ABSTRACT
Major advances have been made over the last decade in our understanding of the molecular basis of several cardiac conditions. Hypertrophic cardiomyopathy (HCM) was the first cardiac disorder in which a genetic basis was identified and as such, has acted as a paradigm for the study of an inherited cardiac disorder. HCM can result in clinical symptoms ranging from no symptoms to severe heart failure and premature sudden death. HCM is the commonest cause of sudden death in those aged less than 35 years, including competitive athletes. At least ten genes have now been identified, defects in which cause HCM. All of these genes encode proteins which comprise the basic contractile unit of the heart, i.e. the sarcomere. While much is now known about which genes cause disease and the various clinical presentations, very little is known about how these gene defects cause disease, and what factors modify the expression of the mutant genes. Studies in both cell culture and animal models of HCM are now beginning to shed light on the signalling pathways involved in HCM, and the role of both environmental and genetic modifying factors. Understanding these mechanisms will ultimately improve our knowledge of the basic biology of heart muscle function, and will therefore provide new avenues for treating cardiovascular disease in man.
Similar content being viewed by others
INTRODUCTION
Over the last decade, major advances have been made in our understanding of the molecular basis of cardiovascular disease. When the double helical structure of DNA was first described in 1953 by Watson and Crick in a two-page article in Nature1, no one could have predicted the tremendous impact this discovery would have in establishing the study of human genetic diseases. This discovery, coupled with the acceptance of Mendel's theory of inheritance, were two important landmarks in the development of the field of cardiovascular molecular genetics. In addition, several recent technological advances have fueled a surge in molecular genetic research in cardiovascular disease. Such advances include the understanding of biochemical components of DNA, the development of cloning techniques and DNA sequencing, the amplification of DNA by polymerase chain reaction (PCR), the identification of restriction enzymes to allow small pieces of DNA to be manipulated, and the undertaking of the Human Genome Project2. Today, molecular cardiology is characterised by the integration of high-technology laboratory studies and clinical medicine. Molecular genetics has redefined the aetiology and diagnostic criteria for numerous diseases and has led to the development of new, individualised treatment regimens for several cardiovascular diseases. This review will focus on progress made in the study of an important inherited cardiomyopathy, a disorder which has acted as a paradigm for the study of a genetic cardiovascular disease.
Hypertrophic cardiomyopathy: The clinical disease
The first cardiovascular disorder in which a genetic basis was identified is hypertrophic cardiomyopathy (HCM). Since the modern description of HCM when the disorder was referred to as a “tumour of the heart” 3, much interest has been generated in this clinically diverse cardiac disorder. HCM is a primary disorder of the myocardium characterized by hypertrophy, usually affecting the left ventricle, in the absence of other loading conditions such as hypertension, aortic valve stenosis or thyroid disease4 (Fig 1A). Although previously thought of as a rare disorder, recent population-based studies suggest the prevalence of the condition to be as high as 0.2% (or 1 in 500) in the general population5. The disorder has a wide clinical spectrum, ranging from a benign, asymptomatic course, to symptoms of heart failure6. The most serious complication is sudden cardiac death, with HCM being the commonest cause of sudden cardiac death in individuals aged less than 35 years, including competitive athletes7, 8, 9, 10. Clinical diagnosis is based on unexplained cardiac hypertrophy on echocardiography, which can be subtle or massive, that is classically asymmetric with particular involvement of the interventricular septum, but can also be concentric, diffuse or focal. Asymmetric septal hypertrophy causes a resting or provocative left ventricular outflow tract obstruction in approximately 25% of affected individuals. Although unexplained cardiac hypertrophy is an important pathologic hallmark of disease, altered cardiac morphology is an age-dependent phenotype, often lacking in children aged less than 10 years11, 12. Histopathology features include disorganized myocyte architecture, including disarray of myocyte fibres, intertwined hypertrophied myocytes with bizarre-shaped nuclei, and focal or widespread interstitial fibrosis (Fig 1B).
Pathology of hypertrophic cardiomyopathy A. Post-mortem examination of the heart from an individual ith HCM demonstrating massive asymmetric left ventricular hypertrophy, with associated reduction in left ventricular cavity size, compared to a normal heart. B. Histopathology of heart sections stained with Massons trichrome showing significant myofibre disarray and interstitial fibrosis in HCM.
The clinical management of HCM is complex, in part due to the heterogeneous symptoms exhibited by affected individuals as well as the marked variability in the natural history of this disease. As previously mentioned, HCM can occur without symptoms, but most individuals experience some dypsnoea, angina and palpitations6. The natural history of disease is usually a gradual progression of symptoms, but in some, sudden death or severe heart failure is superimposed. Cardiac death in HCM is often unexpected, occurs throughout life, and even in asymptomatic individuals. Considerable data indicate ventricular arrhythmias significantly contribute to the incidence of sudden death, but clinical tests are often inaccurate in assessing risk. Atrial arrhythmias, embolic events, and congestive heart failure also contribute to the premature morbidity and mortality of this disorder13. Treatment options range from lifestyle changes, e.g. avoiding competitive sports, to drug therapies such as beta-blockers, and interventions such as dual chamber pacemakers, alcohol septal ablation, and implantable cardioverter-defibrillators [for review see ref 14.
Hypertrophic cardiomyopathy: The genetic disease
The first genetic linkage study of HCM was reported in a large French-Canadian family in 198915. Since then, major advances have been made in understanding the molecular basis for HCM. Family studies demonstrate that HCM is a heritable disorder that is transmitted as an autosomal dominant trait, i.e. affected individuals are heterozygous: that is, they have one normal and one mutant copy of the gene. Offspring of affected individuals will therefore have a one in two risk of inheriting the mutation. HCM is a genetically heterogenous disease with at least 10 causative genes now identified. All these genes encode sarcomere proteins, and include the cardiac β-myosin heavy chain, cardiac troponin T gene, α-tropomyosin, myosin-binding protein C, cardiac troponin I, essential and regulatory myosin light chain, and more recently, the cardiac a-myosin heavy chain, titin and actin genes (Tab 1). These genetic data demonstrate that HCM is a disease of the sarcomere.
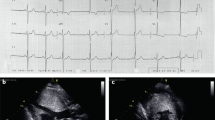
Some of the clinical heterogeneity of HCM can be accounted for by the substantial diversity of gene defects that cause the condition. Approximately 35% of HCM result from cardiac β-myosin heavy chain gene missense mutations. More than 80 unique mutations have been reported that alter one amino acid residue in the globular head or head-rod junction of the b-myosin heavy chain. These mutations are expressed with a high degree of penetrance and most affected individuals (∼90%) exhibit significant myocardial hypertrophy on two-dimensional echocardiography studies by age 20 years16 (Fig 2A). Despite nearly complete disease penetrance and significant hypertrophy, survival in HCM caused by a β-cardiac myosin heavy chain mutation varies considerably and is, in part, mutation-specific. For example, individuals with the Arg403Gln mutation have markedly shortened life expectancies (average age of death = 45 years), whereas survival is near normal in individuals carrying the Val606Met mutation17, 18 (Fig 2B). The change of amino acid charge appears to influence outcome in HCM and conservative mutations appear to be associated with a better prognosis than non-conservative mutations. Although more data is required to provide a complete profile of the phenotypes associated with each myosin mutation, preliminary data clearly indicate that genotype may assist in risk stratification for premature death in HCM.
Genotype-phenotype studies in hypertrophic cardiomyopathy A. Comparison of disease penetrance in families with either β-myosin heavy chain (β-MHC; □) or myosin-binding protein C (MyBP-C; ▪)gene mutations. Penetrance is significantly earlier in families with β-myosin heavy chain gene mutations. B. Comparison of specific gene mutations in HCM. Kaplan-Meier survival curves showing near- normal life expectancy in patients with HCM due to β-MHC mutation Val606Met, or MyBP-C mutation InsG791. In contrast, survival is markedly reduced in HCM caused by β-MHC mutation Arg403Gln or MyBP-C Glu451Gln.
Studies of cardiac myosin-binding protein C, a 137kDa polypeptide with structural and regulatory functions in the sarcomere, have demonstrated mutations that cause HCM19,20. During cardiac embryogenesis, cardiac myosin-binding protein C may participate in the alignment of thick filaments21 whereas in the adult myocardium, phosphorylation of cardiac myosin-binding protein C by a catecholamine-sensitive pathway, may provide dynamic regulation of cardiac contraction in response to adrenergic stimuli22. Whether these or other unknown functions are perturbed by mutations that cause HCM remains unknown. A broad range of gene defects, including missense and splice signal mutations, insertions and deletions, in cardiac myosin-binding protein C can cause HCM19,20,23. As in cardiac troponin T defects, penetranceis often incomplete, and is strikingly age-dependent20. Cardiac myosin-binding protein C mutations may be clinically quiescent until late in adult life, with cardiac hypertrophy manifest only after the fourth and fifth decades of life (Fig 2A). This has implications for clinical screening programs that consider the absence of hypertrophy by age 20 years as indicative of an unaffected individual. Clearly, individuals can develop disease much later in life and thus, need regular clinical screening well into their 40s and 50s. In general the prognosis associated with these mutations is markedly better than that associated with cardiac troponin T defects (detailed below), although the risk of sudden death is increased in at least one cardiac myosin-binding protein C mutation (Fig 2B). It is estimated that approximately 30% of genetically-defined HCM is caused by mutations in cardiac myosin-binding protein C. However the findings of reduced disease penetrance associated with these gene defects, combined with good survival, may have diminished accurate estimates of the true incidence that these mutations have in HCM. Continued definition of cardiac myosin-binding protein C mutations may help to delineate the relationship between gene defects and late-onset HCM.
A variety of cardiac troponin T gene defects (missense mutations, small deletions and mutations in splice signals) cause approximately 15% of all HCM24. The cardiac phenotype produced by these gene defects is characterized by substantially less hypertrophy than that observed with myosin mutations. The mean maximal left ventricular wall thickness resulting from six different cardiac troponin T mutations was 17 ±6 mm (versus 24±8 mm in age-matched patients with β-myosin heavy chain gene mutations), and some adults with these mutations had normal cardiac wall thickness24, 25. Despite this reduced penetrance, most but not all26 reported cardiac troponin T mutations are associated with markedly reduced survival26. Genetic diagnosis may therefore be particularly important in identifying individuals at risk for sudden death in HCM caused by cardiac troponin T defects.
α-tropomyosin gene mutations account for less than 5% of HCM and unlike defects in other sarcomere genes, the spectrum of mutations in α-tropomyosin that cause HCM appears to be limited24. Few disease-causing missense mutations have been identified. The Asp175Asn defect may reflect a mutational hot spot within the gene, as it has arisen independently in multiple families27. Intriguingly, the hypertrophic response to the α-tropomyosin mutation Asp175Asn varies considerably between different families, suggesting that modifying genes and/or environment influence this cardiac phenotype. Despite this, survival is near normal in HCM caused by α-tropomyosin mutation Asp175Asn. Mutations in myosin regulatory or essential light chains rarely cause HCM although the limited number of families reported with these gene defects has hindered correlation of genotype and phenotype28. Distinctive cardiac morphologies have however been reported with some mutations (ventricular regulatory myosin Ala13Thr and Glu22Lys, and myosin essential light chain Met149Val), including predominance of mid-ventricular hypertrophy that resulted in a systolic mid cavity obliteration (hour glass morphology)29. More typical asymmetric hypertrophy also occurs from mutations in these disease genes. Further study is necessary to ascertain to what extent unusual HCM phenotypes are genetically programmed.
Mutations in troponin I have been considered an infrequent cause of HCM although more mutations have recently been reported30, 31, 32, 33. Unusual patterns of hypertrophy, including a predominant apical involvement have also been reported with some troponin I defects30, 31, 32. Based on this morphologic pattern of disease, troponin I mutations may be prevalent in populations with a high incidence of apical HCM, as has been reported in Japan30,34. Furthermore, some mutations have been identified in families which cause both HCM and Wolff-Parkinson-White syndrome30. Mutations in cardiac actin35,36, and titin37are other rare causes of HCM and collectively account for less than 1% of reported cases. Little is known about the clinical correlates of these disease genes.
Genetic data on HCM of the elderly implicate sarcomere protein gene mutations as a cause of this poorly understood disorder. The distribution of mutations that produce hypertrophy late in life is however substantially different from that of familial, early-onset HCM. Whereas defects in β-cardiac myosin heavy chain, cardiac troponin T, and a-tropomyosin account for over 45% of familial HCM, mutations in these were not identified in one study of elderly onset disease. Rather, mutations in cardiac myosin-binding protein C, troponin I, and a-cardiac myosin heavy chain have been defined as causes of elderly-onset HCM38.
A disease locus on chromosome 7q3, previously thought to cause HCM has recently been reclassified as a glycogen storage disease of the heart39, 40, 41. Affected individuals exhibit cardiac hypertrophy as well as electrophysiologic abnormalities including Wolff-Parkinson-White syndrome, atrial fibrillation and progressive atrioventricular block. Cardiac histopathology also distinguishes this disorder from HCM in that myocyte and myofiber disarray is absent, whilst intracytoplasmic vacuoles are found in myocytes41. These vacuoles contain densely packed fine granular and fibrillar electron dense material, with staining properties consistent with polyglucan and amylopectin, a non-soluble product of glycogen metabolism. Mutations in the PRKAG2 gene, encoding the gamma-2 subunit of AMP-activated protein kinase, a molecule involved in energy metabolism and glucose utilization, have been identified in multiple families39, 40, 41. Given these distinctive histopathology, electrophysiologic features and absence the involvement of the mutation in sarcomere function, cardiac hypertrophy resulting from PRKAG2 mutations are more appropriately considered to cause not HCM but a polysaccharide storage disorder of the myocardium.
Molecular pathogenesis of hypertrophic cardiomyopathy
The sarcomere is the basic contractile unit of the heart, and is comprised of both thick and thin filaments. The sarcomere units are aligned in parallel, and are attached to each other through the Z discs. The main component of the thick filament is the myosin heavy chain, while the thin filament is composed of actin, α-tropomyosin and the troponins I, C and T. Myosin-binding protein C and titin are major components of the sarcomere and are involved in both stabilisation of the sarcomere structure, and the generation and transmission of force. Following activation by Ca2+, a series of events involving the troponin-tropomyosin complex results in sliding of the thin and thick filaments, resulting in sarcomere shortening and muscle contraction.
The demonstration that HCM results from defects in genes which encode sarcomeric proteins has raised many questions about the cell biology of cardiac muscle. What is the mechanism by which a cardiac-specific phenotype results from mutations in sarcomere genes? What intracellular signalling pathways are triggered by expression of these gene defects? Is the hypertrophic response compensatory or pathologic? What factors, either genetic or environmental, modify the expression of the mutant gene? Answers to these questions may provide insights into how and why mutated contractile proteins predispose affected individuals to sudden death.
Answering such questions is limited in human studies, due to a variety of factors including broad genetic backgrounds, environmental stimuli which may vary between individuals (e.g. diet, exercise, life-style), small numbers of individuals with the same mutation, and the relative difficulty in obtaining human cardiac samples as well as inadequate methods of maintaining human heart tissue in cell culture systems. For these reasons, a variety of biochemical, cell and animal models have been engineered to more fully dissect the consequences of human sarcomere mutations on muscle structure and function. The development of animal models in HCM have been particularly useful, where there is essentially an unlimited supply of “patients” with the same mutation, where genetic and environmental backgrounds can be controlled, and where access to tissue samples is unlimited.
Cellular studies
One avenue that has advanced our understanding of disease mechanisms has been analyses of HCM mutations in skeletal muscle cells. β-cardiac myosin heavy chains that are abundant in the myocardium, are also found in lower levels in slow twitch skeletal muscles. Abnormal contractile properties of muscle fibers expressing a β-cardiac myosin heavy chain mutation and histologic evidence of central core disease have been demonstrated42,43. The clinical relevance of dysfunction detected ex vivo remains unclear since skeletal myopathy is a rare clinical manifestation of HCM even that caused by a-tropomyosin mutations, which are widely expressed in both skeletal and cardiac muscle. The mechanism by which mutations in this peptide produce a cardiac-specific phenotype may relate to interactions with partner proteins. For example, HCM mutations in α-tropomyosin occur in regions that interact with troponin T, which is expressed as a tissue-specific isoform. Interactions with cardiac troponin T but not skeletal muscle troponin T may be perturbed, thereby restricting dysfunction to the myocardium. Alternatively some α-tropomyosin mutations may alter a unique or critical function of cardiac myocytes. By analogy, essential and regulatory light chain mutations may also produce a predominantly cardiac phenotype by particularly impairing function of cardiac myosin heavy chains.
Structure-function analyses of the location of HCM mutations in sarcomeric proteins suggest that no one common function is perturbed by these mutations. For example, various human β- myosin heavy chain gene mutations have been located on the three-dimensional structure of the chicken skeletal muscle myosin44(Fig 3). These mutations are widely dispersed throughout the molecule, implying that no one function of myosin (such as actin binding, ATP hydrolysis, force transmission or propagation) accounts for disease. The consequences of mutations have also been assayed in reconstituted thick filaments45 and following stable expression into cells43 and results generally indicate that mutant myosins have diminished motor activity without changes in enzymatic activity. Comparable biochemical studies of mutant cardiac troponin T peptides also show adverse effects of hypertrophic mutations on sarcomere filament function, but responses vary depending on calcium concentrations and whether mixtures of mutant and wildtype peptides are present, as would occur in humans who carry heterozygous mutations47, 48. Such data underscore the need to assess the consequences of HCM mutations in the context of cardiac muscle and whole organ physiology.
Location of hypertrophic cardiomyopathy mutations in β-myosin heavy chain Blue spheres represent residues at which mutations which cause HCM have been identified. Green = actin-binding domain; yellow = ATP-binding site; violet = regulatory light chain; orange = essential light chain. Mutations are located throughout the molecule suggesting no single functional abnormality (e.g. ATP binding) is responsible for HCM. (Modified from Rayment et al; ref 44)
Studies in cardiac myocytes have also yielded useful data in understanding disease process in HCM. Expression of mutant troponin T (missense mutation at codon 92) in adult feline cardiac myocytes led to decreased peak, and rate of, cell shortening49. This observation was also seen in rat cardiac myocytes, where overexpression of mutant troponin T resulted in reduced contractile properties 50. Similarly, expression of a truncated troponin T protein in cultured quail myotubes impaired contractile performance51. Furthermore, studies of mutant tropomyosin protein expression in cultured rat myocytes have shown an increased force output at submaximal Ca2+ concentrations52. Taken together, such findings suggest that mutant sarcomeric proteins cause an abnormal power output in isolated cardiac myocytes.
Animal models
While isolated cell models have made important contributions, animal models of HCM have been of greatest value in addressing the issues of molecular pathogenesis, signaling mechanisms, and modifying factors. Genetically engineered mice and rabbits that express human HCM mutations have been particularly useful. Transgenic models that over-express mutant forms of myosin heavy chains53,54, cardiac troponin T55, 56, myosin-binding protein C57, or cardiac troponin I58 as well as a model that physiologically expresses a particular myosin (Arg403Gln) mutation59 have been studied. All these models exhibit histopathology comparable to that observed in human HCM including myocyte disarray, hypertrophy and myocardial fibrosis. The first and most extensively studied mouse model of HCM (Arg403Gln) illustrates how the human disease is replicated in mice (Tab 2). This mouse model was generated by introducing an Arg403Gln mutation into the a-cardiac myosin heavy chain gene by gene targeting and homologous recombination. The mutation is well characterized in humans with HCM, is associated with high penetrance (> 90% express the phenotype by age 20 years) and early sudden death (Fig 2B). The heterozygous mouse genetically recapitulates the human situation, i.e. the mutation is present in one allele only, is under endogenous regulatory mechanisms, and is expressed at physiological levels. Analysis of the phenotype in these heterozygous Arg403Gln mice has revealed features which parallel the human disease (summarised in Tab 2). In brief, Arg403Gln mice develop classical histopathological changes of HCM (myocyte hypertrophy, disarray and fibrosis) by age 15 weeks, and echocardiographically detectable left ventricular hypertrophy by 30 weeks60. This mimics the human disease very closely. Further studies in a second mouse model of HCM in which the myosin-binding protein C gene is truncated using a targeted construct with a Neo cassette, confirms the human finding that disease occurs much later in life, with the myosin-binding protein C mice showing hypertrophy in only 30% of mice at age 30 weeks (versus 100% with hypertrophy in Arg403Gln myosin mice at the same age) but all mice developing disease by age 2 years60. Such comparison between models will be invaluable in addressing the issues of disease heterogeneity and penetrance, as well as the effects of aging in the progression of cardiomyopathy.
Most fundamentally, these animal models of human HCM provide evidence that a mutation in a sarcomeric gene is indeed the primary cause of HCM. These animal models also collectively indicate that cardiac disease is not the consequence of haploinsufficiency, but rather occurs from the dominant effects of mutant proteins on sarcomere function. In transgenic models, levels of mutant sarcomere protein expressed within the heart show some correlation with the severity of myocyte dysfunction57. Analyses of the mechanics of Arg403Gln myosin derived from heart muscle indicate enhanced actin-activated ATPase activity, increased generated force, and accelerated actin filament sliding61. Taken together with data derived from in vitro systems, these findings collectively indicate HCM myosin mutations produce a gain of function. While such defects might actually improve motor performance of individual molecules, in-homogeneity of mutant and wild-type peptides within the sarcomere may uncouple mechanical coordination. This could then increase energy consumption from enhanced ATPase activity and demands of the hypertrophic heart and therefore further diminish any benefit from mutations causing a gain of function.
Longitudinal studies of murine models have also demonstrated that haemodynamic dysfunction precedes histopathology, suggesting that the hypertrophic phenotype occurs as a compensatory response to sarcomere mutation60. Elucidation of intracellular signals that trigger myocyte growth and death are therefore likely to provide novel avenues for intervention and prevention. Such animal models have therefore been invaluable in elucidating various aspects of disease pathogenesis, e.g. related to Ca2+ handling. In the Arg403Gln mouse model of HCM, myofibrillar protein extract, immunohistochemical and ryanodine receptor phosphorylation studies suggest an important early cellular event in HCM is dysregulation of the release of Ca2+ from the sarcoplasmic reticulum, possibly secondary to Ca2+ becoming “trapped” in the mutated sarcomere62. Such studies have led to the identification of pharmacological agents which can both promote severe hypertrophy and sudden death63, as well as prevent disease62, 64, 65. For example, cyclosporin and minoxidil have separately been shown to severely exacerbate cardiac hypertrophy (independent of any blood pressure effects) in Arg403Gln HCM mice, associated with a high incidence of premature death63. The mechanisms by which sarcomere defects increase cardiac interstitial fibrosis have also been probed; L-type Ca2+ channel blockade with diltiazem in HCM mice62; losartan blockade of angiotensin II in HCM mice64, and simvastatin therapy in HCM rabbits65 have shown salutary effects by attenuating or preventing profibrotic effects and reducing collagen deposits. Furthermore, these animal models have provided a basis to systematically study the role of genetic and environmental modifiers in HCM. Based on clinical studies, many individuals within the same family, and therefore carrying the same gene mutation, have vastly different clinical manifestations, strongly supporting the existence of modifying factors (Fig 4A). Animal models of HCM have allowed investigators to begin to evaluate the role of modifiers, primarily due to the unique ability in mice to control both environmental influences, as well as to alter the genetic background by breeding the mutant mice in different mouse strains. Indeed, breeding of the Arg403Gln mice in different genetic backgrounds, both inbred and outbred, has led to the identification of phenotypic differences in terms of hypertrophy (Fig 4B), exercise capacity and histopathology, indicating the presence of a gene modifier in HCM66. Definition of such modifying factors by gene mapping strategies, and other signalling pathways triggered by sarcomere protein gene mutations using animal models has great promise for defining novel targets for therapeutic interventions in human HCM.
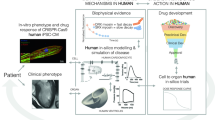
From gene mutation to clinical disease A. Gene mutations cause clinical disease. Two important questions remain the focus of research now and in the future; what is the signalling pathway leading from a gene defect to the clinical phenotype, and secondly, how is this process modified by either genetic and/or environmental factors, e.g. exercise, diet. B. Role of gene modifiers in mice with HCM. Left ventricular wall thickness (LVWT) measurements in 3 groups of mice; wildtype (wt), inbred mice (SvEv) with the Arg403Gln mutation and outbred mice (BSwiss) with the Arg403Gln mutation. An LVWT assessed by echocardiography greater than 1.0 mm (dashed horizontal line) indicates hypertrophy. Approximately 50% of mutant mice in the BSwiss genetic background are protected from developing hypertrophy.
Furthermore, interesting results have arisen by breeding both these heterozygote mouse lines to homozygosity. In both myosin heavy chain and myosin-binding protein C mouse models, homozygosity leads to a dilated cardiomyopathy (DCM), i.e. dilatation of heart chambers with reduced contractile function67, 68. In the Arg403Gln homozygote mice, DCM occurs in neonates and all mice die of heart failure by age 7 days67. In contrast, homozygous myosin-buding protein c mice develop DCM by age 3 weeks, but subsequently develop compensatory hypertrophy and indeed have a normal lifespan68. The ability of being able to breed these mice to both heterozygosity and homozygosity has resulted in clinically relevant models of human HCM and DCM and provide a platform for further studies to both understand pathogenesis, and to potentially identify therapeutic options and targets. The usefulness of such mouse models of human disease is limitless.
The utility of these animal models have been substantially increased by miniaturization of many diagnostic procedures that evaluate cardiac function in humans. For example, the role of vigorous exercise in sudden death and HCM can be evaluated in exercise protocols and provocative electrophysiologic testing in mice. Profound exertion in the Arg403Gln mouse appears to recapitulate sudden death events in some athletes and provocative electrophysiologic testing has demonstrated marked increases in arrhythmogenicity in mutant compared with wild type mice69. In addition, recent studies in a myosin-binding protein C mouse model of HCM show that while these mice may exhibit very mild disease based on cardiac function studies and histopathology analysis, electrophysiological testing suggests that there is a significantly increased vulnerability to ventricular arrhythmias, and therefore sudden death70. M-mode and two-dimensional echocardiography in mice has also enabled accurate assessment of left ventricular hypertrophy, changes in cardiac dimensions, and systolic function (fractional shortening)62, 66, 67, 68. More recently application of magnetic resonance imaging (MRI) has enabled detailed assessment of heart structure and even congenital malformations (e.g. atrial and ventricular septal defects) in mice71. Murine models of human cardiac disease therefore appear to be valuable reagents for delineating the mechanisms of disease, analyses of complex events such as sudden death and important tools for evaluating pharmacologic therapies and devices.
Future perspectives
Understanding how sarcomere mutations perturb biophysical events of muscle contraction and cell signalling pathways within the myocyte, will inevitably allow advances in future treatment interventions. With increased understanding of the genetic mechanisms, it may be possible to target therapy to mitigate the genetic defect or conceivably, to correct the molecular abnormality, i.e. to actually correct the single base abnormality in the mutant allele, and effectively cure the disease. Alternatively, the dominant negative mechanism by which these gene mutations act impacts on the potential for somatic gene therapy for HCM which theoretically would have to be directed toward specific inactivation of the mutant allele.
An exponential growth in our knowledge of gene defects that cause inherited cardiomyopathies, and in particular HCM, will undoubtedly continue72. Molecular genetic causes provide a solid basis for both classifying and understanding many cardiac disorders previously classified as “idiopathic”. Integration of molecular insights into clinical practice has potential advantages to patients afflicted by these intriguing disorders and to the physicians who care for them. Identification of human mutations allows early and accurate diagnosis, enabling modification of specific lifestyle factors (i.e., avoidance of competitive athletics), as well as early therapeutics that can reduce the serious morbidity and mortality associated with these disorders (e.g. implantation of cardioverter-defibrillators). Continued study and development of animal models of disease should further enable studies of the integrative physiology of multiple organ systems involved in the development of cardiomyopathies. Elucidation of signalling events leading from mutant protein to clinical phenotype, and the identification of factors, both genetic and environmental that modify the response to mutations, may provide new insights that can enhance our fundamental understanding of the pathogenesis of HCM and other inherited cardiomyopathies and therefore improve therapeutic intervention.
References
Watson J, Crick F . Molecular structure of nucleic acids. Nature 1953; 171:737–8.
Pennisi E . Finally, the Book of Life and instructions for navigating it. Science 2000; 288:2304–7.
Teare D . Asymmetrical hypertrophy of the heart in young adults. Br Heart J 1958; 20:1–8.
Seidman CE, Seidman JG . Molecular genetic studies of familial hypertrophic cardiomyopathy. Basic Res Cardiol 1998; 93:13–6.
Maron BJ, Gardin JM, Flack JM, Gidding SS, Bild D . Prevalence of hypertrophic cardiomyopathy in a general population of young adults: echocardiographic analysis of 4111 subjects in the CARDIA Study. Circulation 1995; 92:785–9.
Spirito P, Seidman CE, McKenna WJ, Maron BJ . The management of hypertrophic cardiomyopathy. N Engl J Med 1997; 336:775–85.
Maron BJ, Epstein SE, Roberts WC . Causes of sudden death in competitive athletes. J Am Coll Cardiol 1986; 7: 204–14.
Maron BJ, Shirani J, Poliac LC, Mathenge R, Roberts WC, Mueller FO . Sudden death in young competitive athletes-clinical, demographic and pathological profiles. JAMA 1996; 276:199–204.
Semsarian C, Richmond DR . Sudden cardiac death in familial hypertrophic cardiomyopathy: an Australian experience. Aust NZ J Med 1999; 29:368–70.
Semsarian C, Maron BJ . Sudden cardiac death in the young. Med. J. Aust 2002; 176:148–9.
Maron BJ, Spirito P, Wesley Y, et al. Development and progression of left ventricular hypertrophy in children with hypertrophic cardiomyopathy. N Engl J Med 1986; 315:610.
Semsarian C, Jeremy RW, French J, Trent R, Richmond DR . Natural history of wall thickening in hypertrophic cardiomyopathy. Aust NZ J Med 1997; 27:51–8.
Maron BJ, Olivotto I, Spirito P, et al. Epidemiology of hypertrophic cardiomyopathy -related death: revisited in a large non-referral-based patient population. Circulation 2000; 102:858–64.
Maron BJ . Hypertrophic cardiomyopathy: a systematic review. JAMA 2002; 287:1308–20.
Jarcho JA, McKenna WJ, Pare JAP, et al. Mapping a gene for familial hypertrophic cardiomyopathy to chromosome 14q1. N Engl J Med 1989; 321:1372.
Anan R, Greve G, Thierfelder L, et al. Prognostic implications of novel b-cardiac myosin heavy chain gene mutations that cause familial hypertrophic cardiomyopathy. J Clin Invest 1994; 93:280–5.
Watkins H, Rosenzweig A, Hwang D, et al. Characteristics and prognostic implications of myosin missense mutations in familial hypertrophic cardiomyopathy. N Engl J Med 1992; 326:1108–14.
Fananapazir L, Epstein ND . Genotype-phenotype correlations in hypertrophic cardiomyopathy. Circulation 1994; 89:22–9.
Watkins H, Conner D, Thierfelder L, et al. Mutations in the cardiac myosin binding protein-C gene on chromosome 11 cause familial hypertrophic cardiomyopathy. Nat Genet 1995; 11: 434–7.
Nimura H, Bachinski LL, Sangwatanaroj S, et al. Mutations in the gene for cardiac myosin-binding protein C and late-onset familial hypertrophic cardiomyopathy. N Engl J Med 1998; 338:1248–57.
Gilbert R, Kelly MG, Mikawa T, et al. The carboxyl terminus of myosin binding protein C (MyBP-C, C-protein) specifies incorporation into the A-band of striated muscle. J Cell Science 1996; 109:101–11.
Gautel M, Zuffardi O, Freiburg A, et al. Phosphorylation switches specific for the cardiac isoform of myosin binding protein-C: a modulator of cardiac contraction? EMBO J 1995; 14:1952–60.
Yu B, French J, Carrier L, Jeremy RW, et al. Molecular pathology of familial hypertrophic cardiomyopathy caused by mutations in the cardiac myosin binding protein C gene. J Med Genet 1998; 35:205–10.
Watkins H, McKenna WJ, Thierfelder L, et al. The role of cardiac troponin T and ??tropomyosin mutations in hypertrophic cardiomyopathy. N Engl J Med 1995; 332:1058.
Varnava AM, Elliott PM, Baboonian C, Davison F, Davies MJ, McKenna WJ . Hypertrophic cardiomyopathy: histopathological features of sudden death in cardiac troponin T disease. Circulation. 2001; 104:1380–4.
Anan R, Shono H, Kisanuki A, Arima S, Nakao S, Tanaka H . Patients with familial hypertrophic cardiomyopathy caused by a Phe110Ile missense mutation in the cardiac troponin T gene have variable cardiac morphologies and a favourable prognosis. Circulation 1998; 98:391–7.
Coviello DA, Maron BJ, Spirito P, et al. Clinical features of hypertrophic cardiomyopathy caused by mutation of a “hot spot” in the tropomyosin gene. J Am Coll Cardiol 1996; 29:635–40.
Poetter K, Jiang H, Hassanzadeh S, et al. Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nat Genet 1996; 13:63–9.
Lee W-H, Hwang TH, Kimura A, et al. Different expressivity of a ventricular essential myosin light chain gene Ala57Gly mutation in familial hypertrophic cardiomyopathy. Am Heart J 2001; 141:184–9.
Kimura A, Harada H, Park J-E et al. Mutations in the cardiac troponin I gene associated with hypertrophic cardiomyopathy. Nat Genet 1997; 16:379–82.
Kokado H, Shimizu, M, Hirouki Y, et al. Clinical features of hypertrophic cardiomyopathy caused by a Lys183 deletion mutation in the cardiac troponin I gene. Circulation 2000; 102:663–9.
Woo A, Rakowski H, Liew J, et al. Hypertrophic cardiomyopathy: genotypic and phenotypic heterogeneity. Circulation 2000; 102:169–78.
Pascale R, Charron P, Carrier L, et al. Distribution of disease gene in 102 genotyped families with hypertrophic cardiomyopathy. Circulation 2001; 104: II521.
Yamaguchi H, Ishimura T, Nishiyama S, et al. Hypertrophic nonobstructive cardiomyopathy with giant negative T waves (Apical hypertrophy):ventriculographic echocardiographic features in 30 patients. Am J Cardiol 1979; 44:401–12.
Mogensen J, Klausen IC, Pedersen AK, et al. a-cardiac actin is a novel gene in familial hypertrophic cardiomyopathy. J Clin Invest 1999; 103:39R–43R.
Oson TM, Doan TP, Kishimoto NY, et al. Inherited and de novo mutations in the cardiac actin gene cause hypertrophic cardiomyopathy. J Mol Cell Cardiol 2000; 32:1687–94.
Satoh M, Takahashi M, Sakamoto T, Hiroe M, Marumo F, Kimura A: Structural analysis of the titin gene in hypertrophic cardiomyopathy: identification of a novel disease gene. Biochem Biophys Res Commun 1999; 262:411–7.
Niimura H, Patton K, McKenna, W, et al. Sarcomere protein gene mutations in hypertrophic cardiomyopathy of the elderly. Circulation 2002; 105:446–51.
Gollob MH, Green MS, Tang AS et al. Identification of a gene responsible for Wolff- Parkinson-White syndrome. N Engl J Med 2001; 344:1823–31.
Blair E, Redwood C, Ashrafian H, et al. Mutations in the gamma(2) subunit of AMP- activated protein kinase cause familial hypertrophic cardiomyopathy: evidence for the central role of energy compromise in disease pathogenesis. Hum Molec Genet 2001; 10:1215–20.
Arad M, Benson W, Perez-Atayde AR et al. Constitutively active AMP kinase mutations cause glycogen storage disease mimicking hypertrophic cardiomyopathy. J Clin Invest 2002; 109:357–62.
Lankdord EB, Epstein ND, Fananapazir L, et al. Abnormal contractile properties of muscle fibers expression b-myosin chain mutations in patients with hypertrophic cardiomyopathy. J Clin Invest 1995; 95:1409–14.
Fananapazir L, Dalakis MC, Cyran F, et al. Missense mutations in the β- myosin heavy chain cause central core disease in hypertrophic cardiomyopathy. Proc Natl Acad Sci USA 1993; 90:3993–7.
Rayment I, Holden HM, Sellers JR, et al. Structural interpretation of the mutations in the b-cardiac myosin that have been implicated in familial hypertrophic cardiomyopathy. Proc Natl Acad Sci USA 1995; 92:3868–94.
Sweeney HL, Straceski AJ, Leinwand LA, et al. Heterologous expression of a cardiomyopathic myosin that is defective in its actin interaction. J Biol Chem 1994; 269:1603–5.
Sata M, Ikebe M: Functional analysis of the mutations in the human cardiac?- myosin that are responsible for familial hypertrophic cardiomyopathy. J Clin Invest 1996; 98:2866–73.
Lin D, Bobkova A, Homsher E, et al. Altered cardiac troponin T in vitro function in the presence of a mutation implicated in familial hypertrophic cardiomyopathy. J Clin Invest 1996; 97:2842–8.
Watkins H, Seidman CE, Seidman JG, et al. Expression and functional assessment of a truncated cardiac troponin T that cause hypertrophic cardiomyopathy: evidence for a dominant negative action. J Clin Invest 1996; 11:2456–61.
Marian AJ, Zhao G, Seta Y, Roberts R, Yu QT . Expression of mutant (Arg92Gln) human cardiac troponin T, known to cause hypertrophic cardiomyopathy, impairs adult cardiac myocyte contractility. Circ Res 1997; 81:76–85.
Rust EM, Albayya FP, Metzger JM . Identification of a contractile deficit in adult cardiac myocytes expressing hypertrophic cardiomyopathy-associated mutant troponin T proteins. J Clin Invest 1999; 103:1459–67.
Watkins H, Seidman CE, Seidman JG, Feng HS, Sweeney HL . Expression and functional assessment of a truncated cardiac troponin T that causes hypertrophic cardiomyopathy: evidence for a dominant negative action. J Clin Invest 1996; 98:2456–61.
Michele DE, Albayya FP, Metzger JM . Direct, convergent hypersensitivity of calcium-activated force generation produced by hypertrophic cardiomyopathy mutant a-tropomyosins in adult cardiac myocytes. Nat Med 1999; 5:1413–7.
Vikstrom KL, Factor SM, Leinwand LA . A murine model of hypertrophic cardiomyopathy. Zeitschrift fur Kardiologie 1995; 84:49–54.
Marian AJ, Wu Y, McCluggage M, et al. A transgenic rabbit model for human hypertrophic cardiomyopathy. J Clin Invest 1999; 104:1683–92.
Oberst L, Zhao G, Park JT, et al. Dominant-negative effect of a mutant cardiac troponin T on cardiac structure and function in transgenic mice. J Clin Invest 1998; 102:1498–505.
Tardiff JC, Factor SM, Tompkins BD, et al. A truncated cardiac troponin T molecule in transgenic mice suggests multiple cellular mechanisms for familial hypertrophic cardiomyopathy. J Clin Invest 1998; 101:2800–11.
Yang Q, Sanbe A, Osinka H, et al. A mouse model of myosin binding protein C human familial hypertrophic cardiomyopathy. J Clin Invest 1998; 102:1292–300.
James J, Zhang Y, Osinska H, et al. Transgenic modeling of a cardiac troponin I mutation linked to familial hypertrophic cardiomyopathy. Circ Res 2000; 87:805–11.
Geisterfer-Lowrance AAT, Christe M, Conner DA, et al. A murine model of familial hypertrophic cardiomyopathy. Science 1996; 272:731–4.
McConnell BK, Fatkin D, Semsarian C, et al. Comparison of two murine models of familial hypertrophic cardiomyopathy. Circ Res 2001; 88:383–9.
Tyska MJ, Hayes E, Giewat M, et al. Single molecule mechanics of R403Q cardiac myosin isolated from the mouse model of familial hypertrophic cardiomyopathy. Circ Res 2000; 86:737–44.
Semsarian C, Ahmad I, Giewat M, et al. The L-type calcium-channel inhibitor diltiazem prevents cardiomyopathy in a mouse model. J Clin Invest 2002; 109:1013–20.
Fatkin D, McConnell BK, Mudd JO, et al. An abnormal Ca2+ response in mutant sarcomere protein-mediated familial hypertrophic cardiomyopathy. J Clin Invest 2000; 106:1351–9.
Lim DS, Lutucuta S, Bachireddy P, et al. Angiotensin II blockade reverses myocardial fibrosis in a transgenic mouse model of human hypertrophic cardiomyopathy. Circulation 2001; 103:789–91.
Patel R, Nagueh SF, Tsybouleva N, et al. Simvastatin induces regression of cardiac hypertrophy and fibrosis and improves cardiac function in a transgenic rabbit model of human hypertrophic cardiomyopathy. Circulation 2001; 104:317–24.
Semsarian C, Fatkin D, Healey MJ, et al. A polymorphic modifier gene alters the hypertrophic response in a murine model of familial hypertrophic cardiomyopathy. J Mol Cell Cardiol 2001; 33:2055–60.
Fatkin D, Christe ME, Aristizabal O, et al. Neonatal cardiomyopathy in mice homozygous for the Arg403Gln mutation in the alpha cardiac myosin heavy chain gene. J Clin Invest.; 103:147–53.
McConnell BK, Jones KA, Fatkin D, et al. Dilated cardiomyopathy in homozygous myosin-binding protein-C mutant mice. J Clin Invest 1999; 104:1235–44.
Berul C, Christie M, Aronovitz MJ et al. Electrophysiological abnormalities and arrhythmias in ? MHC mutant familial hypertrophic cardiomyopathy mice. J Clin Invest 1997; 99:570–6.
Berul CI, McConnell BK, Wakimoto H, et al. Ventricular arrhythmia vulnerability in cardiomyopathic mice with homozygous mutant myosin-binding protein C gene. Circulation 2001; 104:2734–9.
Bruneau, BG, Nemer G, Schmitt JP, et al. Murine model of Holt-Oram syndrome defines roles of the T-box transcription facto Tbx5 in cardiogenesis and disease. Cell 2001; 106:709–22.
Semsarian C, Seidman CE . Molecular medicine in the 21st century. Intern Med J 2001; 31:53–9.
Acknowledgements
CS is the recipient of a National Heart Foundation of Australia Fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
CHUNG, MW., TSOUTSMAN, T. & SEMSARIAN, C. Hypertrophic cardiomyopathy: from gene defect to clinical disease. Cell Res 13, 9–20 (2003). https://doi.org/10.1038/sj.cr.7290146
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7290146
Keywords
This article is cited by
-
Genetic, clinical, molecular, and pathogenic aspects of the South Asian–specific polymorphic MYBPC3Δ25bp variant
Biophysical Reviews (2020)
-
Multi-targeting chemoprevention of Chinese herb formula Yanghe Huayan decoction on experimentally induced mammary tumorigenesis
BMC Complementary and Alternative Medicine (2019)
-
Transient active force generation and stress fibre remodelling in cells under cyclic loading
Biomechanics and Modeling in Mechanobiology (2019)
-
Tumor Necrosis Factor-α in Heart Failure: an Updated Review
Current Cardiology Reports (2018)
-
Precision medicine approach to genetic cardiomyopathy
Herz (2017)