Abstract
Recently, mutations of the additional sex comb-like 1 (ASXL1) gene were identified in patients with myelodysplastic syndrome (MDS), but the interaction of this mutation with other genetic alterations and its dynamic changes during disease progression remain to be determined. In this study, ASXL1 mutations were identified in 106 (22.7%) of the 466 patients with primary MDS based on the French-American-British (FAB) classification and 62 (17.1%) of the 362 patients based on the World Health Organization (WHO) classification. ASXL1 mutation was closely associated with trisomy 8 and mutations of RUNX1, EZH2, IDH, NRAS, JAK2, SETBP1 and SRSF2, but was negatively associated with SF3B1 mutation. Most ASXL1-mutated patients (85%) had concurrent other gene mutations at diagnosis. ASXL1 mutation was an independent poor prognostic factor for survival. Sequential studies showed that the original ASXL1 mutation remained unchanged at disease progression in all 32 ASXL1-mutated patients but were frequently accompanied with acquisition of mutations of other genes, including RUNX1, NRAS, KRAS, SF3B1, SETBP1 and chromosomal evolution. On the other side, among the 80 ASXL1-wild patients, only one acquired ASXL1 mutation at leukemia transformation. In conclusion, ASXL1 mutations in association with other genetic alterations may have a role in the development of MDS but contribute little to disease progression.
Similar content being viewed by others
Introduction
The myelodysplastic syndromes (MDSs) are a heterogenous group of diseases, characterized by cytopenia, but usually hypercellular bone marrow with dysplastic hematopoiesis, and a propensity to acute leukemia transformation.1, 2 The pathogenesis that causes these pre-leukemic disorders is not quite clear yet, but immune deregulation, abnormal microenvironment and accumulation of genetic alterations may all have some roles.3, 4, 5, 6, 7 Mutations in ASXL1 (additional sex combs 1) have been identified in MDS8 and other myeloid malignancies, like acute myeloid leukemia (AML),9 chronic myelomonocytic leukemia (CMML)10 and myeloproliferative neoplasms.11 The function of ASXL1 protein is not fully delineated,12 but it is suggested that it may be involved in DNA and/or histone modification.13 ASXL1 mutations are all disclosed in exon 12 of the gene and are believed to lead to the truncation of the plant homeodomain10 at the C-terminus of the protein, which is involved in chromatin modification.14, 15 Mutations in ASXL1 result in global decrease of histone 3 lysine 27 methylation, a histone marker associated with repression of transcription.16 Recently, the animal models showed that C-terminal-truncating ASXL1 mutations or deletion/loss of ASXL1 lead to MDS-like disease in mice.17, 18, 19 Further, the deficiency of the BAP1, a nuclear-localized deubiquitinating enzyme, resulted in a CMML-like phenotype, and the interaction with ASXL1 is critical for the enzymatic activity of BAP1.16, 20, 21 ASXL1 mutation is found in a substantial proportion (11–18.5%) of WHO-defined MDS patients8, 10, 22, 23, 24 and is correlated with unfavorable outcome.23, 24 However, the association of ASXL1 mutation with other genetic alterations in the pathogenesis of MDS and their dynamic changes during disease progression remain unclear. In this large cohort of MDS patients, we found that ASXL1 mutation was statistically closely associated with trisomy 8 and mutations of RUNX1, EZH2, IDH, NRAS, JAK2, SETBP1 and SRSF2, suggesting cooperation of these gene alterations with ASXL1 mutation may contribute to the development of MDS. Moreover, sequential analyses showed all ASXL1-mutated patients retained the original ASXL1 mutation during disease progression, but frequently acquired other novel genetic alterations, including RUNX1, NRAS, KRAS, SF3B1 and SETBP1 mutations and chromosomal evolution, at the same time.
Materials and methods
Patients
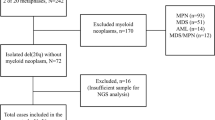
Four hundred and sixty-six adult patients who were diagnosed as having de novo MDS according to the FAB classification at the National Taiwan University Hospital (NTUH) and had cryopreserved bone marrow cells for study were recruited for gene mutation analyses. Among them, the disease of 362 patients fulfilled the criteria of MDS according to the 2008 WHO classification. All patients signed informed consents for sample collection in accordance with the Declaration of Helsinki. This study was approved by the Institutional Review Board of the NTUH.
Mutation analysis
The ASXL1 exon 12 until the stop codon was amplified by three pairs of primers and sequenced by another six internal primers, as described by Gelsi-Boyer et al.,10 with mild modification.9 The PCR reaction included 95 °C for 2 min, followed by 35 cycles of 95 °C for 30 s, 61 °C for 30 s and 72 °C for 1 min. The mutations were confirmed at least twice. When the mutations were not obvious because of location near the sequencing primers, sequencing from the other direction was done to solve this issue. Mutation analyses of other 15 relevant molecular genes, including class I mutations, such as FLT3/ITD,25 NRAS,26 KRAS26 and JAK226 mutations, and class II mutations, such as MLL/PTD,27 RUNX128 and WT1 mutations,29 as well as mutations of genes involving in epigenetic modifications, such as IDH1,30 IDH2,31 including R140 and R172 mutations, DNMT3A32 and EZH233 mutations, splicing machinery mutations, such as U2AF1,34 SRSF235 and SF3B134 mutations, and SETBP1 mutation,36 were performed as previously described. Sequential studies of all these genes were also performed in 305 samples from 112 patients during clinical follow-ups.
Cytogenetics
Bone marrow cells were harvested directly or after 1–3 days of unstimulated culture, and the metaphase chromosomes were banded by the G-banding method as described earlier.37
TA-cloning analysis
For the patients with discrepancy of the mutation status of the ASXL1 in paired samples, Taq polymerase-amplified (TA) cloning was performed in the samples without detectable mutant by direct sequencing. The DNA spanning the mutation spots of ASXL1 detected at either diagnosis or during subsequent follow-ups was amplified and the PCR products were then cloned into the TA-cloning vector pGEM-T Easy (Promega, Madison, WI, USA). More than 10 clones were selected for sequencing as previously described.38
Statistics
The χ2-test was performed to calculate the significance of association between ASXL1 mutation and other parameters, including sex, the FAB subtypes, the 2008 WHO classification, karyotypes, international prognostic scoring system (IPSS) score,1 revised IPSS (IPSS-R) score39 and mutations of other genes. Fisher’s exact test was used if any expected value of the contingency table was less than 5. The Mann–Whitney test method was used to compare continuous variables and medians of distributions. Overall survival (OS) was measured from the date of first diagnosis to the date of last follow-up or death from any cause. Multivariate Cox proportional hazard regression analysis was used to investigate independent prognostic factors for OS. The Kaplan–Meier estimation was used to plot survival curves, and log-rank tests were used to calculate the difference of OS between groups. A P-value<0.05 was considered statistically significant. All statistical analyses were performed with SPSS 17 software (SPSS Inc., Chicago, IL, USA) and Statsdirect (Cheshire, UK).
Results
Mutation status of ASXL1 in patients with MDS
Among the 466 MDS patients according to the FAB classification, 106 (22.7%) patients had ASXL1 mutations, including 96 patients with frameshift mutations and 10 with nonsense mutations. The most common mutation was c.1934dupG that occurred in 66 patients. All 106 patients showed single heterozygous mutation. (Supplementary Table 1) All these mutations resulted in truncation of the plant homeodomain of ASXL1. Patients with CMML had a high incidence (53.8%) of ASXL1 mutations (Table 1). Patients with refractory anemia (RA) and RA with ring sideroblasts had a lower incidence (10.5% and 11.8%, respectively) of ASXL1 mutation than patients with RA with excess blasts (RAEB, 25.5%) or RAEB in transformation (30.8%; P<0.001).
Among the 362 patients with MDS according to the 2008 WHO classification, 17.1% patients had ASXL1 mutation. MDS patients with RAEB1/RAEB2 had a significantly higher incidence of ASXL1 mutation than those with other subtypes (25% vs 10.7%, P<0.001, Table 1).
Clinical features and biological characteristics of ASXL1-mutated patients
Most of the patients received conservative and supportive care and only 91 (19.5%) patients received AML-directed chemotherapy, including 56 patients (12%) who underwent allogeneic hematopoietic stem cell transplantation. We could not find the treatment difference between the patients with ASXL1 mutations and those without ASXL1 mutations (data not shown). The comparison of clinical and hematologic characteristics of patients with and without mutation of ASXL1 is shown in Table 1. The patients with ASXL1 mutation were predominantly male, older (median, 71 years vs 64 years, P=0.001) and had higher white blood cell counts (P<0.001) and higher IPSS-R score (P=0.002) at diagnosis. There was no difference in hemoglobin levels and platelet counts between the patients with and without ASXL1 mutation.
Among the 435 patients with cytogenetic data for analysis, clonal chromosomal abnormalities were detected in 44.4% of the MDS patients based on the FAB classification and in 44.2% of those based on the 2008 WHO classification. The mutation rate was especially high in the patients with trisomy 8 (45.5%, P=0.02; Supplementary Table 2).
Association of ASXL1 mutation with other genetic alterations
Ninety (85%) of the ASXL1-mutated patients had concurrently other gene mutations (Supplementary Table 1 and Table 2). The patients with ASXL1 mutation had significantly higher incidences of concurrent RUNX1 mutation (32.4% vs 6.8%, P<0.001), EZH2 mutation (22.6% vs 1.1%, P<0.001), IDH mutation (11.4% vs 2.5%, P<0.001), NRAS mutation (10.4% vs 3.1%, P=0.007), JAK2 mutation (3.8% vs 0.3%, P=0.011), SETBP1 mutation (10.5% vs 0.6%, P<0.001) and SRSF2 mutation (34.3% vs 6.7%, P<0.001), but had a lower incidence of concurrent SF3B1 mutation (2.9% vs 12.9%, P=0.003) than those with wild-type ASXL1. There was no correlation of ASXL1 mutation with other gene mutations studied (Table 2).
Analysis of ASXL1 mutation in sequential samples
To investigate the role of ASXL1 mutation in disease progression, sequential analyses of the gene mutation were performed in 305 samples from 112 patients, including 32 patients with ASXL1 mutation at diagnosis and 80 patients without the mutation. Among the 32 ASXL1-mutated patients, 27 had disease progression including 19 with AML transformation. Two patients (patients 60 and 92) lost the original ASXL1 mutation at remission status following transplantation. Among the remaining 30 patients, the same mutations were retained in 29 patients during follow-ups (Table 3) but could not be detected by direct sequencing in one patient at the time of disease progression (patient 1). As direct sequencing might not be sensitive enough to detect low level of ASXL1 mutant, we therefore did TA cloning of the sample obtained at RAEB from this patient. The original ASXL1 mutation was detected in 2 of the 10 clones analyzed.
On the other hand, among the 80 ASXL1-wild patients who were sequentially studied, 36 patients had disease progression, including 20 patients with AML transformation. Two of them (patients 107 and 108) acquired ASXL1 mutations when the disease progressed to AML and RAEB, respectively. Actually, we could not find any ASXL1 mutation in the 44 clones analyzed at diagnosis using cloning technique in patient 107. Patient 108 who was diagnosed as having RA had EZH2 mutation initially. He acquired RUNX1, ASXL1 and SETBP1 mutations when the disease progressed to RAEB 90 months later. Interestingly, using a more sensitive cloning technique, we could identify ASXL1 mutation in one of the 11 clones, but no RUNX1 and SETBP1 mutations in the 41 clones and 40 clones analyzed at diagnosis, respectively. Therefore, a total of 31 patients had ASXL1 mutations at both MDS diagnosis and subsequent follow-ups (Table 3 and Supplementary Table 3). Among them, eight patients acquired mutations of other genes, including RUNX1 in four (patients 22, 51, 83 and 108; Table 3), NRAS in three (patients 26, 56 and 80), KRAS in one (patient 35), SF3B1 in one (patient 22) and SETBP1 in one (patient 108) during disease progression, whereas other six patients had chromosomal evolution (patients 5, 24, 38, 71, 98 and 100).
Influence of ASXL1 mutation on clinical outcome
With a median follow-up duration of 58.2 months (range, 0.1–250.7 months), there was a close correlation between ASXL1 mutations and acute leukemia transformation (39.0% vs 17.7%; P<0.001). If the analysis was restricted to the 362 MDS patients based on the 2008 WHO classification, those with ASXL1 mutation also had a significant higher incidence of acute leukemia transformation (32.5% vs 13.0%; P<0.001) than those without the mutation. MDS patients, based either on the FAB or 2008 WHO classification, had a significantly shorter OS if they harbored ASXL1 mutation than those who did not (median, 18.5 vs 42.4 months, P<0.001; and 21 vs 69.9 months, P<0.001, respectively; Figure 1). The difference remained significant when the analysis was performed in the subgroup of patients with lower-risk MDS defined by the FAB classification (including RA and RA with ring sideroblasts), the WHO classification (other than RAEB, subtypes with blasts<5%), IPSS-R (including very low, low and intermediate groups) and those with favorable-risk cytogenetics (median, 36.1 vs 170.2 months, P<0.001, 36.1 vs 170.2 months, P<0.001, 33.8 vs 113.7 months, P<0.001 and 18.5 vs 69.3 months, P<0.001, respectively; Figure 2). However, there was no prognostic impact of ASXL1 mutation on the MDS patients with higher-risk MDS based on the FAB/WHO classification, IPSS-R or poor-risk cytogenetics.
Kaplan–Meier survival curves in the subgroup of patients with lower-risk MDS. (a) Lower-risk group (RA and RA with ring sideroblasts (RARS)) defined by the FAB classification; (b) lower-risk group (other than RAEB, subtypes with bone marrow (BM) blasts<5%) defined by the 2008 WHO classification; (c, d) patients with lower IPSS-R score; (e, f) patients with favorable/intermediate-risk cytogenetics. *Lower IPSS-R groups include very low, low and intermediate subgroups. **Favorable cytogenetics include very good, good and intermediate-risk cytogenetic changes.
In univariate analysis, older age, unfavorable cytogenetics, higher IPSS-R score, IDH, ASXL1, RUNX1, NRAS, EZH2 and SRSF2 mutations were poor prognostic factors for the OS (Supplementary Table 4). The patients with DNMT3A mutations had a trend of poorer OS than those without (P=0.060). In multivariate analysis using covariables including age⩾50 years, IPSS-R and mutations of ASXL1, RUNX1, NRAS, IDH, SRSF2 and DNMT3A (Table 4), ASXL1 mutation was an independent poor prognostic factor for OS. Interestingly, in the 242 MDS patients with normal karyotype, ASXL1 mutation was also an independent poor prognostic factor (hazard ratio=2.307, 95% confidence interval=1.410–3.775, P=0.001) in addition to older age and NRAS mutation (Table 5).
Discussion
In the present study, ASXL1 mutations were detected in 22.7% and 17.1% of MDS patients defined either by the FAB or the 2008 WHO classification, respectively. A majority of ASXL1-mutated patients had concurrently other gene mutations, most commonly RUNX1 and EZH2 mutations. All ASXL1 mutations detected at diagnosis remained unchanged during disease progression but were frequently accompanied with acquisition of other novel genetic alterations. Moreover, ASXL1 mutation was associated with distinct clinical and biological features and a poorer outcome.
We only counted frameshift mutations and nonsense mutations of ASXL1 gene as true mutations in this study as previously reported.9, 10 Missense mutations were excluded because their significance could not be verified owing to lack of normal tissue for comparison, not being reported previously or not at the sites well conserved among different species (data not shown). Thol et al.23 analyzed the prognostic effect of ASXL1 point and frameshift mutations separately. They found that frameshift mutations, but not point mutations, had an independent prognostic effect in MDS patients. The frequency of ASXL1 mutation in this study (22.7% in MDS patients defined by the FAB classification and 17.1% in those by the 2008 WHO classification) was comparable to that in the West (19.3%22 and 11–18.5%).8, 10, 22, 23, 24 The mutation rate in CMML was also similar between this study and others. (45.5% vs 43–49%).8, 40 So were the incidences of ASXL1 mutation among patients with higher-risk MDS (RAEB by the 2008 WHO classification) and those with lower-risk MDS (other subtypes with bone marrow blasts less than 5; 25.5 vs 18.9–31% and 10.5% vs 8.7–14.2%).22, 23 If the comparison was made separately for these two groups, the mutation occurred more frequently in higher- than in lower-risk MDS.
The report concerning interaction of ASXL1 mutation with other genetic alterations in the pathogenesis of MDS and its progression is very limited. In the report of Rocquain et al.,8 three of 65 MDS patients had both ASXL1 and RUNX1 mutations.8, 41 Another study of 24 patients with CMML showed the co-occurrence of ASXL1 and TET2 mutations in seven cases and ASXL1 and EZH2 mutations in two cases.41 However, because of small patient number in these two studies, no statistical analyses were done to evaluate whether there was a significant association of ASXL1 mutation with these mutations. Our study showed 85% (90 of 106) of ASXL1-mutated patients had concurrent other gene alterations. Furthermore, ASXL1 mutation was closely associated with mutations of RUNX1, EZH2, IDH, NRAS, JAK2, SETBP1 and SRSF2 in MDS patients. In other words, the ASXL1 mutation coincided with mutations of genes involved in the signal transduction pathway (JAK2 and NRAS), transcription factor (RUNX1), epigenetic modification (IDH and EZH2) or splicing machinery (SRSF2) in MDS patients. The manner in which the ASXL1 mutation was associated with these genetic aberrations in the MDS pathogenesis needs further investigation.
The reports regarding sequential studies of ASXL1 mutation in MDS are even less. In one report,8 the ASXL1 mutation detected in one patient with RAEB2 at diagnosis was retained at the time of AML transformation. To the best of our knowledge, this is the largest study to evaluate the dynamic change of ASXL1 mutation during disease progression in MDS. We found that with the exception of the two patients who received transplantation, the remaining 30 ASXL1-mutated patients analyzed retained the same mutation during serial follow-ups, including the one (patient 1) whose original mutation could be detected only by a sensitive gene-cloning technique, but not by direct sequencing, at the time of disease progression. These findings suggest that ASXL1 mutation may constitute an early hit in the pathogenesis of MDS. Similar to our findings, most ASXL1 mutations detected at leukemic transformation of myeloproliferative neoplasm patients were already present at chronic phase.14 Interestingly, during disease progression, ASXL1-mutated patients frequently acquired other novel genetic alterations, most commonly RUNX1 and NRAS mutations, (Table 3) or had chromosomal evolution indicating the additional genetic aberrations contributed to the progression of MDS in these patients. On the other hand, only one (patient 107) of the 80 patients without ASXL1 mutation at diagnosis acquired novel ASXL1 mutation at the time of AML transformation. Although no ASXL1 mutation could be detected at diagnosis even using a more sensitive cloning technique, we could not exclude the possibility that minor subpopulations of cells with the ASXL1 mutant existed initially. Another one patient (patient 108) with no detectable ASXL1 mutation at diagnosis by direct sequencing had in fact low level of ASXL1 mutant as shown by a more sensitive TA-cloning technique and the mutant expanded at disease progression. Altogether, these findings imply that ASXL1 mutations may have little, if any, role in the progression of MDS in ASXL1-wild patients.
ASXL1 mutation was shown to predict poor outcome in WHO-defined MDS and CMML patients,10, 42 and was associated with a reduced time to AML transformation.40 Thol et al.23 and Bejar et al.24 further demonstrated ASXL1 mutation as an independent poor prognostic factor in MDS patients. In our study, ASXL1-mutated MDS patients had poorer OS and higher rates of AML progression, especially in lower-risk patients, but not in the higher-risk ones. ASXL1 mutation was an independent poor prognostic factor irrespective of age, IPSS-R and mutations of RUNX1, NRAS, DNMT3A, IDH and SRSF2. Intriguingly, in MDS patients with normal karyotype, ASXL1 mutation was also an independent poor prognostic factor. ASXL1 mutation is thus helpful for risk stratification of MDS patients with normal karyotype, which is categorized as an intermediate-risk cytogenetic group.
In summary, ASXL1 mutations were detected in a substantial portion of MDS patients and were closely associated with trisomy 8 and mutations of RUNX1, EZH2, IDH, NRAS, JAK2, SETBP1 and SRSF2. The presence of ASXL1 mutations predicted shorter survival, especially in the patients with lower-risk MDS. For patients with normal karyotype, the mutation was also an independent poor prognostic factor for OS. Sequential study during the clinical course showed ASXL1-mutated patients retained the original ASXL1 mutation, but frequently acquired other novel genetic alterations during disease evolution.
References
Greenberg P, Cox C, LeBeau MM, Fenaux P, Morel P, Sanz G et al. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood 1997; 89: 2079–2088.
Hofmann WK, Koeffler HP . Myelodysplastic syndrome. Annu Rev Med 2005; 56: 1–16.
Acquaviva C, Gelsi-Boyer V, Birnbaum D . Myelodysplastic syndromes: lost between two states? Leukemia 2010; 24: 1–5.
Kulasekararaj AG, Mohamedali AM, Mufti GJ . Recent advances in understanding the molecular pathogenesis of myelodysplastic syndromes. Br J Haematol 2013; 162: 587–605.
Bejar R, Levine R, Ebert BL . Unraveling the molecular pathophysiology of myelodysplastic syndromes. J Clin Oncol 2011; 29: 504–515.
Sloand EM, Rezvani K . The role of the immune system in myelodysplasia: implications for therapy. Semin Hematol 2008; 45: 39–48.
Raaijmakers MH . Myelodysplastic syndromes: revisiting the role of the bone marrow microenvironment in disease pathogenesis. Int J Hematol 2012; 95: 17–25.
Rocquain J, Carbuccia N, Trouplin V, Raynaud S, Murati A, Nezri M et al. Combined mutations of ASXL1, CBL, FLT3, IDH1, IDH2, JAK2, KRAS, NPM1, NRAS, RUNX1, TET2 and WT1 genes in myelodysplastic syndromes and acute myeloid leukemias. BMC Cancer 2010; 10: 401.
Chou WC, Huang HH, Hou HA, Chen CY, Tang JL, Yao M et al. Distinct clinical and biological features of de novo acute myeloid leukemia with additional sex comb-like 1 (ASXL1) mutations. Blood 2010; 116: 4086–4094.
Gelsi-Boyer V, Trouplin V, Adelaide J, Bonansea J, Cervera N, Carbuccia N et al. Mutations of polycomb-associated gene ASXL1 in myelodysplastic syndromes and chronic myelomonocytic leukaemia. Br J Haematol 2009; 145: 788–800.
Martinez-Aviles L, Besses C, Alvarez-Larran A, Torres E, Serrano S, Bellosillo B . TET2, ASXL1, IDH1, IDH2, and c-CBL genes in JAK2- and MPL-negative myeloproliferative neoplasms. Ann Hematol 2012; 91: 533–541.
Graubert T, Walter MJ . Genetics of myelodysplastic syndromes: new insights. Hematology Am Soc Hematol Educ Program 2011; 2011: 543–549.
Fisher CL, Randazzo F, Humphries RK, Brock HW . Characterization of Asxl1, a murine homolog of additional sex combs, and analysis of the Asx-like gene family. Gene 2006; 369: 109–118.
Abdel-Wahab O, Manshouri T, Patel J, Harris K, Yao J, Hedvat C et al. Genetic analysis of transforming events that convert chronic myeloproliferative neoplasms to leukemias. Cancer Res 2010; 70: 447–452.
Pena PV, Davrazou F, Shi X, Walter KL, Verkhusha VV, Gozani O et al. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 2006; 442: 100–103.
Abdel-Wahab O, Adli M, LaFave LM, Gao J, Hricik T, Shih AH et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 2012; 22: 180–193.
Inoue D, Kitaura J, Togami K, Nishimura K, Enomoto Y, Uchida T et al. Myelodysplastic syndromes are induced by histone methylation-altering ASXL1 mutations. J Clin Invest 2013; 123: 4627–4640.
Abdel-Wahab O, Gao J, Adli M, Dey A, Trimarchi T, Chung YR et al. Deletion of Asxl1 results in myelodysplasia and severe developmental defects in vivo. J Exp Med 2013; 210: 2641–2659.
Wang J, Li Z, He Y, Pan F, Chen S, Rhodes S et al. Loss of Asxl1 leads to myelodysplastic syndrome-like disease in mice. Blood 2013, e-pub ahead of print 19 November 2013.
Dey A, Seshasayee D, Noubade R, French DM, Liu J, Chaurushiya MS et al. Loss of the tumor suppressor BAP1 causes myeloid transformation. Science 2012; 337: 1541–1546.
Scheuermann JC, de Ayala Alonso AG, Oktaba K, Ly-Hartig N, McGinty RK, Fraterman S et al. Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature 2010; 465: 243–247.
Boultwood J, Perry J, Pellagatti A, Fernandez-Mercado M, Fernandez-Santamaria C, Calasanz MJ et al. Frequent mutation of the polycomb-associated gene ASXL1 in the myelodysplastic syndromes and in acute myeloid leukemia. Leukemia 2010; 24: 1062–1065.
Thol F, Friesen I, Damm F, Yun HY, Weissinger EM, Krauter J et al. Prognostic significance of ASXL1 mutations in patients with myelodysplastic syndromes. J Clin Oncol 2011; 29: 2499–2506.
Bejar R, Stevenson K, Abdel-Wahab O, Galili N, Nilsson B, Garcia-Manero G et al. Clinical effect of point mutations in myelodysplastic syndromes. N Engl J Med 2011; 364: 2496–2506.
Chou WC, Tang JL, Lin LI, Yao M, Tsay W, Chen CY et al. Nucleophosmin mutations in de novo acute myeloid leukemia: the age-dependent incidences and the stability during disease evolution. Cancer Res 2006; 66: 3310–3316.
Chen CY, Lin LI, Tang JL, Tsay W, Chang HH, Yeh YC et al. Acquisition of JAK2, PTPN11, and RAS mutations during disease progression in primary myelodysplastic syndrome. Leukemia 2006; 20: 1155–1158.
Shiah HS, Kuo YY, Tang JL, Huang SY, Yao M, Tsay W et al. Clinical and biological implications of partial tandem duplication of the MLL gene in acute myeloid leukemia without chromosomal abnormalities at 11q23. Leukemia 2002; 16: 196–202.
Tang JL, Hou HA, Chen CY, Liu CY, Chou WC, Tseng MH et al. AML1/RUNX1 mutations in 470 adult patients with de novo acute myeloid leukemia: prognostic implication and interaction with other gene alterations. Blood 2009; 114: 5352–5361.
Hou HA, Huang TC, Lin LI, Liu CY, Chen CY, Chou WC et al. WT1 mutation in 470 adult patients with acute myeloid leukemia: stability during disease evolution and implication of its incorporation into a survival scoring system. Blood 2010; 115: 5222–5231.
Chou WC, Hou HA, Chen CY, Tang JL, Yao M, Tsay W et al. Distinct clinical and biologic characteristics in adult acute myeloid leukemia bearing the isocitrate dehydrogenase 1 mutation. Blood 2010; 115: 2749–2754.
Chou WC, Lei WC, Ko BS, Hou HA, Chen CY, Tang JL et al. The prognostic impact and stability of Isocitrate dehydrogenase 2 mutation in adult patients with acute myeloid leukemia. Leukemia 2011; 25: 246–253.
Hou HA, Kuo YY, Liu CY, Chou WC, Lee MC, Chen CY et al. DNMT3A mutations in acute myeloid leukemia: stability during disease evolution and clinical implications. Blood 2012; 119: 559–568.
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet 2010; 42: 722–726.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 2011; 478: 64–69.
Wu SJ, Kuo YY, Hou HA, Li LY, Tseng MH, Huang CF et al. The clinical implication of SRSF2 mutation in patients with myelodysplastic syndrome and its stability during disease evolution. Blood 2012; 120: 3106–3111.
Hou HA, Kuo YY, Tang JL, Chou WC, Yao M, Lai YJ et al. Clinical implications of the SETBP1 mutation in patients with primary myelodysplastic syndrome and its stability during disease progression. Am J Hematol 2013, e-pub ahead of print 11 October 2013; doi:10.1002/ajh.23611.
Tien HF, Wang CH, Lin MT, Lee FY, Liu MC, Chuang SM et al. Correlation of cytogenetic results with immunophenotype, genotype, clinical features, and ras mutation in acute myeloid leukemia: a study of 235 Chinese patients in Taiwan. Cancer Genet Cytogenet 1995; 84: 60–68.
Chen CY, Lin LI, Tang JL, Ko BS, Tsay W, Chou WC et al. RUNX1 gene mutation in primary myelodysplastic syndrome—the mutation can be detected early at diagnosis or acquired during disease progression and is associated with poor outcome. Br J Haematol 2007; 139: 405–414.
Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Sole F et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood 2012; 120: 2454–2465.
Gelsi-Boyer V, Trouplin V, Roquain J, Adelaide J, Carbuccia N, Esterni B et al. ASXL1 mutation is associated with poor prognosis and acute transformation in chronic myelomonocytic leukaemia. Br J Haematol 2010; 151: 365–375.
Abdel-Wahab O, Pardanani A, Patel J, Wadleigh M, Lasho T, Heguy A et al. Concomitant analysis of EZH2 and ASXL1 mutations in myelofibrosis, chronic myelomonocytic leukemia and blast-phase myeloproliferative neoplasms. Leukemia 2011; 25: 1200–1202.
Calzada A, Sacristan M, Sanchez E, Bueno A . Cdc6 cooperates with Sic1 and Hct1 to inactivate mitotic cyclin-dependent kinases. Nature 2001; 412: 355–358.
Acknowledgements
This work was partially sponsored by grants NSC 100–2314-B002-057-MY3, NSC 101-2325-B002-028 and NSC 100-2628 -B-002-003-MY3 from the National Science Council (Taiwan), DOH99-TD-C-111-001 from the Department of Health (Taiwan) and NTUH 102P06 and UN101-014 and 102-015 from the Department of Medical Research, National Taiwan University Hospital.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Author contributions
T-C Chen was responsible for literature collection, data management and interpretation, statistical analysis and manuscript writing; H-AH was responsible for study design, literature collection, data management and interpretation, statistical analysis and manuscript writing; C-YL was responsible for statistical analysis and interpretation of the statistical findings; Y-YK and M-C L were responsible for mutation analysis and interpretation; C-YC, W-CC, M-Y, S-YH, J-LT, B-SK, S-CH, S-JW, WT and Y-CC contributed patient samples and clinical data; M-CL, M-HT, C-FH, Y-CC, C-YL, F-YL and M-CL performed the gene mutation and chromosomal studies and H-FT planned, designed and coordinated the study over the entire period and wrote the manuscript.
Supplementary Information accompanies this paper on Blood Cancer Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Chen, TC., Hou, HA., Chou, WC. et al. Dynamics of ASXL1 mutation and other associated genetic alterations during disease progression in patients with primary myelodysplastic syndrome. Blood Cancer Journal 4, e177 (2014). https://doi.org/10.1038/bcj.2013.74
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bcj.2013.74
Keywords
This article is cited by
-
SRSF2 mutation cooperates with ASXL1 truncated alteration to accelerate leukemogenesis
Leukemia (2024)
-
ASXL1 mutations with serum EPO levels predict poor response to darbepoetin alfa in lower-risk MDS: W-JHS MDS01 trial
International Journal of Hematology (2022)
-
Distinct clinico-biological features in AML patients with low allelic ratio FLT3-ITD: role of allogeneic stem cell transplantation in first remission
Bone Marrow Transplantation (2022)
-
Risk factors affect accurate prognosis in ASXL1-mutated acute myeloid leukemia
Cancer Cell International (2021)
-
Prognostic value of ASXL1 mutations in patients with primary myelofibrosis and its relationship with clinical features: a meta-analysis
Annals of Hematology (2021)