Abstract
Background:
Sinonasal mucosal melanoma (SNMM) comprises <1% of all melanomas and lacks well-characterised molecular markers. Our aim was to determine the frequencies of common mutations and examine their utility as molecular markers in a large series of primary SNMMs.
Methods:
SNMM patients seen at our institution from August 1991 through July 2016 were identified. Genomic DNA was extracted from 66 formalin-fixed paraffin-embedded tumours and screened for mutations by direct sequencing. We investigated the association of mutations with clinicopathological features and survival outcomes.
Results:
Overall, 41% (27 out of 66) of the SNMMs harboured mutations. BRAF and KIT mutations were identified in 8% (five patients) and 5% (three patients) of SNMMs, respectively, whereas NRAS mutations were detected in 30% (20 patients) of SNMMs. Mutation rates in these oncogenes were similar between SNMMs located in the paranasal sinuses and those in the nasal cavity (30% and 13%, respectively, P=0.09). In a multivariate analysis, patients with negative margins had significantly better overall survival (hazard ratio 5.43, 95% confidence interval 1.44–21.85, P=0.01) and disease-specific survival (hazard ratio 21.9, 95% confidence interval 3.71–180, P=0.0004). The mutation status of the tumours showed no association with survival outcomes.
Conclusions:
In SNNM, mutation status does not affect survival outcomes, but NRAS mutations are relatively frequent and could be targeted in this disease by MEK inhibitors.
Similar content being viewed by others
Main
Mucosal melanoma represents approximately 1.3% of all melanomas (Gal et al, 2011). While mucosal melanoma can arise from any mucosa-lined body surface, approximately half of all mucosal melanomas occur in the head and neck, most frequently in the sinonasal cavity (Lourenco et al, 2014; Sun et al, 2014). Sinonasal mucosal melanomas (SNMMs) account for ∼4% of sinonasal malignancies and <1% of all melanomas (Moreno et al, 2010; Gal et al, 2011; Lourenco et al, 2014).
Sun exposure is a well-known risk factor for cutaneous melanoma, but the risk factors for SNMM are less well defined (Spencer and Mehnert, 2016). Patients usually present later in life, with no obvious sex predilection (Spencer and Mehnert, 2016). The mitogen-activated protein kinase (MAPK) pathway has been shown to be important in the development of melanoma (Curtin et al, 2005a). In cutaneous melanoma, between 22 and 72% of cases have BRAF mutations, and 0to 50% have NRAS mutations (Lee et al, 2011); however, molecular markers in mucosal melanoma are less well characterised. While recent studies suggest that BRAF inhibition has a promising effect in cutaneous melanoma, its role in SNMM has yet to be defined (Zebary et al, 2013a; Spagnolo et al, 2016).
SNMM is an aggressive tumour, and patients with SNMM often present with advanced disease (Ledderose and Leunig, 2015). Despite advances in treatment, survival is poor, with a 5-year survival rate of ∼20–30% (Moreno et al, 2010). Single-modality therapy with surgery is rarely adequate for this disease, particularly for SNMMs, in which anatomical and quality-of-life constraints make obtaining adequate margins very difficult and sometimes impossible (Samstein et al, 2016). Therefore, adjuvant therapy is a keystone in the treatment of SNMM. As more options arise for targeted therapy, the need to characterise molecular markers in SNMM has become increasingly important.
This study quantifies molecular features and attempts to identify molecular markers in SNMM. We also investigated the correlation of molecular features with clinicopathological features and survival outcomes to determine their prognostic utility in this disease.
Materials and methods
This retrospective review was approved by the Institutional Review Board at The University of Texas MD Anderson Cancer Center (Protocol RCR04-0636). We surveyed 170 consecutive patients seen at our institution from August 1991 through July 2016 with a pathologically confirmed diagnosis of head and neck mucosal melanoma involving the sinonasal cavity. The inclusion criteria for the analysis were: (1) pathologically confirmed mucosal melanoma; (2) sinonasal origin; (3) available outcome data; (4) available tissue for molecular analysis; and (5) adequate genetic material for analysis. Patient demographic features (age, sex, smoking status and alcohol intake), disease stage, tumour characteristics, treatment modalities used, pathological data (ulceration, perineural and lymphovascular invasion, bony invasion and number of mitotic figures, surgical margin status), and survival outcomes were collected. All staging was completed according to the American Joint Committee on Cancer Staging Manual, 7th edn (Edge et al, 2010).
The primary aim was the incidence of hotspot mutations. The secondary aim was the association between hotspot mutations and survival outcomes—overall survival (OS), disease-specific survival (DSS), disease-free survival (DFS) and distant metastasis-free survival (DMFS)—and with clinicopathological features. The index date for survival outcomes for OS and DSS was set as the date of treatment initiation. DFS was defined as the time from the date of completion of primary treatment to the earliest evidence of disease recurrence. DMFS was defined as the time from the date of completion of primary treatment to the earliest evidence of distant metastasis.
Mutation analysis
Tumour cells were identified in regions with >20% nuclei. Genomic DNA was extracted from formalin-fixed paraffin-embedded tumours and subjected to PCR sequencing using a next-generation sequencing platform to screen for mutations in the coding sequences of 50 key signalling genes in melanoma (see Supplementary Table 1 for the full list of covered genes, exons and codons). The results of the next-generation sequencing were confirmed by a second independent PCR and sequencing reaction. The genomic reference sequence used was genome GRCh37/hg19. The sensitivity of the assay is related in part to depth of coverage, percentage of tumour cells with the mutation, and allelic frequency of the mutation. We determined the effective lower limit of detection of this assay (that is, analytical sensitivity) for single-nucleotide variations to be in the range of 5% (one mutant allele per 19 wild-type alleles) to 10% (one mutant allele per nine wild-type alleles) by considering the depth of coverage at a given base and the ability to confirm low-level mutations using independent conventional platforms. The variants detected by our assay were determined on the basis of both analytic findings, such as allelic frequency, and the currently available information in the curated reference databases COSMIC version 64 (Catalogue Of Somatic Mutations In Cancer, Wellcome Trust Sanger Institute, Hinxton, UK) and dbSNP version 137 (National Institute of Health, Bethesda, MD, USA).
Statistical analysis
Basic baseline descriptive statistics were generated. Continuous data were compared according to mutation status using the Student t-test, and categorical variables were compared according to mutation status using the χ2-test. The Kaplan–Meier method was employed for all survival analyses. Survival curves were stratified according to the presence of mutations and compared using the log-rank test. Univariate and multivariate Cox proportional hazards models were used to compare survival outcomes according to mutation status and clinicopathological features. All statistical tests were two-tailed. Significance was defined by an alpha set to 0.05. All statistical testing was completed on SAS JMP Pro version 12.1.0 (SAS Institute, Cary, NC, USA).
Results
Clinicopathological features
Sixty-six patients met all inclusion and exclusion criteria. Patient and tumour characteristics are summarised in Table 1. There were 33 women and 33 men with a median age at diagnosis of 64 years (range 34–85 years). The tumour epicentre was located in the nasal cavity in 53 (80%) patients and in the paranasal sinuses in 13 patients (eight in the maxillary sinus, three in the sphenoid sinus, one in an ethmoid sinus and one in a frontal sinus). Thirty-five (53%) patients had T3 disease, 23 (35%) had T4a disease and eight (12%) had T4b disease. Nodal metastases were present in seven patients (11%). Surgery was the mainstay of treatment in 58 (88%) cases, and in 26 (39%) patients surgery was the only treatment modality. Adjuvant radiotherapy was administered in 30 (45%) patients, and four (6%) patients were treated with adjuvant chemoradiotherapy.
Mutation analysis
Of the 66 primary SNMMs analysed, 27 (41%) harboured at least one identified mutation, and 39 (60%) had no identified mutations. The most common mutation was NRAS mutation (n=20, 30%, P<0.001). Mutations in BRAF, KIT and TP53 occurred in five (8%), three (5%) and two (3%) patients, respectively (Table 2). In 24 patients (89% of the patients with at least one mutation), mutations in KIT, NRAS and BRAF were mutually exclusive.
The NRAS mutations involved codons 12 (G12A, G12R and G12V), 13 (G13R, G13C and G13D) and 61 (Q61K, Q61L, and Q61R). Eleven of the NRAS mutations were located in exon 1. The three KIT mutations were missense; of those, one was the hotspot mutation p.M541L in exon 10 with simultaneous BRAF V600K mutation (patient 26, Table 2). One tumour harboured a KIT mutation in exon 13 simultaneously with ERBB2, NOTCH1, PI3KR1 and TP53 mutations. No mutations were observed in exon 17 of KIT. Among the five BRAF mutations, four were in codon 600 (BRAFV600E and BRAFV600K), and one was in codon 594 (D594G). Both TP53 mutations were in exon 5; interestingly, one of the patients with TP53 mutation carried a germline polymorphism, but not mutation, of KIT (c.1621A>C p.M541L).
Association of mutations with clinicopathological features
The clinicopathological features of tumours with NRAS, KIT, TP53 or BRAF mutations and tumours lacking these mutations are compared in Table 3. Tumours with these mutations were more likely to be located in the paranasal sinuses (30%), whereas the lesions without identified mutations were more often found in the nasal cavity (13%); however, the difference in location was not statistically significant (P=0.09). Mutated tumours had a significantly higher rate of mitosis compared with lesions without identified mutations (63% and 31% respectively, had mitosis rates of ⩾1 mm−2; P=0.01). The distribution of SNMM cell morphological types (Thompson et al, 2003), including epithelioid, spindle, pleomorphic, rhabdoid pagetoid and undifferentiated (small) cells, was similar for patients with and without identified mutations. There were no differences between the mutation groups with respect to age at diagnosis, sex, smoking status, T classification, N classification or bone invasion. The occurrence rates of perineural and lymphovascular invasion were too low for analysis (n=2 for both).
Association of mutations and clinicopathological features with survival outcomes
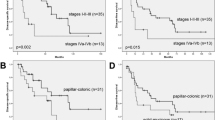
In the whole cohort, the 5-year OS rate was 39%, and the 5-year DSS rate was 54%. The 5-year OS rate was 43% in patients carrying a mutation and 37% in those without an identified mutation (log-rank P=0.55; Figure 1A). The 5-year DSS rate was 54% for both mutation groups (log-rank P=0.91; Figure 1B).
Comparison of survival outcomes in patients with sinonasal mucosal melanoma according to mutation status. (A) Ten-year overall survival, (B) disease-specific survival and (C) disease-free survival by mutation status and (D) 10-year distant metastasis rate calculated using the Kaplan–Meier analysis in patients with (blue line) and without (red line) identified mutations.
Recurrence occurred in 59 (89%) patients over the follow-up period; of these, 27 (40%) had distant metastases. The 5-year DFS was 24% in patients carrying a mutation and 11% for those without an identified mutation (log-rank P=0.64; Figure 1C). A subgroup analysis of patients with NRAS mutations showed no association of NRAS mutations with DFS or DMFS (log-rank P=0.31 and P=0.57, respectively). In patients without identified mutations there was a trend toward a higher 5-year distant metastasis rate compared with patients carrying NRAS, KIT, TP53 or BRAF mutations (78% and 55%, respectively; log-rank P=0.07; Figure 1D).
Univariate analysis comparing patients with and without detected mutations in their tumours showed no association of mutation status with either OS or DSS. To further assess the ability of mutation status to predict outcome in a more homogeneous population and to account for the potential impact of adjuvant treatment, we performed subgroup analyses of each of the following treatment groups: patients undergoing surgery alone (n=30), patients undergoing postoperative radiotherapy (n=30), and patients undergoing adjuvant chemoradiotherapy (n=6). In all treatment groups, mutation status was not an independent predictor of OS or DSS (log-rank analysis, Supplementary Figure 1).
Patients with T3 disease had a significantly better prognosis than those with a T4a or T4b disease, with 5-year OS rates at 58%, 48% and 18%, respectively (log-rank P=0.02, Figure 2A). Similarly, patients with negative margins had a better 5-year OS rate than patients with positive margins (54% and 27%, respectively; log-rank P=0.009; Figure 2B). Of note, patients with tumours in the nasal cavity had a marginally better 5-year OS rate than those with tumours in the paranasal sinuses (48% and 22%, respectively; log-rank P=0.06; Figure 2C). Multivariate Cox regression modelling of these data revealed that only margin status was a significant prognostic factor for OS (hazard ratio 5.43, 95% confidence interval 1.44–21.85, P=0.01) and DSS (hazard ratio 21.9, 95% confidence interval 3.71–180, P=0.0004). To control for margin status, we performed survival analyses separately in patients with positive and negative margins. This analysis revealed no difference in OS and DSS between patients with and without detected mutations in their tumours (Supplementary Figure 2).
Independent risk factors in sinonasal mucosal melanoma. Kaplan–Meier curves of overall survival according to (A) T classification, (B) margin status and (C) tumour site. T classification and surgical margin status reliably distinguished between patients in each subgroup by risk for treatment failure (P<0.05).
Discussion
In this study, we comprehensively screened primary SNMMs for over a hundred different mutations in more than 50 key genes in melanoma and found that NRAS mutations were prevalent (30%). In this retrospective, single-institution analysis, we did not find an association between mutation status and survival outcomes but did find that tumours with identified mutations had a higher mitosis rate.
Genomic aberrations are present in most melanomas (Hodis et al, 2012; Akbani et al, 2015). An increasing understanding of melanocyte biology and melanoma pathogenesis has led to the development of targeted therapies and the potential for major improvements in the care of patients with advanced melanoma. For now, large-scale genomic data in melanoma, derived mainly from cutaneous melanoma, focus on specific genes such as NRAS and its downstream mediator BRAF (Omholt et al, 2003). Targeting these pathways in patients with previously untreated melanoma with these mutations showed promising outcomes (Chapman et al, 2011). However, despite these breakthroughs, the prognosis of patients presenting with SNMM remains poor. Thus, we sought to characterise potential molecular markers in patients with these uncommon melanomas.
Published studies have reported slightly lower overall mutation rates in head and neck mucosal melanoma (10–25%) (Chraybi et al, 2013; Zebary et al, 2013b; Lyu et al, 2016; Ozturk Sari et al, 2017) than that in the current study (40%); however, there was a considerably similar distribution of specific mutation rates in these studies: NRAS, 14%–60%; BRAF, 0%–6%; and KIT, 3%–12% (Cohen et al, 2004; Beadling et al, 2008; Carvajal et al, 2011; Turri-Zanoni et al, 2013; Zebary et al, 2013b). The Cancer Genome Atlas and other large-scale genomic analysis efforts in melanoma have identified hotspot NRAS mutations, thought to be important drivers of oncogenesis, in 25–30% of cutaneous melanomas (Akbani et al, 2015; Krauthammer et al, 2015). Our data show a similar rate (30%) of NRAS mutations. However, in cutaneous melanoma, mutations at codon 61 (Q61R and Q61K) represent the two most common NRAS mutations. In the current study, only 40% of the patients carrying NRAS mutations had Q61R or Q61K mutations, whereas 55% of these patients had mutations in codons 12 (G12V, G12A, G12R and G12D) and 13 (G13R, G13C and G13D). These NRAS mutations at codons 12 and 13 are also prevalent in haematological malignancies (Ward et al, 2012). The different patterns of NRAS mutations in mucosal melanoma compared with cutaneous melanoma support an aetiology other than sun exposure. Another important risk factor in head and neck cancer is smoking. We found a trend towards a lower mutation rate in smokers; however, this difference did not reach significance.
The most common somatic event in cutaneous melanoma is mutation of the serine-threonine kinase BRAF, which is a component of the RAS-RAF-MEK-MAPK signalling pathway. Overall, point mutations in BRAF occur in 40–50% of melanomas (Curtin et al, 2005b). Over 90% of the mutations in BRAF result in substitution of the valine at position 600, resulting in activation of the downstream effectors of the RAS-RAF-MEK-MAPK pathway. Recently, a combination of anti-BRAF and anti-MEK agents have led to an increased response rate and longer duration of response in cutaneous melanoma patients (Larkin et al, 2014; Long et al, 2014). However, the use of these targeted agents is limited to the ∼40% of patients who have melanoma with a BRAFV600 mutation. We identified BRAFV600E and BRAFV600K mutations in only four out of 66 SNMMs. This frequency is similar to the incidence of BRAF mutations in mucosal melanomas from other sites such as the vulva, vagina and anorectum (Omholt et al, 2003; Curtin et al, 2005b).
Most melanoma samples that harboured a hotspot mutation in NRAS, KIT or BRAF did so in a mutually exclusive fashion. The two exceptions harboured BRAFV600 mutations together with an oncogenic NRAS or KIT mutation. Two cases harboured a TP53 missense mutation in exon 5. Interestingly, one patient presented with NOTCH1, PI3KR1, TP53 and KIT mutations, all of which have been previously shown to have a role in melanoma oncogenesis (Liu et al, 2006).
We found a mutation in KIT in only three out of 66 SNMMs. Of those cases, two had additional identified mutation (patients 2 and 26, Table 2). KIT mutations are associated with chronic sun damage in cutaneous melanoma, which is not an aetiological risk factor in SNMMs (Curtin et al, 2005b). However, previous observations suggested that KIT is the most commonly mutated gene in mucosal melanoma, with up to 45% of vulvovaginal and anorectal melanomas carrying a mutation in KIT (Omholt et al, 2011; Schoenewolf et al, 2012). These findings suggest that KIT mutations differ between mucosal melanomas at different sites and are very rare in SNMMs.
We found a significantly higher mitosis rate in patients carrying an identified mutation. There also were trends toward a higher rate of mutations in tumours originating in the paranasal sinuses rather than the nasal cavity and worse prognosis in patients with disease originating from the sinuses compared with those with tumours originating from the nasal cavity. Our finding that mutation status, for all known mutations or for NRAS alone, did not affect survival in the setting of SNMM is in keeping with studies conducted before the availability of MEK inhibitors and immune checkpoint inhibitor antibodies (Ellerhorst et al, 2011). The high proportion of NRAS-mutated tumours suggests that further studies investigating the use of MEK inhibitors, which have shown promising phase II results in cutaneous melanoma with NRAS mutations, may be worthwhile in SNMMs (Ascierto et al, 2013). A phase III study comparing the MEK inhibitor binimetinib with dacarbazine in patients with NRAS-mutant cutaneous melanoma showed longer progression-free survival in patients treated with binimetinib (Dummer, 2016). However, the adverse events profile of these agents, including cardiomyopathy, hypertension, coagulopathies and rash, makes them good candidates for a combined treatment regimen rather than single-agent therapy.
In the present study, we included only patients seen at a single tertiary cancer centre. Although mutation testing was done prospectively in patients with SNMM, data were collected and analysed retrospectively, which might limit our ability to control for patient comorbidities and different treatments administered. Also, matched non-tumour tissue was not tested, so the possibility of a detected mutation being a germline mutation cannot be completely ruled out. In our cohort, 24 patients had one mutation, two patients had two mutations, and one patient had eight mutations. Because of the low number of events, we could not analyse the correlation between the number of mutations and the outcome. However, our study represents the largest single-institution cohort to date of SNMM patients undergoing characterisation of mutation status. The role of mutation status, particularly NRAS mutations in G12 and G13, as a biomarker for response to MEK inhibition in SNMM needs to be addressed in future studies.
In conclusion, NRAS, BRAF and KIT mutations do not affect survival outcomes in SNMM. As MEK inhibitors have shown promise in the treatment of cutaneous melanoma, their prognostic impact in SNMM should be further investigated, especially in the relatively frequent cases with NRAS mutations.
Change history
06 June 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Akbani R, Akdemir KC, Aksoy BA, Albert M, Ally A, Amin SB, Arachchi H, Arora A, Auman JT, Ayala B, Baboud J, Balasundaram M, Balu S, Barnabas N, Bartlett J, Bartlett P, Bastian BC, Baylin SB, Behera M, Belyaev D, Benz C, Bernard B, Beroukhim R, Bir N, Black AD, Bodenheimer T, Boice L, Boland GM, Bono R, Bootwalla MS, Bosenberg M, Bowen J, Bowlby R, Bristow CA, Brockway-Lunardi L, Brooks D, Brzezinski J, Bshara W, Buda E, Burns WR, Butterfield YSN, Button M, Calderone T, Cappellini GA, Carter C, Carter SL, Cherney L, Cherniack AD, Chevalier A, Chin L, Cho J, Cho RJ, Choi YL, Chu A, Chudamani S, Cibulskis K, Ciriello G, Clarke A, Coons S, Cope L, Crain D, Curley E, Danilova L, D’Atri S, Davidsen T, Davies MA, Delman KA, Demchok JA, Deng QA, Deribe YL, Dhalla N, Dhir R, DiCara D, Dinikin M, Dubina M, Ebrom, Egea S, Eley G, Engel J, Eschbacher JM, Fedosenko KV, Felau I, Fennell T, Ferguson ML, Fisher S, Flaherty KT, Frazer S, Frick J, Fulidou V, Gabriel SB, Gao J, Gardner J, Garraway LA, Gastier-Foster JM, Gaudioso C, Gehlenborg N, Genovese G, Gerken M, Gershenwald JE, Getz G, Gomez-Fernandez C, Gribbin T, Grimsby J, Gross B, Guin R, Gutschner T, Hadjipanayis A, Halaban R, Hanf B, Haussler D, Haydu LE, Hayes DN, Hayward NK, Heiman DI, Herbert L, Herman JG, Hersey P, Hoadley KA, Hodis E, Holt RA, Hoon DSB, Hoppough S, Hoyle AP, Huang FW, Huang M, Huang S, Hutter CM, Ibbs M, Iype L, Jacobsen A, Jakrot V, Janning A, Jeck WR, Jefferys SR, Jensen MA, Jones CD, Jones SJM, Ju Z, Kakavand H, Kang H, Kefford RF, Khuri FR, Kim J, Kirkwood JM, Klode J, Korkut A, Korski K, Krauthammer M, Kucherlapati R, Kwong LN, Kycler W, Ladanyi M, Lai PH, Laird PW, Lander E, Lawrence MS, Lazar AJ, Qazniak R, Lee D, Lee JE, Lee J, Lee K, Lee S, Lee W, Leporowska E, Leraas KM, Li HI, Lichtenberg TM, Lichtenstein L, Lin P, Ling SY, Liu J, Liu O, Liu W, Long GV, Lu Y, Ma S, Ma Y, Mackiewicz A, Mahadeshwar HS, Malke J, Mallery D, Manikhas GM, Mann GJ, Marra MA, Matejka B, Mayo M, Mehrabi S, Meng S, Meyerson M, Mieczkowski PA, Miller JP, Miller ML, Mills GB, Moiseenko F, Moore RA, Morris S, Morrison C, Morton DL, Moschos S, Mose LE, Muller FL, Mungall AJ, Murawa D, Murawa P, Murray BA, Nezi L, Ng S, Nicholson D, Noble MS, Osunkoya A, Owonikoko TK, Ozenberger BA, Pagani E, Paklina OV, Pantazi A, Parfenov M, Parfitt J, Park PJ, Park WY, Parker JS, Passarelli F, Penny R, Perou CM, Pihl TD, Potapova O, Prieto VG, Protopopov A, Quinn MJ, Radenbaugh A, Rai K, Ramalingam SS, Raman AT, Ramirez NC, Ramirez R, Rao U, Rathmell WK, Ren XJ, Reynolds SM, Roach J, Robertson AG, Ross MI, Roszik J, Russo G, Saksena G, Saller C, Samuels Y, Sander C, Sander C, Sandusky G, Santoso N, Saul M, Saw RPM, Schadendorf D, Schein JE, Schultz N, Schumacher SE, Schwallier C, Scolyer RA, Seidman J, Sekhar PC, Sekhon HS, Senbabaoglu Y, Seth S, Shannon KF, Sharpe S, Sharpless NE, Shaw KRM, Shelton C, Shelton T, Shen R, Sheth M, Shi Y, Shiau CJ, Shmulevich I, Sica GL, Simons JV, Sinha R, Sipahimalani P, Sofia HJ, Soloway MG, Song XZ, Sougnez C, Spillane AJ, Spychaa A, Stretch JR, Stuart J, Suchorska WM, Sucker A, Sumer SO, Sun YC, Synott M, Tabak B, Tabler TR, Tam A, Tan DH, Tang JB, Tarnuzzer R, Tarvin K, Tatka H, Taylor BS, Teresiak M, Thiessen N, Thompson JF, Thorne L, Thorsson V, Trent JM, Triche TJ, Tsai KY, Tsou P, Van den Berg DJ, Van Allen EM, Veluvolu U, Verhaak RG, Voet D, Voronina O, Walter V, Walton JS, Wan YH, Wang YL, Wang ZN, Waring S, Watson IR, Weinhold N, Weinstein JN, Weisenberger DJ, White P, Wilkerson MD, Wilmott JS, Wise L, Wiznerowicz M, Woodman SE, Wu CJ, Wu CC, Wu JY, Wu Y, Xi RB, Xu AW, Yang D, Yang LM, Yang LX, Zack TI, Zen-Klusen JC, Zhang HL, Zhang JH, Zhang W, Zhao XB, Zhu JC, Zhu K, Zimmer L, Zmuda E, Zou LH Network CGA (2015) Genomic classification of cutaneous melanoma. Cell 161 (7): 1681–1696.
Ascierto PA, Schadendorf D, Berking C, Agarwala SS, van Herpen CML, Queirolo P, Blank CU, Hauschild A, Beck JT, St-Pierre A, Niazi F, Wandel S, Peters M, Zubel A, Dummer R (2013) MEK162 for patients with advanced melanoma harbouring NRAS or Val600 BRAF mutations: a non-randomised, open-label phase 2 study. Lancet Oncol 14 (3): 249–256.
Beadling C, Jacobson-Dunlop E, Hodi FS, Le C, Warrick A, Patterson J, Town A, Harlow A, Cruz F 3rd, Azar S, Rubin BP, Muller S, West R, Heinrich MC, Corless CL (2008) KIT gene mutations and copy number in melanoma subtypes. Clin Cancer Res 14 (21): 6821–6828.
Carvajal RD, Antonescu CR, Wolchok JD, Chapman PB, Roman RA, Teitcher J, Panageas KS, Busam KJ, Chmielowski B, Lutzky J, Pavlick AC, Fusco A, Cane L, Takebe N, Vemula S, Bouvier N, Bastian BC, Schwartz GK (2011) KIT as a therapeutic target in metastatic melanoma. JAMA 305 (22): 2327–2334.
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, Jouary T, Schadendorf D, Ribas A, O’Day SJ, Sosman JA, Kirkwood JM, Eggermont AMM, Dreno B, Nolop K, Li J, Nelson B, Hou J, Lee RJ, Flaherty KT, McArthur GA, Grp B-S (2011) Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 364 (26): 2507–2516.
Chraybi M, Abd Alsamad I, Copie-Bergman C, Baia M, Andre J, Dumaz N, Ortonne N (2013) Oncogene abnormalities in a series of primary melanomas of the sinonasal tract: NRAS mutations and cyclin D1 amplification are more frequent than KIT or BRAF mutations. Hum Pathol 44 (9): 1902–1911.
Cohen Y, Rosenbaum E, Begum S, Goldenberg D, Esche C, Lavie O, Sidransky D, Westra WH (2004) Exon 15 BRAF mutations are uncommon in melanomas arising in nonsun-exposed sites. Clin Cancer Res 10 (10): 3444–3447.
Curtin JA, Fridlyand J, Kageshita T, Patel HN, Busam KJ, Kutzner H, Cho KH, Aiba S, Brocker EB, LeBoit PE, Pinkel D, Bastian BC (2005a) Distinct sets of genetic alterations in melanoma. N Engl J Med 353 (20): 2135–2147.
Curtin JA, Fridlyand J, Kageshita T, Patel HN, Busam KJ, Kutzner H, Cho KH, Aiba S, Brocker EB, LeBoit PE, Pinkel D, Bastian BC (2005b) Distinct sets of genetic alterations in melanoma. N Engl J Med 353 (20): 2135–2147.
Dummer R (2016) Results of NEMO: Aphase III trial of binimetinib (BINI) vs dacarbazine (DTIC) in NRAS-mutant cutanous melanoma. J Clin Oncol 34 (suppl): abstr 9500.
Edge SBDR, Byrd DR, Compton CC, Fritz AG, Greene FL, Trotti A (2010) AJCC Cancer Staging Manual, 7th edn. Springer: New York, USA.
Ellerhorst JA, Greene VR, Ekmekcioglu S, Warneke CL, Johnson MM, Cooke CP, Wang LE, Prieto VG, Gershenwald JE, Wei QY, Grimm EA (2011) Clinical correlates of NRAS and BRAF mutations in primary human melanoma. Clin Cancer Res 17 (2): 229–235.
Gal TJ, Silver N, Huang B (2011) Demographics and treatment trends in sinonasal mucosal melanoma. Laryngoscope 121 (9): 2026–2033.
Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP, Nickerson E, Auclair D, Li LR, Place C, DiCara D, Ramos AH, Lawrence MS, Cibulskis K, Sivachenko A, Voet D, Saksena G, Stransky N, Onofrio RC, Winckler W, Ardlie K, Wagle N, Wargo J, Chong K, Morton DL, Stemke-Hale K, Chen G, Noble M, Meyerson M, Ladbury JE, Davies MA, Gershenwald JE, Wagner SN, Hoon DSB, Schadendorf D, Lander ES, Gabriel SB, Getz G, Garraway LA, Chin L (2012) A landscape of driver mutations in melanoma. Cell 150 (2): 251–263.
Krauthammer M, Kong Y, Bacchiocchi A, Evans P, Pornputtapong N, Wu C, McCusker JP, Ma SG, Cheng E, Straub R, Serin M, Bosenberg M, Ariyan S, Narayan D, Sznol M, Kluger HM, Mane S, Schlessinger J, Lifton RP, Halaban R (2015) Exome sequencing identifies recurrent mutations in NF1 and RASopathy genes in sun-exposed melanomas. Nat Genet 47 (9): 996–1002.
Larkin J, Ascierto PA, Dreno B, Atkinson V, Liszkay G, Maio M, Mandala M, Demidov L, Stroyakovskiy D, Thomas L, de la Cruz-Merino L, Dutriaux C, Garbe C, Sovak MA, Chang I, Choong N, Hack SP, McArthur GA, Ribas A (2014) Combined vemurafenib and cobimetinib in BRAF-mutated melanoma. N Engl J Med 371 (20): 1867–1876.
Ledderose GJ, Leunig A (2015) Surgical management of recurrent sinonasal mucosal melanoma: endoscopic or transfacial resection. Eur Arch Otorhinolaryngol 272 (2): 351–356.
Lee JH, Choi JW, Kim YS (2011) Frequencies of BRAF and NRAS mutations are different in histological types and sites of origin of cutaneous melanoma: a meta-analysis. Br J Dermatol 164 (4): 776–784.
Liu ZJ, Xiao M, Balint K, Smalley KSM, Brafford P, Qiu RH, Pinnix CC, Li XL, Herlyn M (2006) Notch1 signaling promotes primary melanoma progression by activating mitogen-activated protein kinase/phosphatidylinositol 3-kinase-Akt pathways and up-regulating N-cadherin expression. Cancer Res 66 (8): 4182–4190.
Long GV, Stroyakovskiy D, Gogas H, Levchenko E, de Braud F, Larkin J, Garbe C, Jouary T, Hauschild A, Grob JJ, Sileni VC, Lebbe C, Mandala M, Millward M, Arance A, Bondarenko I, Haanen JBAG, Hansson J, Utikal J, Ferraresi V, Kovalenko N, Mohr P, Probachai V, Schadendorf D, Nathan P, Robert C, Ribas A, DeMarini DJ, Irani JG, Casey M, Ouellet D, Martin AM, Le N, Patel K, Flaherty K (2014) Combined BRAF and MEK inhibition versus BRAF inhibition alone in melanoma. N Engl J Med 371 (20): 1877–1888.
Lourenco SV, Fernandes JD, Hsieh R, Coutinho-Camillo CM, Bologna S, Sangueza M, Nico MM (2014) Head and neck mucosal melanoma: a review. Am J Dermatopathol 36 (7): 578–587.
Lyu J, Wu Y, Li C, Wang R, Song H, Ren G, Guo W (2016) Mutation scanning of BRAF, NRAS, KIT, and GNAQ/GNA11 in oral mucosal melanoma: a study of 57 cases. J Oral Pathol Med 45 (4): 295–301.
Moreno MA, Roberts DB, Kupferman ME, DeMonte F, El-Naggar AK, Williams M, Rosenthal DS, Hanna EY (2010) Mucosal melanoma of the nose and paranasal sinuses, a contemporary experience from the M. D. Anderson Cancer Center. Cancer 116 (9): 2215–2223.
Omholt K, Grafstrom E, Kanter-Lewensohn L, Hansson J, Ragnarsson-Olding BK (2011) KIT pathway alterations in mucosal melanomas of the vulva and other sites. Clin Cancer Res 17 (12): 3933–3942.
Omholt K, Platz A, Kanter L, Ringborg U, Hansson J (2003) NRAS and BRAF mutations arise early during melanoma pathogenesis and are preserved throughout tumor progression. Clin Cancer Res 9 (17): 6483–6488.
Ozturk Sari S, Yilmaz I, Taskin OC, Narli G, Sen F, Comoglu S, Firat P, BI B, Yilmazbayhan D, Ozluk Y, Buyukbaban IN (2017) BRAF, NRAS, KIT, TERT, GNAQ/GNA11 mutation profile analysis of head and neck mucosal melanomas: a study of 42 cases. Pathology 49 (1): 55–61.
Samstein RM, Carvajal RD, Postow MA, Callahan MK, Shoushtari AN, Patel SG, Lee NY, Barker CA (2016) Localized sinonasal mucosal melanoma: outcomes and associations with stage, radiotherapy, and positron emission tomography response. Head Neck 38 (9): 1310–1317.
Schoenewolf NL, Bull C, Belloni B, Holzmann D, Tonolla S, Lang R, Mihic-Probst D, Andres C, Dummer R (2012) Sinonasal, genital and acrolentiginous melanomas show distinct characteristics of KIT expression and mutations. Eur J Cancer 48 (12): 1842–1852.
Spagnolo F, Picasso V, Lambertini M, Ottaviano V, Dozin B, Queirolo P (2016) Survival of patients with metastatic melanoma and brain metastases in the era of MAP-kinase inhibitors and immunologic checkpoint blockade antibodies: a systematic review. Cancer Treat Rev 45: 38–45.
Spencer KR, Mehnert JM (2016) Mucosal melanoma: epidemiology, biology and treatment. Cancer Treat Res 167: 295–320.
Sun CZ, Li QL, Hu ZD, Jiang YE, Song M, Yang AK (2014) Treatment and prognosis in sinonasal mucosal melanoma: a retrospective analysis of 65 patients from a single cancer center. Head Neck 36 (5): 675–681.
Thompson LD, Wieneke JA, Miettinen M (2003) Sinonasal tract and nasopharyngeal melanomas: a clinicopathologic study of 115 cases with a proposed staging system. Am J Surg Pathol 27 (5): 594–611.
Turri-Zanoni M, Medicina D, Lombardi D, Ungari M, Balzarini P, Rossini C, Pellegrini W, Battaglia P, Capella C, Castelnuovo P, Palmedo G, Facchetti F, Kutzner H, Nicolai P, Vermi W (2013) Sinonasal mucosal melanoma: molecular profile and therapeutic implications from a series of 32 cases. Head Neck 35 (8): 1066–1077.
Ward AF, Braun BS, Shannon KM (2012) Targeting oncogenic Ras signaling in hematologic malignancies. Blood 120 (17): 3397–3406.
Zebary A, Jangard M, Omholt K, Ragnarsson-Olding B, Hansson J (2013a) KIT, NRAS and BRAF mutations in sinonasal mucosal melanoma: a study of 56 cases. Br J Cancer 109 (3): 559–564.
Zebary A, Omholt K, Vassilaki I, Hoiom V, Linden D, Viberg L, Kanter-Lewensohn L, Johansson CH, Hansson J (2013b) KIT, NRAS, BRAF and PTEN mutations in a sample of Swedish patients with acral lentiginous melanoma. J Dermatol Sci 72 (3): 284–289.
Acknowledgements
The University of Texas MD Anderson Cancer Center is supported in part by the National Institutes of Health through Cancer Center Support Grant P30CA016672.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Amit, M., Tam, S., Abdelmeguid, A. et al. Mutation status among patients with sinonasal mucosal melanoma and its impact on survival. Br J Cancer 116, 1564–1571 (2017). https://doi.org/10.1038/bjc.2017.125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.125
Keywords
This article is cited by
-
Development and validation of a nomogram for predicting overall survival in patients with sinonasal mucosal melanoma
BMC Cancer (2024)
-
Head and Neck Mucosal Melanoma: Where Are We Now?
Current Oncology Reports (2024)
-
Rh-endostatin combined with chemotherapy in patients with advanced or recurrent mucosal melanoma: retrospective analysis of real-world data
Investigational New Drugs (2022)
-
Alterations in key signaling pathways in sinonasal tract melanoma. A molecular genetics and immunohistochemical study of 90 cases and comprehensive review of the literature
Modern Pathology (2022)
-
Analyses of molecular and histopathologic features and expression of PRAME by immunohistochemistry in mucosal melanomas
Modern Pathology (2019)