Abstract
Plastins/fimbrins are conserved actin-bundling proteins contributing to motility, cytokinesis and other cellular processes by organizing strikingly different actin assemblies as in aligned bundles and branched networks. We propose that this ability of human plastins stems from an allosteric communication between their actin-binding domains (ABD1/2) engaged in a tight spatial association. Here we show that ABD2 can bind actin three orders of magnitude stronger than ABD1, unless the domains are involved in an equally strong inhibitory engagement. A mutation mimicking physiologically relevant phosphorylation at the ABD1–ABD2 interface greatly weakened their association, dramatically potentiating actin cross-linking. Cryo-EM reconstruction revealed the ABD1–actin interface and enabled modeling of the plastin bridge and domain separation in parallel bundles. We predict that a strong and tunable allosteric inhibition between the domains allows plastins to modulate the cross-linking strength, contributing to remodeling of actin assemblies of different morphologies defining the unique place of plastins in actin organization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The cryo-EM map of ABD1PLS3–F-actin complex was deposited with accession code EMD-25371 in the EMDB. Figures using protein structures were generated using PDB accession numbers: 6ANU, 1AOA, and 6VEC and AlphaFold entry: https://alphafold.ebi.ac.uk/entry/P13797. Source data are provided with this paper.
References
Shinomiya, H. Plastin family of actin-bundling proteins: its functions in leukocytes, neurons, intestines, and cancer. Int. J. Cell Biol. 2012, 213492 (2012).
Delanote, V., Vandekerckhove, J. & Gettemans, J. Plastins: versatile modulators of actin organization in (patho)physiological cellular processes. Acta Pharmacol. Sin. 26, 769–779 (2005).
Grimm-Günter, E. M. et al. Plastin 1 binds to keratin and is required for terminal web assembly in the intestinal epithelium. Mol. Biol. Cell 20, 2549–2562 (2009).
Lin, C. S., Shen, W., Chen, Z. P., Tu, Y. H. & Matsudaira, P. Identification of I-plastin, a human fimbrin isoform expressed in intestine and kidney. Mol. Cell. Biol. 14, 2457–2467 (1994).
Drenckhahan, D. et al. Three different actin filament assemblies occur in every hair cell: each contains a specific actin crosslinking protein. J. Cell Biol. 112, 641–651 (1991).
Sobin, A. & Flock, A. Immunohistochemical identification and localization of actin and fimbrin in vestibular hair cells in the normal guinea pig and in a strain of the waltzing guinea pig. Acta Otolaryngol. 96, 407–412 (1983).
Diaz-Horta, O. et al. Novel variant p.E269K confirms causative role of PLS1 mutations in autosomal dominant hearing loss. Clin. Genet 96, 575–578 (2019).
Morgan, A. et al. Mutations in PLS1, encoding fimbrin, cause autosomal dominant nonsyndromic hearing loss. Hum. Mutat. 40, 2286–2295 (2019).
Jones, S. L., Wang, J., Turck, C. W. & Brown, E. J. A role for the actin-bundling protein L-plastin in the regulation of leukocyte integrin function. Proc. Natl Acad. Sci. USA 95, 9331–9336 (1998).
Ma, T., Sadashivaiah, K., Madayiputhiya, N. & Chellaiah, M. A. Regulation of sealing ring formation by L-plastin and cortactin in osteoclasts. J. Biol. Chem. 285, 29911–29924 (2010).
Morley, S. C. The actin-bundling protein L-plastin: a critical regulator of immune cell function. Int. J. Cell Biol. 2012, 935173 (2012).
Morley, S. C. et al. The actin-bundling protein L-plastin dissociates CCR7 proximal signaling from CCR7-induced motility. J. Immunol. 184, 3628–3638 (2010).
Todd, E. M., Deady, L. E. & Morley, S. C. The actin-bundling protein L-plastin is essential for marginal zone B cell development. J. Immunol. 187, 3015–3025 (2011).
Wabnitz, G. H. et al. Costimulation induced phosphorylation of L-plastin facilitates surface transport of the T cell activation molecules CD69 and CD25. Eur. J. Immunol. 37, 649–662 (2007).
Wabnitz, G. H. et al. Sustained LFA-1 cluster formation in the immune synapse requires the combined activities of L-plastin and calmodulin. Eur. J. Immunol. 40, 2437–2449 (2010).
Lin, C. S., Park, T., Chen, Z. P. & Leavitt, J. Human plastin genes: comparative gene structure, chromosome location, and differential expression in normal and neoplastic cells. J. Biol. Chem. 268, 2781–2792 (1993).
Chaijan, S., Roytrakul, S., Mutirangura, A. & Leelawat, K. Matrigel induces L-plastin expression and promotes L-plastin-dependent invasion in human cholangiocarcinoma cells. Oncol. Lett. 8, 993–1000 (2014).
Foran, E., McWilliam, P., Kelleher, D., Croke, D. T. & Long, A. The leukocyte protein L-plastin induces proliferation, invasion and loss of E-cadherin expression in colon cancer cells. Int. J. Cancer 118, 2098–2104 (2006).
Klemke, M. et al. Phosphorylation of ectopically expressed L-plastin enhances invasiveness of human melanoma cells. Int. J. Cancer 120, 2590–2599 (2007).
Riplinger, S. M. et al. Metastasis of prostate cancer and melanoma cells in a preclinical in vivo mouse model is enhanced by L-plastin expression and phosphorylation. Mol. Cancer 13, 1–12 (2014).
Garbett, D. et al. T-Plastin reinforces membrane protrusions to bridge matrix gaps during cell migration. Nat. Commun. 11, 4818 (2020).
Schwebach, C. L. et al. Osteogenesis imperfecta mutations in plastin 3 lead to impaired calcium regulation of actin bundling. Bone Res. 8, 21 (2020).
Li, N. et al. Actin-bundling protein plastin 3 is a regulator of ectoplasmic specialization dynamics during spermatogenesis in the rat testis. FASEB J. 29, 3788–3805 (2015).
Xue, F., Janzen, D. M. & Knecht, D. A. Contribution of filopodia to cell migration: a mechanical link between protrusion and contraction. Int. J. Cell Biol. 2010, 507821 (2010).
Hagiwara, M. et al. Interaction of activated Rab5 with actin-bundling proteins, L- and T-plastin and its relevance to endocytic functions in mammalian cells. Biochem. Biophys. Res. Commun. 407, 615–619 (2011).
Charras, G. T., Hu, C. K., Coughlin, M. & Mitchison, T. J. Reassembly of contractile actin cortex in cell blebs. J. Cell Biol. 175, 477–490 (2006).
Schwebach, C. L., Kudryashova, E. & Kudryashov, D. S. Plastin 3 in X-linked osteoporosis: imbalance of Ca(2+)-dependent regulation is equivalent to protein loss. Front Cell Dev. Biol. 8, 635783 (2020).
Hosseinibarkooie, S. et al. The power of human protective modifiers: PLS3 and CORO1C unravel impaired endocytosis in spinal muscular atrophy and rescue SMA phenotype. Am. J. Hum. Genet 99, 647–665 (2016).
Ralser, M. et al. Ataxin-2 and huntingtin interact with endophilin-A complexes to function in plastin-associated pathways. Hum. Mol. Genet. 14, 2893–2909 (2005).
Lyon, A. N. et al. Calcium binding is essential for plastin 3 function in Smn-deficient motoneurons. Hum. Mol. Genet. 23, 1990–2004 (2014).
Oprea, G. E. et al. Plastin 3 is a protective modifier of autosomal recessive spinal muscular atrophy. Science 320, 524–527 (2008).
Morandell, J. et al. Cul3 regulates cytoskeleton protein homeostasis and cell migration during a critical window of brain development. Nat. Commun. 12, 3058 (2021).
Begue, E. et al. Inducible expression and pathophysiologic functions of T-plastin in cutaneous T-cell lymphoma. Blood 120, 143–154 (2012).
Velthaus, A. et al. The actin binding protein plastin-3 is involved in the pathogenesis of acute myeloid leukemia. Cancers (Basel) 11, 1663 (2019).
Xin, Z. et al. PLS3 predicts poor prognosis in pancreatic cancer and promotes cancer cell proliferation via PI3K/AKT signaling. J. Cell. Physiol. 235, 8416–8423 (2020).
Ma, Y. et al. Plastin 3 down-regulation augments the sensitivity of MDA-MB-231 cells to paclitaxel via the p38 MAPK signalling pathway. Artif. Cells Nanomed. Biotechnol. 47, 685–695 (2019).
Korenbaum, E. & Rivero, F. Calponin homology domains at a glance. J. Cell Sci. 115, 3543–3545 (2002).
Schwebach, C. L., Agrawal, R., Lindert, S., Kudryashova, E. & Kudryashov, D. S. The roles of actin-binding domains 1 and 2 in the calcium-dependent regulation of actin filament bundling by human plastins. J. Mol. Biol. 429, 2490–2508 (2017).
Ishida, H., Jensen, K. V., Woodman, A. G., Hyndman, M. E. & Vogel, H. J. The calcium-dependent switch helix of L-plastin regulates actin bundling. Sci. Rep. 7, 40662 (2017).
Namba, Y., Ito, M., Zu, Y., Shigesada, K. & Maruyama, K. Human T cell L-plastin bundles actin filaments in a calcium dependent manner. J. Biochem. 112, 503–507 (1992).
Shirayama, S. & Numata, O. Tetrahymena fimbrin localized in the division furrow bundles actin filaments in a calcium-independent manner. J. Biochem. 134, 591–598 (2003).
Kovar, D. R., Staiger, C. J., Weaver, E. A. & McCurdy, D. W. AtFim1 is an actin filament crosslinking protein from Arabidopsis thaliana. Plant J. 24, 625–636 (2000).
Nakano, K., Satoh, K., Morimatsu, A., Ohnuma, M. & Mabuchi, I. Interactions among a fimbrin, a capping protein, and an actin-depolymerizing factor in organization of the fission yeast actin cytoskeleton. Mol. Biol. Cell 12, 3515–3526 (2001).
Klein, M. G. et al. Structure of the actin crosslinking core of fimbrin. Structure 12, 999–1013 (2004).
Goldsmith, S. C. et al. The structure of an actin-crosslinking domain from human fimbrin. Nat. Struct. Biol. 4, 708–712 (1997).
Galkin, V. E., Orlova, A., Cherepanova, O., Lebart, M. C. & Egelman, E. H. High-resolution cryo-EM structure of the F-actin-fimbrin/plastin ABD2 complex. Proc. Natl Acad. Sci. USA 105, 1494–1498 (2008).
Volkmann, N., DeRosier, D., Matsudaira, P. & Hanein, D. An atomic model of actin filaments cross-linked by fimbrin and its implications for bundle assembly and function. J. Cell Biol. 153, 947–956 (2001).
Dyson, M. R., Shadbolt, S. P., Vincent, K. J., Perera, R. L. & McCafferty, J. Production of soluble mammalian proteins in Escherichia coli: identification of protein features that correlate with successful expression. BMC Biotechnol. 4, 32 (2004).
Kudryashov, D. S. et al. Cofilin cross-bridges adjacent actin protomers and replaces part of the longitudinal F-actin interface. J. Mol. Biol. 358, 785–797 (2006).
Kudryashov, D. S., Phillips, M. & Reisler, E. Formation and destabilization of actin filaments with tetramethylrhodamine-modified actin. Biophys. J. 87, 1136–1145 (2004).
Kudryashov, D. S. et al. Connecting actin monomers by iso-peptide bond is a toxicity mechanism of the Vibrio cholerae MARTX toxin. Proc. Natl Acad. Sci. USA 105, 18537–18542 (2008).
Smith, H. et al. Rounding out the understanding of ACD toxicity with the discovery of cyclic forms of actin oligomers. Int. J. Mol. Sci. 22, 718 (2021).
Niesen, F. H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Avery, A. W. et al. Structural basis for high-affinity actin binding revealed by a beta-III-spectrin SCA5 missense mutation. Nat. Commun. 8, 1350 (2017).
Hanein, D. et al. An atomic model of fimbrin binding to F-actin and its implications for filament crosslinking and regulation. Nat. Struct. Biol. 5, 787–792 (1998).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Xu, X. et al. Mst1 kinase regulates the actin-bundling protein L-plastin to promote T cell migration. J. Immunol. 197, 1683–1691 (2016).
Lin, C. S., Lau, A. & Lue, T. F. Analysis and mapping of plastin phosphorylation. DNA Cell Biol. 17, 1041–1046 (1998).
Hornbeck, P. V. et al. PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res. 40, D261–D270 (2012).
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Klammer, M. et al. Phosphosignature predicts Dasatinib response in non-small cell lung cancer. Mol. Cell. Proteom. 11, 651–668 (2012).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Mertins, P. et al. Ischemia in tumors induces early and sustained phosphorylation changes in stress kinase pathways but does not affect global protein levels. Mol. Cell. Proteom. 13, 1690–1704 (2014).
Zhou, H. et al. Toward a comprehensive characterization of a human cancer cell phosphoproteome. J. Proteome Res 12, 260–271 (2013).
Watanabe, N. & Mitchison, T. J. Single-molecule speckle analysis of actin filament turnover in lamellipodia. Science 295, 1083–1086 (2002).
Kudryashova, E. et al. Actin cross-linking toxin is a universal inhibitor of tandem-organized and oligomeric G-actin binding proteins. Curr. Biol. 28, 1536–1547 e1539 (2018).
Skau, C. T. et al. Actin filament bundling by fimbrin is important for endocytosis, cytokinesis, and polarization in fission yeast. J. Biol. Chem. 286, 26964–26977 (2011).
Tsai, F. C. & Meyer, T. Ca2+ pulses control local cycles of lamellipodia retraction and adhesion along the front of migrating cells. Curr. Biol. 22, 837–842 (2012).
Song, J. et al. A cryo-tomography-based volumetric model of the actin core of mouse vestibular hair cell stereocilia lacking plastin 1. J. Struct. Biol. 210, 107461 (2020).
Leite, J. et al. Equatorial non-muscle myosin II and plastin cooperate to align and compact F-actin bundles in the cytokinetic ring. Front. Cell Dev. Biol. 8, 573393 (2020).
Zhu, C., Chen, Y. & Ju, L. A. Dynamic bonds and their roles in mechanosensing. Curr. Opin. Chem. Biol. 53, 88–97 (2019).
Martino, F., Perestrelo, A. R., Vinarsky, V., Pagliari, S. & Forte, G. Cellular mechanotransduction: from tension to function. Front. Physiol. 9, 824 (2018).
Anderson, C. A., Kovar, D. R., Gardel, M. L. & Winkelman, J. D. LIM domain proteins in cell mechanobiology. Cytoskeleton (Hoboken) 78, 303–311 (2021).
Kamioka, H., Sugawara, Y., Honjo, T., Yamashiro, T. & Takano-Yamamoto, T. Terminal differentiation of osteoblasts to osteocytes is accompanied by dramatic changes in the distribution of actin-binding proteins. J. Bone Miner. Res. 19, 471–478 (2004).
Weinbaum, S., Duan, Y., Satlin, L. M., Wang, T. & Weinstein, A. M. Mechanotransduction in the renal tubule. Am. J. Physiol. Ren. Physiol. 299, F1220–F1236 (2010).
Tseng, H. Y. et al. LCP1 preferentially binds clasped alphaMbeta2 integrin and attenuates leukocyte adhesion under flow. J. Cell Sci. 131, jcs218214 (2018).
Wabnitz, G. H. et al. LFA-1 cluster formation in T-cells depends on L-plastin phosphorylation regulated by P90(RSK) and PP2A. Cell. Mol. Life Sci. 78, 3543–3564 (2021).
Mei, L. et al. Structural mechanism for bi-directional actin crosslinking by T-plastin. Preprint at bioRxiv https://doi.org/10.1101/2021.12.07.471696 (2021).
Spudich, J. A. & Watt, S. The regulation of rabbit skeletal muscle contraction. J. Biol. Chem. 246, 4866–4871 (1971).
Heisler, D. B. et al. ACTIN-DIRECTED TOXIN. ACD toxin-produced actin oligomers poison formin-controlled actin polymerization. Science 349, 535–539 (2015).
Lu, J. & Pollard, T. D. Profilin binding to poly-l-proline and actin monomers along with ability to catalyze actin nucleotide exchange is required for viability of fission yeast. Mol. Biol. Cell 12, 1161–1175 (2001).
Kudryashova, E., Heisler, D., Zywiec, A. & Kudryashov, D. S. Thermodynamic properties of the effector domains of MARTX toxins suggest their unfolding for translocation across the host membrane. Mol. Microbiol. 92, 1056–1071 (2014).
Nolen, B. J. & Pollard, T. D. Structure and biochemical properties of fission yeast Arp2/3 complex lacking the Arp2 subunit. J. Biol. Chem. 283, 26490–26498 (2008).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schindelin, J., Rueden, C. T., Hiner, M. C. & Eliceiri, K. W. The ImageJ ecosystem: an open platform for biomedical image analysis. Mol. Reprod. Dev. 82, 518–529 (2015).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Kudryashova, E. et al. Human defensins facilitate local unfolding of thermodynamically unstable regions of bacterial protein toxins. Immunity 41, 709–721 (2014).
Purde, V., Busch, F., Kudryashova, E., Wysocki, V. H. & Kudryashov, D. S. Oligomerization affects the ability of human cyclase-associated proteins 1 and 2 to promote actin severing by cofilins. Int. J. Mol. Sci. 20, 5647 (2019).
Bairoch, A. The Cellosaurus, a cell-line knowledge resource. J. Biomol. Tech. 29, 25–38 (2018).
Uphoff, C. C. & Drexler, H. G. Detection of Mycoplasma contamination in cell cultures. Curr. Protoc. Mol. Biol. 106, 28.4.1–28.4.14 (2014).
Chu, J. et al. Non-invasive intravital imaging of cellular differentiation with a bright red-excitable fluorescent protein. Nat. Methods 11, 572–578 (2014).
Rizzo, M. A., Davidson, M. W. & Piston, D. W. Fluorescent protein tracking and detection: fluorescent protein structure and color variants. Cold Spring Harb. Protoc. 2009, pdb top63 (2009).
Watanabe, N. Fluorescence single-molecule imaging of actin turnover and regulatory mechanisms. Methods Enzymol. 505, 219–232 (2012).
Hatano, T. et al. Rapid production of pure recombinant actin isoforms in Pichia pastoris. J. Cell Sci. 131, jcs213827 (2018).
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank F. Wang (University of Virginia) for assistance in modeling parallel actin filaments linked by a plastin bridge and S. Dong (OSU) and L. Runyan (OSU) for assistance in protein purification. This work was supported by the National Institute of General Medical Sciences of the NIH under award numbers R01GM114666 (D.S.K.), R35GM122510 (E.H.E.) and a 2018 Pelotonia Graduate Fellowship Award at The Ohio State University Comprehensive Cancer Center (C.L.S.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
D.S.K. conceptualized the study. D.S.K., E.H.E. and C.L.S. acquired funding. D.S.K. and E.H.E. supervised the study and C.L.S, E.K., R.A. and W.Z carried out the investigations. C.L.S., E.K. and W.Z. carried out formal analysis and visualization. C.L.S, D.S.K. and E.K. wrote the original draft. All authors reviewed and edited the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Charles Sindelar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Florian Ullrich was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are avialable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 ABD2PLS2 exists in solution as a monomer and rescues polymerization of ACD-crosslinked actin oligomers.
(a-c) Sedimentation velocity analytical ultracentrifugation (SV-AUC) analysis of ABD2PLS2. Raw sedimentation profiles of absorbance at 280 nm versus radius (a) and residual plots (b) are shown. The distribution of sedimentation coefficients (c) indicates the presence of only monomeric species of ABD2PLS2 protein. (d) Actin nucleation activity of ABD2 of PLS2 was tested by TIRFM in the presence of profilin (PFN1)/F-actin. Error bars represent the SD of the mean; n is number of biologically independent samples examined over two independent experiments. ANOVA followed by multiple comparison tests (two-sided Student’s t-test) with Bonferroni correction was applied: asterisks indicate statistically significant difference (*p<0.017). Scale bars are 10 µm. (e) ABD2PLS2 rescues polymerization of ACD-cross-linked actin oligomers. G-actin was covalently cross-linked by addition of ACD toxin in the absence (Actin) or presence (Actin +ABD2) of ABD2. Following ACD treatment, cross-linked actin was allowed to polymerize by addition of Mg2+ and KCl and subjected to ultracentrifugation to separate non-polymerized soluble (S) and polymerized pellet (P) fractions on SDS-PAGE. ABD2 alone sample (ABD2) treated identically served as a negative control. Data for graph in d are available as source data.
Extended Data Fig. 2 ABD1/ABD2 interaction is not affected by salt.
The ABD1–ABD2 affinity in the presence of 30 and 130 mM KCl was determined by fluorescence anisotropy assays. Error bars represent the SD of the mean; n=3 independent experiments. Data for graph are available as source data.
Extended Data Fig. 3 L475P mutation diminishes F-actin bundling ability of PLS2.
(a) F-actin bundling and binding by PLS2WT and PLS2L475P was assessed by low- (bundling) and high- (binding) speed sedimentation. Low-speed co-sedimentation (panel Actin Bundling) detected F-actin bundles formed only in the presence of PLS2WT in the absence of Ca2+ (PLS2WT Actin Bundling 3s/3p vs 4s/4p), while PLS2L475P was unable to bundle F-actin regardless of Ca2+ [F-actin remained mainly in the supernatant (PLS2L475P Actin Bundling 3s/3p and 4s/4p)]. In high-speed co-sedimentation assays (panel Actin Binding), both constructs were co-pelleted with F-actin in the absence (Actin Binding 3s/3p) and presence of Ca2+ (Actin Binding 4s/4p) implying that L475P mutation does not affect F-actin binding mediated through ABD1. (b,c) For DSF experiments, protein preps of PLS constructs (lines ‘T’ on SDS-PAGE (b)) were further cleared to remove minor contamination of truncated fragments corresponding to RD–ABD1 (red arrow on SDS-PAGE (b)). This contamination resulted in an additional melting peak (Tm2, dotted lines on DSF graphs (c)) but was neither removable by ion exchange chromatography (due to similar pIs) nor by gel filtration (due to insufficiently different sizes). However, following heating to 55 °C and ultracentrifugation at 300,000g for 30 min at 4 °C to remove precipitate (lines ‘P’ on SDS-PAGE (b)), the supernatants (lines ‘S’ on SDS-PAGE (b)) containing cleared FL PLS constructs were free of the contaminants, displayed single DSF peaks (Tm1) and were otherwise indistinguishable from the originally prepped proteins (c). The cleared samples (lines ‘S’ on SDS-PAGE (b)) were used in DSF experiments (Fig. 3). Data for graphs in c are available as source data.
Extended Data Fig. 4 Cryo-EM reconstruction of ABD1/F-actin.
(a-d) Preparation of ABD1/F-actin sample for cryo-EM. To produce F-actin decorated with ABD1, recombinant human β-Actin carrying K50C and C374A mutations was purified from Pichia pastoris and polymerized as described in online Methods. Individual Cys residues were introduced on the Cys-null RD–ABD1PLS3 background (Supplementary Table 1) at the indicated positions resulting in constructs containing single cysteines (Q194C, A216C, L226C, or G229C). Asterisks indicate activation of either actin (Actin*) or RD–ABD1 constructs (194*, 216*, 226*, or 229*) using a corresponding cross-linking reagent [MTS-8-MTS (a), oPDM (c), or pPDM (d)]. Following the activation, 2.5 µM actin was mixed with 10-molar excess of an RD–ABD1 construct. The resulting cross-linked samples were resolved on 9% SDS-PAGE (a, c, d). Note that non-crosslinked actin (42 kDa) and RD–ABD1 constructs (43 kDa) have similar molecular weights resulting in similar mobility on SDS-PAGE. Anti-actin and anti-PLS3 western blotting was performed to confirm identity of the resulting bands; only the immunoblots for the samples boxed in a are shown in b. Red arrowheads indicate RD–ABD1/RD–ABD1 cross-links. Green arrowheads indicate successful formation of RD–ABD1/F-actin cross-links. The cross-linking of MTS-8-MTS-activated actin with the RD–ABD1 (boxed in a) was the most efficient as compared to other tested reagents/combinations, and the sample of MTS-8-MTS-activated actin cross-linked with RD–ABD1Q194C was subjected to cryo-EM. (e) ABD1/F-actin density map is colored according to the resolution in angstroms.
Extended Data Fig. 5 S406E phospho-mimetic mutation moderately increases nucleation activity of PLS2.
Actin nucleation activity of PLS2S406E at 50-nM (a; n=5) and 1-µM (b; n=8) was tested by TIRFM (n is number of biologically independent samples examined over two independent experiments). Error bars represent the SD of the mean. ANOVA followed by multiple comparison tests with Bonferroni correction was applied: asterisk indicates statistically significant difference (*p<0.025) compared to the actin control. Data for graphs in a,b are available as source data.
Extended Data Fig. 6 Intracellular localization of human plastins.
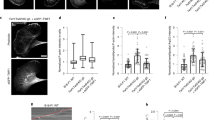
U2OS cells transiently co-transfected with the indicated mEmerald-tagged plastin constructs and a focal adhesion marker mCardinal-paxillin were fixed and counter-stained with TRITC-phalloidin and Hoechst. Boxed areas in (a) are enlarged in (b). Scale bars are 20 µm in (a) and 5 µm in (b).
Extended Data Fig. 7 Intracellular localization of human plastins.
To calculate the lamellipodial and total cell fluorescence density (fluorescence intensity per cell area), XTC cells were transiently transfected with either mEmerald-tagged PLS2-WT or PLS2-S406E (a). Cells were fragmented in ImageJ and the obtained masks were eroded by 12 px=2 µm (using Process → Binary → Options → Erode ImageJ tool); the resulting masks were used to generate selections to outline and measure the total cell fluorescence (b) and lamellipodial fluorescence, that is, fluorescence in the 2-µm-thick band at the cell edge (c). Scale bars are 10 µm.
Supplementary information
Supplementary Information
Supplementary Table 1
Supplementary Video 1
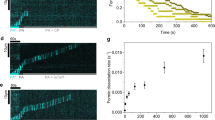
Human plastin isoforms undergo retrograde flow in the lamellipodia. SiMS TIRFM time-lapse imaging of XTC cells transiently transfected with mEmerald-tagged human plastin isoforms. Scale bars, 5 µm.
Supplementary Video 2
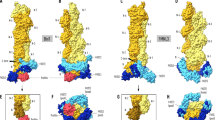
The domain reorientation is required for parallel, inregister actin bundle. Two modes of plastin are shown: the AlphaFold model of plastin core in the unbound state and the model for a crossbridge in actin bundle. CH1 is light blue, CH2 is dark blue, CH3 is light red, CH4 is dark red and actin is gray.
Source data
Source Data Fig. 1
Numerical Source Data.
Source Data Fig. 2
Numerical Source Data.
Source Data Fig. 3
Numerical Source Data.
Source Data Fig. 5
Numerical Source Data.
Source Data Fig. 6
Numerical Source Data.
Source Data Extended Data Fig. 1
Numerical Source Data.
Source Data Extended Data Fig. 2
Numerical Source Data.
Source Data Extended Data Fig. 3
Numerical Source Data.
Source Data Extended Data Fig. 5
Numerical Source Data.
Rights and permissions
About this article
Cite this article
Schwebach, C.L., Kudryashova, E., Agrawal, R. et al. Allosteric regulation controls actin-bundling properties of human plastins. Nat Struct Mol Biol 29, 519–528 (2022). https://doi.org/10.1038/s41594-022-00771-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00771-1
This article is cited by
-
A three-level regulatory mechanism of the aldo-keto reductase subfamily AKR12D
Nature Communications (2024)
-
The Dictyostelium discoideum FimA protein, unlike yeast and plant fimbrins, is regulated by calcium similar to mammalian plastins
Scientific Reports (2023)
-
Green finance in circular economy: a literature review
Environment, Development and Sustainability (2023)