Abstract
Members of the Rho family of GTPases are key regulators of the actin cytoskeleton. In particular, activated Rac1 stimulates membrane dorsal ruffle formation in response to platelet-derived growth factor (PDGF). Abl-interactor (Abi)-1 and βPIX, a guanine nucleotide exchange factor for Rac1, localise at these Rac1-induced actin structures and play important roles in the induction of membrane dorsal ruffling in response to PDGF in fibroblasts. Here, we demonstrate a novel interaction between Abi-1 and βPIX using the yeast two-hybrid system, in vitro pull-down assays, and in vivo co-immunoprecipitation experiments. In vitro, the C-terminal fragment of βPIX interacted with Abi-1, while in vivo the N-terminal fragment of βPIX interacted with Abi-1. The biological function of this interaction was investigated in mouse fibroblasts in response to PDGF stimulation. Abi-1 and βPIX co-localised in the cytoplasm and to membrane dorsal ruffles after PDGF treatment. We show that the co-expression of Abi-1 and truncated forms of βPIX in mouse fibroblasts blocked PDGF-induced membrane dorsal ruffles. Together, these results show that the interaction between Abi-1 and βPIX is involved in the formation of growth factor-induced membrane dorsal ruffles.
Similar content being viewed by others
Introduction
Regulation of the actin cytoskeleton plays a crucial role in many cellular functions such as cell shape change, cell motility, cell adhesion, and cytokinesis 1. Small GTPases of the Rho family are key regulators in the transduction of signals from membrane receptors to the actin cytoskeleton 2. Their activation requires the exchange of GDP for GTP, triggered by the action of guanine nucleotide exchange factors (GEFs) in response to extracellular signals like the platelet-derived growth factor (PDGF) 3. Active RhoA proteins regulate stress fibre formation and the assembly of focal adhesions 4, while active Rac1 regulates membrane ruffle, lamellipodia formation, and focal complexes 5, and active Cdc42 regulates filopodia formation 6.
The Abl-interactor (Abi) proteins were shown to localise to sites of de novo actin polymerisation at the tips of lamellipodia and filopodia in migrating cells 7. The Abi protein family 8 is an emerging class of cytoplasmic molecular adaptors with an Src homology 3 (SH3) domain, similar to Grb2 or Nck 9. The human Abi-1 protein was initially identified in a yeast two-hybrid screen as interaction partner of Eps8, a substrate for growth factor receptor tyrosine kinases 10. Abi-1 contains several PxxP (proline-rich region) motifs that bind to the SH3 domain of c-Abl 11, Grb4 12, spectrin 13, and synaptojanin 14. Abi-1 also contains a PxxDY (unconventional proline-rich region) motif, which has been shown to bind to Eps8 15. Several sequences rich in proline, glutamic acid, serine, and threonine (PEST sequences), which function as signals for rapid intracellular proteolysis, are also present 16. The SH3 domain at the C-terminus of Abi-1 interacts with the proline-rich domains of c-Abl 11, 17, 18 and Sos-1 19. Abi-1 is implicated in the regulation of c-Abl, which is involved in membrane dorsal ruffling in PDGF-treated fibroblasts 20.
Abi-1 has been shown to participate in the formation of membrane dorsal ruffles in response to PDGF 21. One such mechanism has been elucidated. The PDGF signal is transmitted from Ras to Rac1 through a trimeric complex containing Abi-1, Eps8, and Sos-1, in which Abi-1 holds Eps8 and Sos-1 together 19, 21, 22. More specifically, Abi-1 recruits the phosphatidylinositol 3-kinase (PI3-K), through a direct interaction with the p85 subunit of PI3-K, to a multimolecular signalling complex including Eps8 and Sos-1. The recruitment of PI3-K and its catalytic product phosphatidylinositol 3, 4, 5 phosphates (PIP3), which regulates the activity of the GEF protein Sos-1, are indispensable for the activation of Rac1 and actin cytoskeleton remodelling in response to PDGF 23. Thus, it can be easily imagined that as Abi-1 is implicated in the regulation of PDGF-induced membrane dorsal ruffling, other GEF proteins besides Sos-1 could also be involved in this process.
The human βPIX protein, designated as a p21-activated kinase (PAK)-Interacting eXchange protein, belongs to the family of GEFs 24, 25. βPIX, a GEF for Rac1, contains an SH3 domain followed by a Dbl homology (DH) domain in tandem with a pleckstrin homology (PH) domain (Figure 1). The DH domain is directly responsible for the guanine nucleotide exchange activity and activation of the small Rho GTPases 26, 27. However, the DH and PH domains in βPIX have been shown to function as independent units in vitro 27. βPIX also contains several PxxP motifs and a GIT1-binding domain, which is involved in the interaction with various cytoskeleton proteins, such as the G-protein-coupled receptor kinase-interacting target protein (GIT), the paxillin-kinase linker protein (p95PKL) and the cool-associated, tyrosine-phosphorylated protein 28, 29, 30. A leucine zipper motif has been characterised at the C-terminal end of the protein and shown to mediate the formation of βPIX homodimers 31, 32.
Abi-1 and βPIX constructs. Schematic representation of the various constructs used in this work. For yeast two-hybrid analysis, the indicated DNA fragments were cloned into the pB42AD and the pLexA vectors, downstream of the activation domain and the DNA-binding domain, respectively. For pull-down assays, Abi-1 constructs were cloned into the pGex-4T3 bacterial expression vector containing an N-terminal GST tag, and βPIX coding sequences were cloned into the pET28c vector. These constructs were also cloned into the pCEP4 mammalian expression vector for immunoprecipitation experiments and subcellular localisation studies of Abi-1 and βPIX. βPIX constructs were cloned with an N-terminal HA tag and Abi-1 constructs with a N-terminal Flag tag. aa: amino acids; Δ: deletion; FL: full-length; CC: coiled-coiled domain; HHR: homeo-domain homologous region; PxxP: proline-rich region; PxxDY: unconventional proline-rich region; SR: serine-rich region; SH3: Src Homology 3 domain; C-ter: C-terminus; N-ter: N-terminus; DH: Dbl homology domain; PH: pleckstrin homology domain; GBD: GIT1-binding domain; and LZ: leucine zipper. Black underlined lines in Abi-1 represented sequences rich in proline, glutamic acid, serine, and threonine (PEST sequences).
The formation of βPIX homodimers is important for its localisation to the cell periphery in order to drive the formation of membrane ruffles 31, 32. Thus, βPIX is implicated in cytoskeleton remodelling important in cell migration 33. Extracellular signals, e.g., from the PDGF receptor to cytoskeletal rearrangement are transmitted via βPIX through the activation of Rac1 24. However, little is known about the associated signalling cascade and upstream activators of βPIX leading to the activation of Rac1. Binding of βPIX to PAK leads to Rac1 activation phenotype 34. βPIX binds also to p95PKL and GIT1, which link PAK to the central focal complex components, paxillin and focal adhesion kinase. These protein complexes promote focal adhesion turnover and Rac1-dependent mobility 28, 29.
In this study, we demonstrate that Abi-1 and βPIX interact in vitro and in vivo. Since Abi-1 and PIX play important roles in the PDGF-induced Rac1-dependent membrane ruffling, we investigated the biological function of the Abi-1 and βPIX interaction in mouse fibroblasts in response to PDGF stimulation. Our results show that the interaction between Abi-1 and βPIX is involved in PDGF-induced membrane dorsal ruffling.
Materials and Methods
Constructs
The different Abi-1 and βPIX constructs are presented in Figure 1. The cDNA fragments encoding for βPIX and Abi-1 were cloned into EcoRI and XhoI sites of the pB42AD and pLexA yeast expression vectors (Clontech). The cDNA fragment encoding for Abi-1 full-length (Abi-1 FL) sequence was cloned into the EcoRI and XhoI sites of the bacterial expression vector pGex-4T3 (Invitrogen), whereas the cDNA fragments encoding for the different βPIX constructs were cloned into the BamHI and HindIII sites of the bacterial expression vector pET28c (Novagen). The different Abi-1 and βPIX cDNA fragments were cloned into the NheI and XhoI sites of the mammalian expression vector pCEP4 (Invitrogen).
Two-hybrid interaction
The yeast two-hybrid assay was performed using the competent EGY48[p8op-LacZ] yeast strain prepared as follows. An overnight culture was used to inoculate a fresh culture at an OD600 of 0.1, which was grown to an OD600 of 0.6 and harvested by centrifugation. Cells were washed in 1/10 culture volume Buffer A (1 M sorbitol, 1 mM bicine, and 3% ethyleneglycol, pH 8.35), re-suspended in 1/100 volume Buffer A, aliquoted and stored first for 30 min at −20 °C, and then frozen at −70 °C. Frozen competent yeast cells were transformed by heat shock with 1 μg of the bait (pLexA) and prey (pB42AD) plasmid cDNAs, 5 μl denaturised salmon's sperm (10 μg/μl), and 5 μl histamine. After 5 min shaking at 37 °C and addition of 1 ml Buffer B (40% PEG-1000 and 200 mM bicine, pH 8.35), the sample was incubated for 1 h at 30 °C with casual shaking. Cells were harvested by centrifugation, washed in 1 ml Buffer C (150 mM NaCl, 10 mM bicine, pH 8.35), re-suspended in 100 μl Buffer C, and plated on selective medium: Trp-, His-, Ura-, galactose-raffinose-X-Gal-containing medium for the activation of the LacZ reporter gene and Trp-, His-, Ura-, glucose-X-Gal-containing medium for negative control. A positive interaction between the bait and prey proteins results in LacZ activation, conversion of X-Gal substrate, and selectable blue colonies.
Cell cultures, transfections, and microinjection
Transient expression of proteins for biochemical analysis and immunofluorescence was achieved by Ca2+ phosphate transfection. One day prior to transfection, 1×106 293T cells were seeded on 6 cm plates in DMEM (Life Technologies) with 10% FCS, 1% penicillin/streptomycin, 10 mM HEPES (pH 7.4), and 4 mM L-glutamine, or 40 000 NIH3T3 cells were seeded on glass coverslips in 12-well plates in DMEM with 10% FCS, 1% penicillin/streptomycin, 1 mM sodium pyruvate, and 4 mM L-glutamine. Plasmid cDNAs were mixed with one volume of 250 mM CaCl2 and then with one volume of 2×HBS (50 mM HEPES-NaOH (pH 7.0), 1.5 mM Na2HPO4, and 273 mM NaCl). The preparation was added to the cells in medium supplemented with 25 μM chloroquine (Sigma). One day post-transfection, NIH3T3 cells were grown in serum starvation medium (0.5% FCS) for 24 h and then stimulated with 15 ng/ml PDGF-BB (Upstate Biotechnology) for 10 min at 37 °C with 5% CO2. Cells were fixed as described below.
Fifty ng/μl expression plasmid were microinjected into the nucleus of NIH3T3 cells seeded on glass coverslips as described above, using an Eppendorf microinjector. After 3 h, cells were serum starved for 24 h, treated with PDGF, and fixed as described below.
Immunofluorescence
Cells grown on glass coverslips were fixed with 3% PFA for 30 min at 37 °C or overnight at 4 °C. Cells were washed thrice with PBS, blocked and permeabilised with 0.2% Triton X-100, and 10% goat serum in PBS for 30 min at room temperature. Cells were incubated with primary antibodies: anti-HA Y-11 rabbit polyclonal (1:100, Santa Cruz), and/or anti-Flag M2 mouse monoclonal (1:2 500, Sigma), diluted in 3% goat serum and 0.05% Tween 20 in PBS for 1 h at room temperature. Cells were washed twice with PBS, blocked for 5 min, and incubated with the secondary antibodies: Cy3-conjugated goat anti-mouse (1:100, Dianova), and/or Amca-conjugated anti-rabbit (1:100, Jackson), diluted as previously described, for 50 min. Cells were washed twice with PBS, blocked for 5 min, and incubated with Alexa Fluor 488 phalloidin for 45 min (1:100, Molecular Probe). Preparations were analysed under a fluorescence microscope equipped with a 63× and a 100× oil immersion objectives (Leica), and a digital camera. Images were processed using the TCS Software and Adobe Photoshop.
Immunoprecipitation and immunoblotting
For precipitation of endogenous Abi-1, NIH3T3 cells were grown in serum starvation medium (0.5% FCS) for 24 h and were then stimulated with 15 ng/ml PDGF-BB for 10 min at 37 °C with 5% CO2. Cells were lysed in 100 mM NaCl, 10 mM sodium phosphate buffer, 2 mM EGTA, 1 mM EDTA, 10 mM NaF, 10 mM Na2P2O7, 0.5% deoxycholate, 0.05% SDS, 5% glycerol, 1% NP-40, plus protease inhibitor mix (1 μg/ml aprotinin, 0.5 μg/ml leupeptin, 1 mM pefabloc, and 10 μM pepstatin), pH 7.2. Lysates were incubated or not with Abi-1 monoclonal antibody (generous gift from Giorgio Scita, Istituto FIRC di Oncologia Molecolare, Milan, Italy). In case of immunoprecipitation of overexpressed proteins, cells were harvested 48 h post-transfection, washed with cold PBS, and re-suspended in 600 μl Single-Detergent-Buffer-1 (20 mM Tris-HCl, pH 7.5, 150 mM NaCl, and 1% NP-40 with protease inhibitor mix) on ice for 10 min. Cells were then passed through a syringe twice and centrifuged at 13 000 rpm. for 15 min at 4 °C. Cell lysates were pre-incubated and rotated with the indicated agarose-coupled antibody, anti-HA (Santa Cruz) or anti-Flag (M2, Sigma), overnight at 4 °C. Immunoprecipitated proteins bound to the agarose matrix were then washed in Single-Detergent-Buffer-1 without protease inhibitors, boiled in 2× sample solution, separated in SDS-PAGE, and detected by immunoblotting with the antibodies specific for the protein's tag. Anti-HA Y-11 rabbit polyclonal (1:1 000), anti-Flag M2 mouse monoclonal (1:5 000), goat anti-rabbit peroxidase, and goat anti-mouse peroxidase (1:3 000, Amersham Life Science) were used.
In vitro translation
In vitro translation was performed using the TnT® Coupled Reticulocyte Lysate System (Promega) according to the manufacturer's protocol. For a reaction in 50 μl H2O, 1 μg plasmid cDNA coding for different deletion constructs of PIX and 5 μl [35S]methionine (10 μCi/μl, Amersham/Pharmacia) were mixed. Protein translation was reported by SDS-PAGE, Coomassie staining, and exposure of the gels to films.
GST protein expression, purification, and GST pull-down assay
GST-Abi-1 full-length (GST-Abi-1 FL) and GST were expressed in BL21-CodonPlus (DE3)-RP Escherichia coli strain (Stratagene) by the addition of 0.5 mM IPTG. For cell lysis, bacteria were re-suspended on ice in 2 ml cold PBS with protease inhibitor mix and sonicated four times for 30 s. Purification of GST-Abi-1 FL and GST was performed with a 50% gluthatione sepharose 4B matrix (Pharmacia) at 4 °C and rotated for 2 h. The matrix containing the bound protein was washed twice with PBS. The proteins were eluted with 20 mM gluthatione, 50 mM Tris-HCl (pH 8.0), 100 mM NaCl, and dialysed in PBS overnight at 4 °C. The dialysed protein was shock frozen and stored at −70 °C. For pull-down assay the purified GST-Abi-1 FL and GST were bound to gluthatione sepharose 4B matrix, as described previously, washed with cold PBS, and incubated with the different in vitro translated βPIX proteins for 30 min at 4 °C with shaking. Samples were centrifuged and pellets washed thrice with the Single-Detergent-Buffer-2 (20 mM Tris-HCl, pH 7.5, 500 mM NaCl, and 1% NP-40 with protease inhibitor mix). Pellets were re-suspended in 50 μl Single-Detergent-Buffer-2, 10 μl 2× sample solution and samples were analysed by SDS-PAGE. Then, the gels were stained with Coomassie, dried, and exposed overnight.
Results
A new interaction between βPIX and Abi-1
In order to identify new interaction partners for PAKs, a yeast two-hybrid screen was performed, which resulted in the isolation of Abi-1 and βPIX (unpublished data). We also performed yeast two-hybrid screens to find out whether Abi-1 and βPIX not only interact with PAKs but may also be with each other. Different fragments of βPIX were fused to the activation domain of the pB42AD vector. The Abi-1 FL sequence was fused downstream to the DNA-binding domain of the pLexA vector (Figure 1). The expression of all constructs was verified by Western blot analysis (not shown). As a negative control, the different βPIX fusion proteins and the Abi-1 FL fusion protein were each tested for potential self-activation of the LacZ reporter gene and for potential interactions with a non-related protein, lamin C or caspase-9, respectively. No self-activation by any of these fusion proteins was detected, nor did these fusions show any interaction with non-related negative control proteins (data not shown). To prove that the system is useful for detecting proteins interacting with βPIX, the interaction of βPIX with its known partner PAK2 25 was tested and verified.
The different βPIX constructs were co-transformed with the Abi-1 FL construct into yeast and assayed for interaction. Full-length Abi-1 and full-length βPIX clearly interacted as indicated by the positive β-galactosidase reaction (Figure 2A). Neither the βPIX hybrid protein containing the N-terminal domain from amino acids 1 to 410 nor the βPIX hybrid proteins containing the DH and PH domains or the PH domain alone interacted with Abi-1. However, an interaction was detected with the βPIX hybrid protein containing a deletion of the SH3 domain and with the βPIX hybrid protein containing the C-terminal domain from amino acids 410 to 646. Thus, in this assay the interaction of βPIX with Abi-1 was mediated by the C-terminus of βPIX.
Identification of an interaction between Abi-1 and βPIX. The pB42AD vector expressing the indicated βPIX fusions and the pLexA vector expressing the indicated Abi-1 fusions were co-transformed into the yeast strain EGY48[p8op-LacZ] to test for interaction in a two-hybrid assay as described. (A) Interaction of βPIX or its fragments with full-length Abi-1 on X-Gal- (X) or glucose- (G) containing medium. (B) Interaction of Abi-1 or its fragments with full-length βPIX on X-Gal- (X) or glucose- (G) containing medium. The blue coloured colonies were observed when interaction had occurred. Clones containing the bait and the prey plasmids were amplified and the expression of the activation domain fusion proteins was repressed by the presence of glucose (G) in the medium. Abbreviations are explained in the legend of Figure 1.
Yeast two-hybrid assays were also performed with different constructs of Abi-1 described in Figure 1. These constructs were co-transformed in yeast with the βPIX full-length construct, in order to define the domain of interaction in Abi-1. Surprisingly, only full-length Abi-1 interacted with βPIX (Figure 2B), suggesting that the interaction between Abi-1 and βPIX may require a particular conformation of Abi-1, which is destroyed in Abi-1 deletion proteins.
Interaction between βPIX and Abi-1 in vitro
The interaction between Abi-1 and βPIX observed in the yeast two-hybrid assay was confirmed using in vitro GST pull-down assays. GST-Abi-1 FL was constructed and used to pull down various radioactively labelled βPIX constructs. As shown in Figure 3, the interaction between Abi-1 and βPIX full-length was confirmed. Neither the N-terminal fragment, nor the fragment containing the PH domain nor the fragment containing the DH and PH domains of βPIX interacted with full-length Abi-1. However, Abi-1 FL interacted with the C-terminal fragment and the construct that lacked the SH3 domain of βPIX. These results are consistent with the results obtained in the yeast two-hybrid analyses indicating that the C-terminus of βPIX mediates the interaction between βPIX and Abi-1 in vitro.
Characterisation of the interaction between Abi-1 and βPIX in vitro. The interaction between full-length Abi-1 and full-length βPIX or its fragments was assayed in pull-down experiments using a GST-Abi-1 full-length (GST-Abi-1 FL) fusion protein. The full-length Abi-1 protein was fused to the C-terminus of the GST tag and expressed from the pGex-4T3 bacterial expression vector. GST alone or GST-Abi-1 FL were coupled to sepharose and then incubated with radioactively labelled in vitro translated full-length βPIX or its fragments (for the description of constructs see Figure 1). Samples were eluted and separated by SDS-PAGE; gels were dried and exposed to an X-ray film overnight.
Interaction between βPIX and Abi-1 in vivo
In order to determine whether Abi-1 and βPIX could interact in vivo, co-immunoprecipitation of endogenous Abi-1 and βPIX was performed in NIH3T3 cells. NIH3T3 cells were serum starved for 24 h and treated with PDGF before Abi-1 was precipitated with a monoclonal Abi-1 antibody. βPIX clearly co-precipitated with Abi-1 in PDGF-treated cells (Figure 4A), suggesting an interaction of the two proteins in these cells. Since the βPIX anti-serum was inefficient in precipitating mouse βPIX from NIH3T3 cells, the human βPIX was overexpressed and immunoprecipitated from HeLa cells. Endogenous Abi-1 co-precipitated with βPIX (Supplement 1), suggesting that both proteins are organised in the same complex.
In vivo interaction between Abi-1 and βPIX. (A) NIH3T3 were serum starved and treated with (+) or without (−) PDGF before lysis and immunoprecipitation of Abi-1 with monoclonal antibody. The levels of precipitated Abi-1 and co-precipitated βPIX were monitored in the lysates (WCL) and the Abi-1 immunoprecipitations (IP Abi-1) with (+) and without (−) Abi-1 antibody (Abi-1 AB). (B, C) 293T cells were mock transfected (cells) or transfected with 5 μg of each of the different deletion constructs of βPIX and Abi-1 FL in (B) or with each of the different deletion constructs of Abi-1 and βPIX FL in (C). (B) Flag-Abi-1 FL was immunoprecipitated (IP) with the anti-Flag antibody from the total cell lysates, and protein complexes were detected by immunoblotting with anti-Flag and anti-HA antibodies for Flag-Abi-1 FL and HA-βPIX, respectively. (C) HA-βPIX FL was immunoprecipitated with the anti-HA antibody from the total cell lysates and protein complexes were detected by immunoblotting as described above. The black arrow points to Flag-Abi-1 FL and the broken arrow to the antibody heavy chain. The two bottom panels in (B) and (C) show the expression of the diverse constructs, the top panels show the immunoprecipitated or co-immunoprecipitated proteins. The diverse constructs for Abi-1 and βPIX are represented in Figure 1.
To map the domain of interaction in Abi-1 and βPIX, 293T cells were double transfected either with Flag-Abi-1 FL and different HA-tagged βPIX constructs (Figure 4B) or with βPIX full-length and different Abi-1 constructs (Figure 4C). The full-length HA-βPIX protein co-immunoprecipitated with the full-length Flag-Abi-1 protein. Interestingly, co-transfections of different βPIX constructs with Abi-1 FL showed that the βPIX N-terminal fragment, the construct lacking the SH3 domain, and the construct containing the DH and PH domains of βPIX interacted with Abi-1 FL. The C-terminal construct of βPIX, which interacted in vitro with Abi-1 FL (Figure 3), did not interact with Abi-1 FL in vivo (Figure 4B). However, in confirmation of the yeast two-hybrid results, none of the Abi-1 deletion constructs interacted with βPIX full-length (Figure 4C). Thus, whereas further evidence could be obtained that a conformational epitope of Abi-1 is involved in the interaction with βPIX, the βPIX fragments interacting with Abi-1 in vitro and in vivo differ.
Abi-1 and βPIX co-localise to membrane dorsal ruffles in PDGF-induced NIH3T3 cells
Rac1 regulates PDGF-induced membrane dorsal ruffling 5. It has been demonstrated that Abi-1 protein induces membrane dorsal ruffling in fibroblasts through Rac1 activation in response to PDGF 19. Overexpression of the βPIX protein, known to function as a GEF protein for Rac1, induces membrane dorsal ruffling in fibroblasts through Rac1 24, 31, 34. In order to investigate the subcellular distribution of Abi-1 and βPIX in NIH3T3 cells upon PDGF treatment, the expression vectors encoding, either for βPIX full-length or Abi-1 FL, were microinjected into fibroblasts. Serum-starved cells were induced with PDGF, and βPIX and Abi-1 were visualised by indirect immunofluorescence. Abi-1 localised in the cytoplasm in non-treated cells (Figure 5B) and to membrane dorsal ruffles in PDGF-treated cells (Figure 5D). The same localisation pattern was observed for βPIX (Figure 5E-5H), seen localised in the cytoplasm (Figure 5F), and then at membrane dorsal ruffles in response to PDGF treatment (Figure 5H).
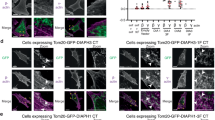
Subcellular localisation of Abi-1 and βPIX upon PDGF induction. Abi-1 (A-D) and βPIX (E-H) localise to membrane dorsal ruffles. Nuclei of mouse fibroblasts (NIH3T3) were microinjected with 50 ng/μl of the expression vector for Abi-1 full-length (Abi-1 FL) or βPIX full-length (βPIX FL). Cells were serum starved and treated with (C, D, G, H) or without (A, B, E, F) PDGF. Abi-1 and βPIX co-localise to membrane dorsal ruffles (I-P). Nuclei of NIH3T3 were co-microinjected with expression vectors for βPIX FL and Abi-1 FL. Cells were serum-starved and treated with (M, N, O, P) or without (I, J, K, L) PDGF. Cells were fixed and stained for immunofluorescence with Alexa-conjugated phalloidin to detect F-actin (green), with the anti-Flag antibody to detect Abi-1 (yellow), and with the anti-HA antibody to detect βPIX (blue). Arrows point to membrane dorsal ruffles. Bars, 10 μm.
Then, the subcellular distribution of Abi-1 and βPIX full-length proteins co-expressed in NIH3T3 cells was analysed. Abi-1 and βPIX exhibited a similar diffuse staining pattern in the cytoplasm of non-treated cells (Figure 5J, 5K, and 5L), suggesting that both proteins localise in the same compartment. In PDGF-treated cells, a substantial portion of Abi-1 and βPIX localised to membrane dorsal ruffles (Figure 5N, 5O, and 5P), compared to the diffuse cytoplasmic localisation of Abi-1 and βPIX in serum-starved, untreated cells (Figure 5J, 5K, and 5L). Taken together, the co-localisation of Abi-1 and βPIX to sites of membrane dorsal ruffling and the interaction between Abi-1 and βPIX suggest a role for this interaction in the formation of membrane dorsal ruffles in response to PDGF.
The interaction between Abi-1 and βPIX is involved in the formation of PDGF-induced membrane dorsal ruffles
To examine the involvement of the interaction between Abi-1 and βPIX in the induction of membrane dorsal ruffles in PDGF-treated NIH3T3 cells, the effect of co-expression of Abi-1 FL, either with the N-terminal or the C-terminal fragments of βPIX, was investigated.
The effects of the N-terminal and C-terminal fragments of βPIX were first studied. Therefore, the vector encoding for the N-terminal or the C-terminal fragment of βPIX was transfected in NIH3T3 cells. Serum-starved transfected cells were treated with PDGF and overexpressed proteins were visualised by immunofluorescence microscopy. As shown in Figure 6A-6H, the N-terminal fragment of βPIX (Figure 6B) as well as the C-terminal fragment (Figure 6F) localised in the cytoplasm of serum-starved, non-treated cells. Cells overexpressing either the N-terminal fragment (Figure 6D) or the C-terminal fragment of βPIX (Figure 6H) did not induce membrane dorsal ruffling after PDGF treatment. Thus, upon treatment with PDGF, the N-terminal and the C-terminal fragment of βPIX inhibited the formation of membrane dorsal ruffles, suggesting both domains are required for this PDGF-mediated response.
Functional interaction of Abi-1 and βPIX. (A-H) The N-terminal fragment (βPIX N-ter) and C-terminal fragment (βPIX C-ter) of βPIX inhibit membrane dorsal ruffle formation in response to PDGF. NIH3T3 were transfected with the expression vector for βPIX N-ter (A-D) or βPIX C-ter (E-H) for 1 day. Cells were serum starved and treated with (C, D, G, H) or without (A, B, E, F) PDGF. (I-P) Co-expression of the N-terminal fragment of βPIX with Abi-1 full-length (Abi-1 FL) inhibits membrane dorsal ruffle formation in response to PDGF. NIH3T3 were co-transfected with expression vectors for βPIX N-ter and Abi-1 FL for 1 day. Cells were serum-starved and treated with (M, N, O, P) or without (I, J, K, L) PDGF. (Q-X) Co-expression of the C-terminal fragment of βPIX with Abi-1 FL inhibits membrane dorsal ruffle formation in response to PDGF. NIH3T3 were co-transfected with expression vectors for βPIX C-ter and Abi-1 FL for 1 day. Cells were serum starved and treated with (U, V, W, X) or without (Q, R, S, T) PDGF. Cells were fixed and stained with Alexa-conjugated phalloidin to detect F-actin (green), with the anti-Flag antibody to detect Abi-1 (yellow), and with the anti-HA antibody to detect βPIX (blue). Arrows point to membrane dorsal ruffles and asterisks identify the transfected cell. Bars, 10 μm. (Y) Quantification of the experiments shown from (I) to (X). NIH3T3 cells expressing Abi-1 FL and the indicated full-length or truncated βPIX proteins were stimulated with PDGF, and were scored for the presence or absence of membrane dorsal ruffling. Data represent the mean of three independent experiments. Bars indicate the standard errors.
Then, NIH3T3 cells were co-transfected either with the vectors encoding for Abi-1 FL and the N-terminal fragment of βPIX (Figure 6I-6P) or with the vectors encoding for Abi-1 FL and the C-terminal fragment of βPIX (Figure 6Q-6X). The number of cells forming ruffles upon PDGF stimulation was similar in cells expressing either Abi-1 FL or βPIX full-length alone (80% and 83%, respectively). Cells co-expressing Abi-1 FL and the N-terminal fragment of βPIX did not exhibit morphological changes in response to PDGF (Figure 6M, 6N, 6O, and 6P). Whereas 78.4% of cells overexpressing Abi-1 FL and βPIX full-length formed membrane dorsal ruffles after PDGF induction, only 11.74% of cells overexpressing Abi-1 FL and the βPIX N-terminal fragment showed this phenotype (Figure 6Y). A similar effect was observed with cells co-expressing Abi-1 FL and the C-terminal fragment of βPIX (Figure 6U, 6V, 6W, and 6X), since only 5.5% of the cells showed this phenotype (Figure 6Y). Thus, both the N-terminal and C-terminal fragment of βPIX functioned in a dominant-negative manner and blocked PDGF-induced membrane dorsal ruffling.
Discussion
We report here a novel interaction between Abi-1 and βPIX. This interaction was shown in yeast two-hybrid experiments, in vitro, by GST pull-down experiments and, in vivo, by co-immunoprecipitation of the endogenous and overexpressed proteins.
Our data indicate that the C-terminal fragment of βPIX interacts with Abi-1 in vitro. The C-terminal fragment of βPIX contains different domains including several PxxP motifs, whereas Abi-1 contains in its C-terminal part an SH3 domain. The fact that SH3 domain and PxxP motifs are typical interaction modules in protein-protein interactions 35 suggests that βPIX interacts with the SH3 domain of Abi-1 through the PxxP motifs present in the C-terminal region. This has been shown for another binding partner of Abi-1, the Sos-1 protein 19, 22.
Interestingly, Abi-1 and Sos-1 interact via different domains in vitro and in vivo. In vivo, Sos-1 interacts with the N-terminus of Abi-1 and not with the SH3 domain contained in the C-terminus 36. We find a similar case for the interaction of βPIX and Abi-1. In contrast to the results of initial in vitro binding studies, in vivo Abi-1 co-immunoprecipitated with the N-terminal fragment of βPIX and not with the C-terminal fragment. Nevertheless, it cannot be excluded that the C-terminal fragment of βPIX interacts with Abi-1 in vivo. The expressed C-terminal fragment may acquire a conformation different from the full-length protein, which is not recognised by Abi-1, or another protein may have masked the Abi-1-interacting domain in the C-terminus of βPIX.
The results obtained in vivo showed that the SH3 domain of βPIX is not required for an interaction with Abi-1, while the fragment of βPIX containing the DH and PH domains bound to Abi-1. DH domains contain the GEF activity, whereas PH domains are known to bind specifically to phosphatidylinositol-4, 5-biphosphate, which targets the protein to membranes 37. This is now realised to be a property of a small number of PH domain-containing proteins, which bind phosphoinositides only very weakly. This suggests that either most PH domains require assistance from other parts of the protein in order to achieve their function or the physiological ligands of PH domains are not limited to phosphoinositides 38.
In the present study, it was not possible to define the domain of interaction in Abi-1 in vitro and in vivo. As soon as the full-length protein Abi-1 is deleted either in its N-terminus or C-terminus, no interaction was detected with βPIX. It can be hypothesised that deletions in Abi-1 destroy a conformation of the protein necessary for binding to βPIX. Another possible explanation for this observation is the limited stability of the truncated Abi-1 proteins. Several bands of low molecular weight reacted with the anti-Flag antibody (Figure 4C) indicating the presence of degraded Abi-1 products, an observation made before by others 13. It is possible that the rapid intracellular proteolysis is mediated by the PEST sequences present in Abi-1, which can function as a signal for proteasome-mediated degradation.
The co-localisation of Abi-1 and βPIX in the cytoplasm confirms the relevance of the interaction between the two proteins. After induction with PDGF, both Abi-1 and βPIX co-localised to membrane dorsal ruffles. To study the function of the interaction between Abi-1 and βPIX in PDGF-induced membrane dorsal ruffles in NIH3T3 cells, the subcellular localisation and the effects of overexpressed βPIX deletion constructs and Abi-1 FL were investigated. The N-terminal fragment of βPIX overexpressed with Abi-1 FL in PDGF-induced cells shows a diffuse cytoplasmic localisation and abrogates membrane dorsal ruffling. The same phenotype was observed when the C-terminal fragment of βPIX was overexpressed with Abi-1 FL.
What is the possible significance of the effect of the N-terminal fragment of βPIX? The N-terminal construct containing the domain of interaction with Abi-1, in vivo, may act in a dominant-negative manner, abrogating the interaction between Abi-1 and endogenous βPIX. The diffuse cytoplasmic localisation of the N-terminal fragment of βPIX after PDGF induction could result from the sequestration of downstream effectors molecules essential for membrane dorsal ruffling. Additionally, βPIX homodimerise in vitro and in vivo through its leucine zipper domain located in the C-terminal extremity of the protein 32. The ability of βPIX to homodimerise is necessary for the βPIX-mediated membrane dorsal ruffle formation in NIH3T3 fibroblasts in response to PDGF 31. Although the N-terminal fragment of βPIX contains the GEF activity required for the activation of Rac1-induced membrane ruffles 24 and interacts with Abi-1 in vivo (Figure 4), the absence of the dimerisation domain in the N-terminal construct could also explain the loss of membrane dorsal ruffling.
How may the C-terminal fragment of βPIX interfere with dorsal ruffle formation? It is possible that the C-terminal fragment of βPIX interacts with Abi-1 in vivo in a way that does not allow co-immunoprecipitation. In this case, the C-terminal fragment of βPIX would also function in a dominant-negative manner by competing for binding of endogenous or overexpressed Abi-1 with endogenous βPIX. As the C-terminal fragment of βPIX does not possess the GEF activity, induction of membrane dorsal ruffles in response to PDGF cannot be achieved directly through the interaction between Abi-1 and the βPIX C-terminal fragment. Other reports where βPIX full-length was overexpressed in fibroblasts with a C-terminal fragment of βPIX have shown the loss of membrane dorsal ruffles in response to PDGF, suggesting the importance of the N-terminal domain of βPIX 31, 32. Furthermore, the C-terminal fragment of βPIX containing PxxP motifs could directly compete for the binding of Sos-1 to the SH3 domain of Abi-1 19. Indeed, in serum-starved fibroblasts, an interaction between Abi-1 and Sos-1 induces membrane dorsal ruffles in response to PDGF 19.
How might this interaction play a role in PDGF-induced actin cytoskeleton reorganisation? We propose two models. The first model implicates the interaction between Abi-1 and βPIX as part of an alternative Ras-dependent pathway. Activation of the PDGF receptor induces a signalling cascade involving the interaction between Abi-1 and Sos-1. Sos-1 functions as a Rac-GEF downstream of Ras in a complex with the proteins Eps8, Abi-1, and PI3-K 19, 21, 23. Within this complex, Abi-1 serves as a scaffold bridging together Eps8 and Sos-1. In an alternative route of the same pathway, Abi-1 may interact and activate the Rac-GEF βPIX and thereby induces membrane dorsal ruffles. A second model of the interaction between Abi-1 and βPIX involves a Ras-independent pathway. Rac1 can also be activated through a Ras-independent pathway involving a direct recruitment of PI-3K to activated receptor tyrosine kinases to induce Rac1-mediated cytoskeletal reorganisation 21, 39. The interaction between the p85 regulatory subunit of PI3-K and Abi-1 has been shown to be important for Abi-1-dependent Rac1 activation and membrane dorsal ruffle formation 23. Thus, Abi-1 may serve as a scaffold bridging PI3-K and βPIX to allow activation of the GEF activity of βPIX.
In summary, a novel interaction between Abi-1 and βPIX was identified in this study, which may coordinate the formation of membrane dorsal ruffles in response to PDGF. The translation of growth signals to changes in the architecture of the cytoskeleton is a highly complex process involving different signalling pathways and multiple protein complexes. Since the whole process is highly dynamic, it is to be expected that the formation and resolution of different signalling complexes initiated by apparent redundant pathways is also highly dynamic. In this scenario, the multiple pathways initiated by growth receptors rather reflect the complexity of signalling machinery required for the coordinate regulation of the cytoskeleton than redundant parallel ways to the same outcome.
References
Etienne-Manneville S, Hall A . Rho GTPases in cell biology. Nature 2002; 420:629–635.
Van Aelst L, D'Souza-Schorey C . Rho GTPases and signaling networks. Genes Dev 1997; 11:2295–2322.
Schmidt A, Hall A . Guanine nucleotide exchange factors for Rho GTPases: turning on the switch. Genes Dev 2002; 16:1587–1609.
Ridley AJ, Hall A . The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell 1992; 70:389–399.
Ridley AJ, Paterson HF, Johnston CL, Diekmann D, Hall A . The small GTP-binding protein rac regulates growth factor-induced membrane ruffling. Cell 1992; 70:401–410.
Nobes CD, Hall A . Rho, rac, and cdc42 GTPases regulate the assembly of multimolecular focal complexes associated with actin stress fibers, lamellipodia, and filopodia. Cell 1995; 81:53–62.
Stradal T, Courtney KD, Rottner K, et al. The Abl interactor proteins localize to sites of actin polymerization at the tips of lamellipodia and filopodia. Curr Biol 2001; 11:891–895.
Ichigotani Y, Fujii K, Hamaguchi M, Matsuda S . In search of a function for the E3B1/Abi2/Argbp1/NESH family (Review). Int J Mol Med 2002; 9:591–595.
Smith JM, Katz S, Mayer BJ . Activation of the Abl tyrosine kinase in vivo by Src homology 3 domains from the Src homology 2/Src homology 3 adaptor Nck. J Biol Chem 1999; 274:27956–27962.
Fazioli F, Minichiello L, Matoska V, et al. Eps8, a substrate for the epidermal growth factor receptor kinase, enhances EGF-dependent mitogenic signals. EMBO J 1993; 12:3799–3808.
Biesova Z, Piccoli C, Wong WT . Isolation and characterization of e3B1, an eps8 binding protein that regulates cell growth. Oncogene 1997; 14:233–241.
Cowan CA, Henkemeyer M . The SH2/SH3 adaptor Grb4 transduces B-ephrin reverse signals. Nature 2001; 413:174–179.
Ziemnicka-Kotula D, Xu J, Gu H, et al. Identification of a candidate human spectrin Src homology 3 domain-binding protein suggests a general mechanism of association of tyrosine kinases with the spectrin-based membrane skeleton. J Biol Chem 1998; 273:13681–13692.
So CW, So CK, Cheung N, et al. The interaction between EEN and Abi-1, two MLL fusion partners, and synaptojanin and dynamin: implications for leukaemogenesis. Leukemia 2000; 14:594–601.
Mongiovi AM, Romano PR, Panni S, et al. A novel peptide-SH3 interaction. EMBO J 1999; 18:5300–5309.
Rogers S, Wells R, Rechsteiner M . Amino acid sequences common to rapidly degraded proteins: the PEST hypothesis. Science 1986; 234:364–368.
Dai Z, Pendergast AM . Abi-2, a novel SH3-containing protein interacts with the c-Abl tyrosine kinase and modulates c-Abl transforming activity. Genes Dev 1995; 9:2569–2582.
Wang B, Mysliwiec T, Krainc D, et al. Identification of ArgBP1, an Arg protein tyrosine kinase binding protein that is the human homologue of a CNS-specific Xenopus gene. Oncogene 1996; 12:1921–1929.
Scita G, Nordstrom J, Carbone R, et al. EPS8 and E3B1 transduce signals from Ras to Rac. Nature 1999; 401:290–293.
Plattner R, Kadlec L, DeMali KA, Kazlauskas A, Pendergast AM . c-Abl is activated by growth factors and Src family kinases and has a role in the cellular response to PDGF. Genes Dev 1999; 13:2400–2411.
Scita G, Tenca P, Areces LB, et al. An effector region in Eps8 is responsible for the activation of the Rac-specific GEF activity of Sos-1 and for the proper localization of the Rac-based actin-polymerizing machine. J Cell Biol 2001; 154:1031–1044.
Innocenti M, Tenca P, Frittoli E, et al. Mechanisms through which Sos-1 coordinates the activation of Ras and Rac. J Cell Biol 2002; 156:125–136.
Innocenti M, Frittoli E, Ponzanelli I, et al. Phosphoinositide 3-kinase activates Rac by entering in a complex with Eps8, Abi1, and Sos-1. J Cell Biol 2003; 160:17–23.
Manser E, Loo TH, Koh CG, et al. PAK kinases are directly coupled to the PIX family of nucleotide exchange factors. Mol Cell 1998; 1:183–192.
Bagrodia S, Taylor SJ, Jordon KA, Van Aelst L, Cerione RA . A novel regulator of p21-activated kinases. J Biol Chem 1998; 273:23633–23636.
Cerione RA, Zheng Y . The Dbl family of oncogenes. Curr Opin Cell Biol 1996; 8:216–222.
Aghazadeh B, Zhu K, Kubiseski TJ, et al. Structure and mutagenesis of the Dbl homology domain. Nat Struct Biol 1998; 5:1098–1107.
Zhao ZS, Manser E, Loo TH, Lim L . Coupling of PAK-interacting exchange factor PIX to GIT1 promotes focal complex disassembly. Mol Cell Biol 2000; 20:6354–6363.
Turner CE, Brown MC, Perrotta JA, et al. Paxillin LD4 motif binds PAK and PIX through a novel 95-kD ankyrin repeat, ARF-GAP protein: A role in cytoskeletal remodeling. J Cell Biol 1999; 145:851–863.
Bagrodia S, Bailey D, Lenard Z, et al. A tyrosine-phosphorylated protein that binds to an important regulatory region on the cool family of p21-activated kinase-binding proteins. J Biol Chem 1999; 274:22393–22400.
Kim S, Lee SH, Park D . Leucine zipper-mediated homodimerization of the p21-activated kinase-interacting factor, beta Pix. Implication for a role in cytoskeletal reorganization. J Biol Chem 2001; 276:10581–10584.
Koh CG, Manser E, Zhao ZS, Ng CP, Lim L . Beta1PIX, the PAK-interacting exchange factor, requires localization via a coiled-coil region to promote microvillus-like structures and membrane ruffles. J Cell Sci 2001; 114:4239–4251.
Ridley AJ, Schwartz MA, Burridge K, et al. Cell migration: integrating signals from front to back. Science 2003; 302:1704–1709.
Obermeier A, Ahmed S, Manser E, et al. PAK promotes morphological changes by acting upstream of Rac. EMBO J 1998; 17:4328–4339.
Mayer BJ . SH3 domains: complexity in moderation. J Cell Sci 2001; 114:1253–1263.
Fan PD, Goff SP . Abl interactor 1 binds to sos and inhibits epidermal growth factor- and v-Abl-induced activation of extracellular signal-regulated kinases. Mol Cell Biol 2000; 20:7591–7601.
Harlan JE, Hajduk PJ, Yoon HS, Fesik SW . Pleckstrin homology domains bind to phosphatidylinositol-4,5-bisphosphate. Nature 1994; 371:168–170.
Lemmon MA, Ferguson KM, Abrams CS . Pleckstrin homology domains and the cytoskeleton. FEBS Lett 2002; 513:71–76.
Wennstrom S, Siegbahn A, Yokote K, et al. Membrane ruffling and chemotaxis transduced by the PDGF beta-receptor require the binding site for phosphatidylinositol 3' kinase. Oncogene 1994; 9:651–660.
Acknowledgements
We thank Giorgio Scita (Istituto FIRC di Oncologia Molecolare, Milan, Italy) for providing the Abi-1 monoclonal antibody used for immunoprecipitation of endogenous Abi-1. We thank Trent Fowler for critical reading of the manuscript. This work was supported by Grant Nos. RU631/2-1 and RU631/2-2 to T.R.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Campa, F., Machuy, N., Klein, A. et al. A new interaction between Abi-1 and βPIX involved in PDGF-activated actin cytoskeleton reorganisation. Cell Res 16, 759–770 (2006). https://doi.org/10.1038/sj.cr.7310091
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310091
Keywords
This article is cited by
-
Low expression of Abelson interactor-1 is linked to acquired drug resistance in Bcr-Abl-induced leukemia
Leukemia (2014)
-
PDGF: the nuts and bolts of signalling toolbox
Tumor Biology (2011)