF-κB activity. Par-4-mediated recruitment of DAPK3 to the actin cytoskeleton in the cytoplasm is known,18 and the NES of Par-4 was suggested to assist in the relocation of the Par-4/DAPK3 complex.2 However, the formation of the complex between Par-4LZ and DAPK3 (Figure 4) would mask Par-4NES and block the translocation of the Par-4/DAPK3 complex to the cytoplasm. Therefore, it is more likely that the interaction of Par-4 with DAPK3 occurs in the cytoplasm.
In summary, there appears to be two very different modes of interaction of Par-4 with signaling proteins: (i) a dimerization via the HDD subdomain, which may appear in two different subtypes by binding of trigger motifs, (ii) a hetero-oligomerization by Par-4LZ binding and blocking of Par-4NES.
Materials and Methods
Production of SeMet-labeled Par-4CC
Details regarding Par-4CC plasmid construct and protein expression have been reported earlier,42 except for changes mentioned below. DYT medium of 750 ml with 200 μg/ml ampicillin was inoculated with a preculture to get an initial OD600 of 0.2 and allowed to grow in a shaker at 37 °C and 110 r.p.m. until it reached an OD600 of 0.8–1.2. The culture was centrifuged at 4000 r.p.m. for 15 min at 4 °C. The cell pellet was rinsed with 5–10 ml of SeMet minimal medium,43 resuspended gently in 20 ml of SeMet minimal medium and inoculated into 750 ml of SeMet minimal medium with 200 μg/ml ampicillin and 100 μg/ml l-SeMet. The culture was immediately induced with 1 mM isopropyl-β-D-thiogalactopyranoside and continued for 4 h before it was harvested and stored at −20 °C.
The SeMet-labeled Par-4CC (SeMet-Par-4CC) was purified as reported for native protein,42 and the protein sample was analyzed for labeling by mass spectroscopy. The SeMet incorporation was confirmed to be complete. The m/z values of the native and the SeMet-Par-4CC proteins were 12 258.401 and 12 402.916 Da, respectively, with a mass difference of 144.5 Da. This was in reasonable agreement with the theoretical value of 140.69 Da for replacement of the three methionines.
Crystallization of SeMet-Par-4CC
Crystallization of the purified SeMet-Par-4CC was performed with our reported crystallization condition:42 crystals were obtained using the condition 42% tert-butanol, 100 mM sodium citrate/citric acid, pH 5.9, within a week. Perfluoropolyether cryo oil (Hampton Research, Aliso Viejo, CA, USA) was overlaid on the crystallization drop immediately after opening the experiment to avoid evaporation for crystal mounting.
Data collection and processing
Multiwavelength anomalous dispersion (MAD) data were collected on beamline ID23-1 at the European Synchrotron Radiation Facility (ESRF) (Grenoble, France). Data measurements were performed at −173 °C. MAD data were collected at wavelengths of 0.9794 Å (peak), 0.9795 Å (inflection point), 0.977 Å (high-energy remote) and 0.9825 Å (low-energy remote), as determined from a fluorescence scan of the Se K edge.
Data were processed in space group P43212 till 3.0 Å using iMOSFLM44 and scaled using SCALA.45 Data collection statistics for SeMet-Par-4CC are summarized in Table 1. The number of molecules in the asymmetric unit was estimated to be seven using MATTHEWS_COEF (CCP4),46, 47 corresponding to a Matthews coefficient (VM) of 2.36 Å3/Da. Substructure solution and experimental phasing were carried out using PHENIX AutoSol wizard 48 (mean figure of merit (FOM) – 0.39) (Table 1) (Supplementary Figure S4) and SHELXCDE.49, 50 Sixteen SeMet sites out of the 21 expected for seven molecules were located. The starting model obtained by SHELXE (CC – 45.83%) consisted of helices. Structure refinement and model building were carried out using Phenix.refine,51 CNS 52 and Coot,53 respectively. Several rounds of simulated annealing, energy minimization and B-factor refinement were carried out using CNS before final refinement, which was performed using Phenix.refine. Torsion-based NCS restraints, which allow local differences between NCS-related molecules, were used for all chains during the refinements. The refinement of the final structure of Par-4 includes residues 254–331 for chains A and C, 255–232 for chains B and E, 256–332 for chain D and residues 255–294 for chains F and G. (For local differences between different chains in the asymmetric unit see Supplementary Results.) All the structure figures were prepared with CCP4mg.54
Because of the presence of crystallographic and non-crystallographic dimers, the data were analyzed for possible pathologies. Twinning in a polar tetragonal space group, which could create the EE′ dimer by the twinning operator, could be excluded. Measurements at room temperature were performed with native Par-4CC crystals on beamline P11 at PETRA III (DESY, Hamburg, Germany) as a control to investigate if cryocooling had any effect on the space group. Room temperature data measurement excluded cryocooling induced phase transformation.
Modeling
The protein sequence of the Par-4CC and its interacting proteins, that is, Amida (rat), AATF (human), DAPK3 (rat) and THAP1 (human), respectively, were analyzed by COILS/PCOILS,55 which calculates the probability of the sequence to adopt a coiled-coil conformation. The amino-acid sequences of those interacting proteins that showed relatively high probability for coiled-coil formation were used for model building using SWISS-MODEL.56, 57, 58 Par-4CC and the modeled proteins were docked using GRAMM-X,59 which is designed exclusively for docking pairs of protein molecules. Docked models with parallel pairing of helices in coiled-coil structures were analyzed (see Supplementary Table S2). The selected models were energy minimized by knowledge-based protein structure refinement using KoBaMIN.60 For protein model quality estimation, the minimized structures were submitted to QMEAN.61, 62 The number of salt bridges and the calculated length of coiled coil by SOCKET analysis were considered for selecting the model.
Accession number
Coordinates and structure factors for the SeMet-Par-4CC crystal have been submitted to the Protein Data Bank under accession code 5FIY.
References
Libich DS, Schwalbe M, Kate S, Venugopal H, Claridge JK, Edwards PJ et al. Intrinsic disorder and coiled-coil formation in prostate apoptosis response factor 4. FEBS J 2009; 276: 3710–3728.
Shrestha-Bhattarai T, Rangnekar VM . Cancer-selective apoptotic effects of extracellular and intracellular Par-4. Oncogene 2010; 29: 3873–3880.
Duan W, Guo Z, Mattson MP . Participation of par-4 in the degeneration of striatal neurons induced by metabolic compromise with 3-nitropropionic acid. Exp Neurol 2000; 165: 1–11.
Guo Q, Fu W, Xie J, Luo H, Sells SF, Geddes JW et al. Par-4 is a mediator of neuronal degeneration associated with the pathogenesis of Alzheimer disease. Nat Med 1998; 4: 957–962.
Pedersen WA, Luo H, Kruman I, Kasarskis E, Mattson MP . The prostate apoptosis response-4 protein participates in motor neuron degeneration in amyotrophic lateral sclerosis. FASEB J 2000; 14: 913–924.
Kruman II, Nath A, Maragos WF, Chan SL, Jones M, Rangnekar VM et al. Evidence that Par-4 participates in the pathogenesis of HIV encephalitis. Am J Pathol 1999; 155: 39–46.
Zhao Y, Rangnekar VM . Apoptosis and tumor resistance conferred by Par-4. Cancer Biol Ther 2008; 7: 1867–1874.
Ranganathan P, Rangnekar VM . Regulation of cancer cell survival by Par-4. Ann NY Acad Sci 2005; 1059: 76–85.
Boghaert ER, Sells SF, Walid AJ, Malone P, Williams NM, Weinstein MH et al. Immunohistochemical analysis of the proapoptotic protein Par-4 in normal rat tissues. Cell Growth Differ 1997; 8: 881–890.
El-Guendy N, Zhao Y, Gurumurthy S, Burikhanov R, Rangnekar VM . Identification of a unique core domain of par-4 sufficient for selective apoptosis induction in cancer cells. Mol Cell Biol 2003; 23: 5516–5525.
Gurumurthy S, Goswami A, Vasudevan KM, Rangnekar VM . Phosphorylation of Par-4 by protein kinase A is critical for apoptosis. Mol Cell Biol 2005; 25: 1146–1161.
Boosen M, Vetterkind S, Kubicek J, Scheidtmann KH, Illenberger S, Preuss U . Par-4 is an essential downstream target of DAP-like kinase (Dlk) in Dlk/Par-4-mediated apoptosis. Mol Biol Cell 2009; 20: 4010–4020.
Goswami A, Burikhanov R, de Thonel A, Fujita N, Goswami M, Zhao Y et al. Binding and phosphorylation of par-4 by akt is essential for cancer cell survival. Mol Cell 2005; 20: 33–44.
Sells SF, Han SS, Muthukkumar S, Maddiwar N, Johnstone R, Boghaert E et al. Expression and function of the leucine zipper protein Par-4 in apoptosis. Mol Cell Biol 1997; 17: 3823–3832.
Diaz-Meco MT, Municio MM, Frutos S, Sanchez P, Lozano J, Sanz L et al. The product of par-4, a gene induced during apoptosis, interacts selectively with the atypical isoforms of protein kinase C. Cell 1996; 86: 777–786.
Johnstone RW, See RH, Sells SF, Wang J, Muthukkumar S, Englert C et al. A novel repressor, par-4, modulates transcription and growth suppression functions of the Wilms' tumor suppressor WT1. Mol Cell Biol 1996; 16: 6945–6956.
Guo Q, Xie J . AATF inhibits aberrant production of amyloid beta peptide 1-42 by interacting directly with Par-4. J Biol Chem 2004; 279: 4596–4603.
Page G, Kogel D, Rangnekar V, Scheidtmann KH . Interaction partners of Dlk/ZIP kinase: co-expression of Dlk/ZIP kinase and Par-4 results in cytoplasmic retention and apoptosis. Oncogene 1999; 18: 7265–7273.
Boosen M, Vetterkind S, Koplin A, Illenberger S, Preuss U . Par-4-mediated recruitment of Amida to the actin cytoskeleton leads to the induction of apoptosis. Exp Cell Res 2005; 311: 177–191.
Roussigne M, Cayrol C, Clouaire T, Amalric F, Girard JP . THAP1 is a nuclear proapoptotic factor that links prostate-apoptosis-response-4 (Par-4) to PML nuclear bodies. Oncogene 2003; 22: 2432–2442.
Chang S, Kim JH, Shin J . p62 forms a ternary complex with PKCzeta and PAR-4 and antagonizes PAR-4-induced PKCzeta inhibition. FEBS Lett 2002; 510: 57–61.
Walshaw J, Woolfson DN . Socket: a program for identifying and analysing coiled-coil motifs within protein structures. J Mol Biol 2001; 307: 1427–1450.
Lee DL, Ivaninskii S, Burkhard P, Hodges RS . Unique stabilizing interactions identified in the two-stranded alpha-helical coiled-coil: crystal structure of a cortexillin I/GCN4 hybrid coiled-coil peptide. Protein Sci 2003; 12: 1395–1405.
Harbury PB, Zhang T, Kim PS, Alber T . A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants. Science 1993; 262: 1401–1407.
Lupas AN, Gruber M . The structure of alpha-helical coiled coils. Adv Protein Chem 2005; 70: 37–78.
Krylov D, Vinson CR . Leucine zipper. eLS 2001.
Rasmussen R, Benvegnu D, O'Shea EK, Kim PS, Alber T . X-ray scattering indicates that the leucine zipper is a coiled coil. Proc Natl Acad Sci USA 1991; 88: 561–564.
Meier M, Lustig A, Aebi U, Burkhard P . Removing an interhelical salt bridge abolishes coiled-coil formation in a de novo designed peptide. J Struct Biol 2002; 137: 65–72.
Burkhard P, Ivaninskii S, Lustig A . Improving coiled-coil stability by optimizing ionic interactions. J Mol Biol 2002; 318: 901–910.
Kammerer RA, Jaravine VA, Frank S, Schulthess T, Landwehr R, Lustig A et al. An intrahelical salt bridge within the trigger site stabilizes the GCN4 leucine zipper. J Biol Chem 2001; 276: 13685–13688.
Azmi AS, Aboukameel A, Bao B, Sarkar FH, Philip PA, Kauffman M et al. Selective inhibitors of nuclear export block pancreatic cancer cell proliferation and reduce tumor growth in mice. Gastroenterology 2013; 144: 447–456.
la Cour T, Kiemer L, Mølgaard A, Gupta R, Skriver K, Brunak S . Analysis and prediction of leucine-rich nuclear export signals. Protein Eng Des Sel 2004; 17: 527–536.
Fu SC, Huang HC, Horton P, Juan HF . ValidNESs: a database of validated leucine-rich nuclear export signals. Nucleic Acids Res 2013; 41: D338–D343.
Xu D, Farmer A, Collett G, Grishin NV, Chook YM . Sequence and structural analyses of nuclear export signals in the NESdb database. Mol Biol Cell 2012; 23: 3677–3693.
Page G, Lodige I, Kogel D, Scheidtmann KH . AATF, a novel transcription factor that interacts with Dlk/ZIP kinase and interferes with apoptosis. FEBS Lett 1999; 462: 187–191.
Dutta K, Alexandrov A, Huang H, Pascal SM . pH-induced folding of an apoptotic coiled coil. Protein Sci 2001; 10: 2531–2540.
Dutta K, Engler FA, Cotton L, Alexandrov A, Bedi GS, Colquhoun J et al. Stabilization of a pH-sensitive apoptosis-linked coiled coil through single point mutations. Protein Sci 2003; 12: 257–265.
Kammerer RA, Schulthess T, Landwehr R, Lustig A, Engel J, Aebi U et al. An autonomous folding unit mediates the assembly of two-stranded coiled coils. Proc Natl Acad Sci USA 1998; 95: 13419–13424.
Schwalbe M, Dutta K, Libich DS, Venugopal H, Claridge JK, Gell DA et al. Two-state conformational equilibrium in the Par-4 leucine zipper domain. Proteins 2010; 78: 2433–2449.
Kawai T, Matsumoto M, Takeda K, Sanjo H, Akira S . ZIP kinase, a novel serine/threonine kinase which mediates apoptosis. Mol Cell Biol 1998; 18: 1642–1651.
Stommel JM, Marchenko ND, Jimenez GS, Moll UM, Hope TJ, Wahl GM . A leucine-rich nuclear export signal in the p53 tetramerization domain: regulation of subcellular localization and p53 activity by NES masking. EMBO J 1999; 18: 1660–1672.
Tiruttani Subhramanyam UK, Kubicek J, Eidhoff UB, Labahn J . Cloning, expression, purification, crystallization and preliminary crystallographic analysis of the C-terminal domain of Par-4 (PAWR). Acta Crystallogr F 2014; 70: 1224–1227.
Guerrero SA, Hecht HJ, Hofmann B, Biebl H, Singh M . Production of selenomethionine-labelled proteins using simplified culture conditions and generally applicable host/vector systems. Appl Microbiol Biotechnol 2001; 56: 718–723.
Battye TG, Kontogiannis L, Johnson O, Powell HR, Leslie AG . iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr D 2011; 67: 271–281.
Evans PR . An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr D 2011; 67: 282–292.
Matthews BW . Solvent content of protein crystals. J Mol Biol 1968; 33: 491–497.
Kantardjieff KA, Rupp B . Matthews coefficient probabilities: improved estimates for unit cell contents of proteins, DNA, and protein–nucleic acid complex crystals. Protein Sci 2003; 12: 1865–1871.
Terwilliger TC, Adams PD, Read RJ, McCoy AJ, Moriarty NW, Grosse-Kunstleve RW et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr D 2009; 65: 582–601.
Schneider TR, Sheldrick GM . Substructure solution with SHELXD. Acta Crystallogr D 2002; 58: 1772–1779.
Sheldrick GM . Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr D 2010; 66: 479–485.
Afonine PV, Grosse-Kunstleve RW, Echols N, Headd JJ, Moriarty NW, Mustyakimov M et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr D 2012; 68: 352–367.
Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr D 1998; 54: 905–921.
Emsley P, Lohkamp B, Scott WG, Cowtan K . Features and development of Coot. Acta Crystallogr D 2010; 66: 486–501.
McNicholas S, Potterton E, Wilson KS, Noble ME . Presenting your structures: the CCP4mg molecular-graphics software. Acta Crystallogr D 2011; 67: 386–394.
Lupas A, Van Dyke M, Stock J . Predicting coiled coils from protein sequences. Science 1991; 252: 1162–1164.
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 2014; 42: W252–W258.
Bordoli L, Kiefer F, Arnold K, Benkert P, Battey J, Schwede T . Protein structure homology modeling using SWISS-MODEL workspace. Nat Protoc 2009; 4: 1–13.
Arnold K, Bordoli L, Kopp J, Schwede T . The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 2006; 22: 195–201.
Tovchigrechko A, Vakser IA . GRAMM-X public web server for protein–protein docking. Nucleic Acids Res 2006; 34: W310–W314.
Rodrigues JP, Levitt M, Chopra G . KoBaMIN: a knowledge-based minimization web server for protein structure refinement. Nucleic Acids Res 2012; 40: W323–W328.
Benkert P, Kunzli M, Schwede T . QMEAN server for protein model quality estimation. Nucleic Acids Res 2009; 37: W510–W514.
Benkert P, Tosatto SC, Schomburg D . QMEAN: a comprehensive scoring function for model quality assessment. Proteins 2008; 71: 261–277.
Acknowledgements
We acknowledge the European Synchrotron Radiation Facility for provision of synchrotron radiation facilities and we thank the local staff for assistance in using beamline ID23-1. The room temperature diffraction measurements were performed on beamline P11, DESY; we are also thankful to the local contact for providing assistance.
Author contributions
UBE, JK and UKTS isolated and crystallized the protein. UKTS generated and analyzed the interaction models. UKTS and JL performed crystallographic experiments, data analysis and completion of manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by P Czabotar
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Tiruttani Subhramanyam, U., Kubicek, J., Eidhoff, U. et al. Structural basis for the regulatory interactions of proapoptotic Par-4. Cell Death Differ 24, 1540–1547 (2017). https://doi.org/10.1038/cdd.2017.76
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2017.76
This article is cited by
-
Prostate apoptosis response-4 and tumor suppression: it’s not just about apoptosis anymore
Cell Death & Disease (2021)