Featured
-
-
Article
| Open Access Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria
Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria
Transcription rates are regulated by the interactions between RNA polymerase, sigma factor, and promoter DNA sequences in bacteria. Here the authors combine massively parallel experiments & machine learning to develop a predictive biophysical model of transcription, validated across 22132 bacterial promoters, and apply it to the design and debugging of genetic circuits.
- Travis L. LaFleur
- , Ayaan Hossain
- & Howard M. Salis
-
Article
| Open Access Specificity of the Hox member Deformed is determined by transcription factor levels and binding site affinities
Specificity of the Hox member Deformed is determined by transcription factor levels and binding site affinities
Despite the central role of Hox genes in controlling morphogenesis, the DNA binding of different Hox members is relatively similar. Here they show that specificity of Hox member Dfd relies on a precise balance of transcription factors and binding site affinities.
- Pedro B. Pinto
- , Katrin Domsch
- & Ingrid Lohmann
-
Article
| Open Access Structural analysis of Red1 as a conserved scaffold of the RNA-targeting MTREC/PAXT complex
Structural analysis of Red1 as a conserved scaffold of the RNA-targeting MTREC/PAXT complex
Unwanted RNA transcripts are targeted for degradation by nuclear complexes such as MTREC/PAXT. Here, the authors structurally and functionally characterized three interfaces of the scaffold protein Red1, providing mechanistic insights into conserved features of MTREC/PAXT architecture.
- Anne-Emmanuelle Foucher
- , Leila Touat-Todeschini
- & Jan Kadlec
-
Article
| Open Access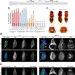 A single WNT enhancer drives specification and regeneration of the Drosophila wing
A single WNT enhancer drives specification and regeneration of the Drosophila wing
The wing is a remarkable evolutionary novelty in insects. Here the authors demonstrate that the specification and regenerative capacity of the wing relies on a single wing-specific enhancer of the wingless gene in Drosophila.
- Elena Gracia-Latorre
- , Lidia Pérez
- & Marco Milán
-
Article
| Open Access Visualizing molecular interactions that determine assembly of a bullet-shaped vesicular stomatitis virus particle
Visualizing molecular interactions that determine assembly of a bullet-shaped vesicular stomatitis virus particle
Vesicular stomatitis virus (VSV) is the most widely studied prototype for negative-sense RNA viruses. Structure determination of VSV particles is particularly challenging because they are polymorphic with different helical symmetries. Here, Jenni et al. apply computational classification approaches to sort different morphologies, and obtain CryoEM reconstructions at 3.5–4.1 A resolution. They show that the matrix protein (M) is present in the virion in two layers, of which the inner layer associates with N protein of vRNPs.
- Simon Jenni
- , Joshua A. Horwitz
- & Stephen C. Harrison
-
Article
| Open Access Structural basis of transcriptional regulation by a nascent RNA element, HK022 putRNA
Structural basis of transcriptional regulation by a nascent RNA element, HK022 putRNA
HK022 put is an RNA element that inhibits transcription termination without aids from protein factors. Here, authors solved cryo-EM structures of put-associated RNA polymerase and showed the structure of putRNA and its binding to the RNA polymerase.
- Seungha Hwang
- , Paul Dominic B. Olinares
- & Jin Young Kang
-
Article
| Open Access Mettl3-mediated mRNA m6A modification controls postnatal liver development by modulating the transcription factor Hnf4a
Mettl3-mediated mRNA m6A modification controls postnatal liver development by modulating the transcription factor Hnf4a
m6A is the most abundant RNA modification of eukaryotic mRNAs and is involved in various physiological and pathological processes. Here the authors show a role for Mettl3-mediated RNA m6A modification in postnatal liver development by regulating the Hnf4a-centered transcriptional network
- Yan Xu
- , Zhuowei Zhou
- & Qi Zhang
-
Article
| Open Access Etv2 regulates enhancer chromatin status to initiate Shh expression in the limb bud
Etv2 regulates enhancer chromatin status to initiate Shh expression in the limb bud
The embryonic limb bud is known to be patterned by a Shh morphogen gradient, though how Shh expression is activated remains less clear. Here the authors show that Etv2 acts as a pioneer transcription factor to mediate accessibility of the ZRS enhancer and initiate Shh expression.
- Naoko Koyano-Nakagawa
- , Wuming Gong
- & Daniel J. Garry
-
Article
| Open Access Pseudomonas aeruginosa SutA wedges RNAP lobe domain open to facilitate promoter DNA unwinding
Pseudomonas aeruginosa SutA wedges RNAP lobe domain open to facilitate promoter DNA unwinding
SutA is a transcription factor which increases transcription activity of an RNA polymerase (RNAP). Here, authors present cryo-EM structures of SutA-bound RNAP-σS holoenzyme and SutA-bound transcription initiation complex, which reveals SutA wedging the RNAP-β lobe open to aid unwinding.
- Dingwei He
- , Linlin You
- & Yu Zhang
-
Article
| Open Access Inactivation of Sirt6 ameliorates muscular dystrophy in mdx mice by releasing suppression of utrophin expression
Inactivation of Sirt6 ameliorates muscular dystrophy in mdx mice by releasing suppression of utrophin expression
Utrophin is a dystrophin-related protein stabilizing the sarcolemma in absence of dystrophin. Here the authors report that inactivation of the protein deacetylase SIRT6, involved in the deacetylation of the epigenetic mark H3K56ac in muscle cells, increases expression of utrophin and ameliorates dystrophic muscle pathology in mice.
- Angelina M. Georgieva
- , Xinyue Guo
- & Thomas Braun
-
Article
| Open Access Molecular dissection of the glutamine synthetase-GlnR nitrogen regulatory circuitry in Gram-positive bacteria
Molecular dissection of the glutamine synthetase-GlnR nitrogen regulatory circuitry in Gram-positive bacteria
How bacteria sense and respond to nitrogen is a key question in microbial physiology. This work unveils the mechanism by which the central nitrogen metabolic enzyme, glutamine synthetase, directly signals nitrogen availability to the GlnR regulator.
- Brady A. Travis
- , Jared V. Peck
- & Maria A. Schumacher
-
Article
| Open Access Lentivector cryptic splicing mediates increase in CD34+ clones expressing truncated HMGA2 in human X-linked severe combined immunodeficiency
Lentivector cryptic splicing mediates increase in CD34+ clones expressing truncated HMGA2 in human X-linked severe combined immunodeficiency
De Ravin et al. report an unplanned interim analysis of a secondary safety outcome for an ongoing clinical trial on lentiviral gene therapy for the treatment of X-linked Severe Combined Immunodeficiency (NCT01306019). Vector induced alternative splicing events are identified that cause aberrant fusion transcripts, leading to clonal dominance in a single patient and clonal expansion in others. This can be mitigated by the removal of the lentivector cryptic splice acceptor.
- Suk See De Ravin
- , Siyuan Liu
- & Xiaolin Wu
-
Article
| Open Access RIViT-seq enables systematic identification of regulons of transcriptional machineries
RIViT-seq enables systematic identification of regulons of transcriptional machineries
Here the authors present their method ‘regulon identification by in vitro transcription-sequencing’ (RIViT-seq), which enables systematic identification of target genes of transcription factors of interest. They applied RIViT-seq to 13 sigma factors from Streptomyces coelicolor A3(2) and successfully identified target genes of 11 of these, expanding the regulatory characterisation in this organism.
- Hiroshi Otani
- & Nigel J. Mouncey
-
Article
| Open Access JMJD3 intrinsically disordered region links the 3D-genome structure to TGFβ-dependent transcription activation
JMJD3 intrinsically disordered region links the 3D-genome structure to TGFβ-dependent transcription activation
Here the authors demonstrate that TGFβ drives multi-enhancer contacts and ultimately gene activation during neuronal commitment, and that this requires the intrinsically disordered region (IDR) of the histone demethylase JMJD3 likely through its role in promoting phase-separated biomolecular condensates.
- Marta Vicioso-Mantis
- , Raquel Fueyo
- & Marian A. Martínez-Balbás
-
Article
| Open Access Extended intergenic DNA contributes to neuron-specific expression of neighboring genes in the mammalian nervous system
Extended intergenic DNA contributes to neuron-specific expression of neighboring genes in the mammalian nervous system
A large part of noncoding DNA is intergenic regions, yet how the size of intergenic regions affects gene expression in a tissue-specific manner is unclear. Here the authors present long intergenic DNA length-dependent neural gene expression, reflecting the complexity in the mammalian nervous system.
- Ravneet Jaura
- , Ssu-Yu Yeh
- & Ho Sung Rhee
-
Article
| Open Access Insulin action and resistance are dependent on a GSK3β-FBXW7-ERRα transcriptional axis
Insulin action and resistance are dependent on a GSK3β-FBXW7-ERRα transcriptional axis
The downstream mechanisms involved in insulin signaling and resistance remain incompletely understood. Here the authors report that insulin-dependent dephosphorylation stabilizes ERRα via the GSK3β/FBXW7 axis, and disruption of this post-translational mechanism results in insulin resistance in mice.
- Hui Xia
- , Charlotte Scholtes
- & Vincent Giguère
-
Article
| Open Access Cooperative interaction between ERα and the EMT-inducer ZEB1 reprograms breast cancer cells for bone metastasis
Cooperative interaction between ERα and the EMT-inducer ZEB1 reprograms breast cancer cells for bone metastasis
The epithelial mesenchymal transition (EMT) is important in the metastatic spread of cancer cells. Here, the authors show that the EMT transcription factor, ZEB1, can modify estrogen receptor α during EMT and facilitate the migration of breast cancer cells to the bone
- Nastaran Mohammadi Ghahhari
- , Magdalena K. Sznurkowska
- & Didier Picard
-
Article
| Open Access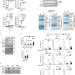 The nuclear receptor ERR cooperates with the cardiogenic factor GATA4 to orchestrate cardiomyocyte maturation
The nuclear receptor ERR cooperates with the cardiogenic factor GATA4 to orchestrate cardiomyocyte maturation
Mature cardiac muscle requires high mitochondrial ATP production and specialized contractile proteins. Here the authors demonstrate that cardiomyocyte-specific contractile maturation involves cooperation between the nuclear receptor ERRγ and cardiogenic transcription factor GATA4, but ERRγ controls metabolic genes independently.
- Tomoya Sakamoto
- , Kirill Batmanov
- & Daniel P. Kelly
-
Article
| Open Access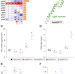 Exploring a blue-light-sensing transcription factor to double the peak productivity of oil in Nannochloropsis oceanica
Exploring a blue-light-sensing transcription factor to double the peak productivity of oil in Nannochloropsis oceanica
Microalgae are promising feedstock for oil production. The authors report that a transcription factor NobZIP77 can regulate oil synthesis by sensing the blue light, and explore these findings to greatly enhance oil productivity via genetic and process engineering in Nannochloropsis oceanica.
- Peng Zhang
- , Yi Xin
- & Jian Xu
-
Article
| Open Access Unveiling RCOR1 as a rheostat at transcriptionally permissive chromatin
Unveiling RCOR1 as a rheostat at transcriptionally permissive chromatin
The classical neuronal-gene corepressor RCOR1/CoREST is paradoxically enriched in transcriptionally active chromatin. Here the authors show RCOR1 is recruited during promoter-proximal pausing and negatively regulates the nascent-transcript synthesis. They also show that an RCOR1-LSD1- HDAC1 complex removes lysine acetylation from RNA polymerase II to repress transcription.
- Carlos Rivera
- , Hun-Goo Lee
- & María Estela Andrés
-
Article
| Open Access Transcription factors modulate RNA polymerase conformational equilibrium
Transcription factors modulate RNA polymerase conformational equilibrium
Pausing of RNA polymerase (RNAP) and transcription is regulated by the NusA and NusG transcription factors in bacteria. Here the authors provide structural evidence for how they interact with RNAP to carry out their pausing roles and also reveal functions for NusA and NusG in transcription termination.
- Chengjin Zhu
- , Xieyang Guo
- & Albert Weixlbaumer
-
Article
| Open Access GLI transcriptional repression is inert prior to Hedgehog pathway activation
GLI transcriptional repression is inert prior to Hedgehog pathway activation
GLI repression has been presumed to be the default transcriptional state and important for pre-patterning tissues. Challenging current models, the authors show that GLI3 repression is inert in the limb bud before the onset of Hedgehog signaling.
- Rachel K. Lex
- , Weiqiang Zhou
- & Steven A. Vokes
-
Article
| Open Access Manipulation of RNA polymerase III by Herpes Simplex Virus-1
Manipulation of RNA polymerase III by Herpes Simplex Virus-1
RNA Polymerase III (Pol III) transcribes non-coding RNA, including tRNAs. Applying different RNA-Seq techniques, Dremel et al. provide the Pol III transcriptional landscape of Herpes simplex virus 1 (HSV-1) infected cells. Infection leads to an increase in tRNA expression from host euchromatin and Pol II re-localization to tRNA loci. They also find that Pol III – associated factors bind to the viral genome.
- Sarah E. Dremel
- , Frances L. Sivrich
- & Neal A. DeLuca
-
Article
| Open Access Androgen receptor and MYC equilibration centralizes on developmental super-enhancer
Androgen receptor and MYC equilibration centralizes on developmental super-enhancer
Androgen receptor in prostate cancer (PCa) transcriptionally represses multiple genes including MYC. Here, the authors suggest that increased MYC in response to androgen deprivation contributes to castration-resistant PCa, while decreased MYC may contribute to responses to supraphysiological androgen therapy.
- Haiyang Guo
- , Yiming Wu
- & Steven P. Balk
-
Article
| Open Access Structural insights into RNA polymerase III-mediated transcription termination through trapping poly-deoxythymidine
Structural insights into RNA polymerase III-mediated transcription termination through trapping poly-deoxythymidine
Termination of eukaryotic RNA polymerase III (Pol III)-mediated transcription occurs when the polymerase reaches a stretch of four or more deoxythymidine nucleotides (poly-dT) on the non-template strand. Here, the authors present the 3.6 Å cryo-EM structure of a human Pol III pre-termination complex (PTC) that was assembled on a 7 dT-containing DNA template and discuss the mechanism of poly-dT-dependent transcription termination of Pol III.
- Haifeng Hou
- , Yan Li
- & Yanhui Xu
-
Article
| Open Access Complex small-world regulatory networks emerge from the 3D organisation of the human genome
Complex small-world regulatory networks emerge from the 3D organisation of the human genome
Gene-regulatory networks are thought to be complex, and yet perturbation of just a few transcription factors (TFs) can have major consequences. Here the authors apply DNA polymer modelling and simulations to predict how 3D genome structure and TF-DNA interactions can give rise to transcriptional regulation operating over broad genomic regions, where small perturbations can have long-reaching effects.
- C. A. Brackley
- , N. Gilbert
- & D. Marenduzzo
-
Article
| Open Access Functional characterization of T2D-associated SNP effects on baseline and ER stress-responsive β cell transcriptional activation
Functional characterization of T2D-associated SNP effects on baseline and ER stress-responsive β cell transcriptional activation
Identifying causal variants at GWAS loci is important to understand disease mechanisms. Here the authors use massively parallel reporter assays to identify type 2 diabetes-associated variants that alter cis-regulatory activity, narrowing in on the causal variants and genetic mechanisms behind the disease.
- Shubham Khetan
- , Susan Kales
- & Michael L. Stitzel
-
Article
| Open Access A TALE/HOX code unlocks WNT signalling response towards paraxial mesoderm
A TALE/HOX code unlocks WNT signalling response towards paraxial mesoderm
Cells in the developing embryo interpret WNT signalling with context-dependence, but the mechanism decoding these cues is unclear. Here, the authors show that combinatorial TALE/HOX activity destabilizes nucleosomes at WNT-responsive regions to activate paraxial mesodermal genes.
- Luca Mariani
- , Xiaogang Guo
- & Elisabetta Ferretti
-
Article
| Open Access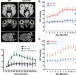 GLIS1 regulates trabecular meshwork function and intraocular pressure and is associated with glaucoma in humans
GLIS1 regulates trabecular meshwork function and intraocular pressure and is associated with glaucoma in humans
Dysfunction of the trabecular meshwork (TM) is the chief cause of elevated intraocular pressure, the major risk factor of glaucoma. Here, the authors identify the transcription factor, GLIS1, as a critical regulator of TM maintenance and intraocular pressure, and as a glaucoma risk gene.
- K. Saidas Nair
- , Chitrangda Srivastava
- & Anton M. Jetten
-
Article
| Open Access Quantitative imaging of transcription in living Drosophila embryos reveals the impact of core promoter motifs on promoter state dynamics
Quantitative imaging of transcription in living Drosophila embryos reveals the impact of core promoter motifs on promoter state dynamics
Here the authors decode how core promoter elements regulate rate limiting steps of transcription using quantitative live imaging, genetics and modeling in early Drosophila embryos. TATA-driven promoters require one rate-limiting step while INR promoters need an extra step associated with Pol II pausing.
- Virginia L. Pimmett
- , Matthieu Dejean
- & Mounia Lagha
-
Article
| Open Access Multiplexed functional genomic analysis of 5’ untranslated region mutations across the spectrum of prostate cancer
Multiplexed functional genomic analysis of 5’ untranslated region mutations across the spectrum of prostate cancer
Mutations in 5’ untranslated regions (UTRs) have a functional role in gene expression in cancer. Here, the authors develop a sequencing-based high throughput functional assay named PLUMAGE and show the effects of these mutations on gene expression and their association with clinical outcomes in prostate cancer.
- Yiting Lim
- , Sonali Arora
- & Andrew C. Hsieh
-
Article
| Open Access Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macrophages
Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macrophages
STAT1a is required for pro-inflammatory responses in macrophages. Here the authors reveal that post-translational modification of STAT1a, ADPribosylation, plays a critical role in enhancer formation and activation, thus modulating IFNγ-stimulated inflammatory responses in macrophages.
- Rebecca Gupte
- , Tulip Nandu
- & W. Lee Kraus
-
Article
| Open Access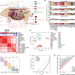 A pig BodyMap transcriptome reveals diverse tissue physiologies and evolutionary dynamics of transcription
A pig BodyMap transcriptome reveals diverse tissue physiologies and evolutionary dynamics of transcription
A comprehensive transcriptomic survey of the pig could enable mechanistic understanding of tissue specialization and accelerate its use as a biomedical model. Here the authors characterize four distinct transcript types in 31 adult pig tissues to dissect their distinct structural and transcriptional features and uncover transcriptomic variability related to tissue physiology.
- Long Jin
- , Qianzi Tang
- & Mingzhou Li
-
Article
| Open Access Acetylation of PAX7 controls muscle stem cell self-renewal and differentiation potential in mice
Acetylation of PAX7 controls muscle stem cell self-renewal and differentiation potential in mice
The acetyltransferase MYST1 stimulated by acetyl-CoA, and the deacetylase SIRT2 stimulated by NAD+, regulate PAX7 acetylation in muscle stem cells, which in turn, regulates stem cell self-renewal and regeneration following injury in mouse skeletal muscle.
- Marie-Claude Sincennes
- , Caroline E. Brun
- & Michael A. Rudnicki
-
Article
| Open Access Landscape of allele-specific transcription factor binding in the human genome
Landscape of allele-specific transcription factor binding in the human genome
Single-nucleotide variants in enhancers or promoters may affect gene transcription by altering transcription factor binding sites. Here the authors present a meta-analysis empowered by a new statistical method covering thousands of ChIP-Seq experiments resulting in the identification of more than 500 thousand allele-specific binding (ASB) events in the human genome.
- Sergey Abramov
- , Alexandr Boytsov
- & Ivan V. Kulakovskiy
-
Article
| Open Access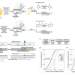 CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
Base editors can inactivate splice sites or introduce stop codons into a gene sequence. Here the authors present SpliceR to design, rank, and test sgRNAs for efficient gene disruption in T cells.
- Mitchell G. Kluesner
- , Walker S. Lahr
- & Branden S. Moriarity
-
Article
| Open Access Circadian clock dysfunction in human omental fat links obesity to metabolic inflammation
Circadian clock dysfunction in human omental fat links obesity to metabolic inflammation
Whether chronic inflammation contributes to metabolic disease through the dysregulation of circadian systems remains incompletely understood in humans. Here the authors show that circadian clock function is perturbed in adipose tissue from individuals with obesity, and that inhibition of NFkB improves clock function.
- Eleonore Maury
- , Benoit Navez
- & Sonia M. Brichard
-
Article
| Open Access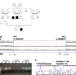 Deletion of CTCF sites in the SHH locus alters enhancer–promoter interactions and leads to acheiropodia
Deletion of CTCF sites in the SHH locus alters enhancer–promoter interactions and leads to acheiropodia
Acheiropodia is associated with homozygous deletions in the LMBR1 gene around ZRS, an enhancer regulating SHH during limb development, but how these deletions lead to this phenotype is unknown. Here the authors use whole-genome sequencing, ChIP-seq, 4C-seq and DNA FISH to show that alterations in CTCF motifs are responsible via altered enhancer–promoter interactions.
- Aki Ushiki
- , Yichi Zhang
- & Nadav Ahituv
-
Article
| Open Access Redundant and non-redundant cytokine-activated enhancers control Csn1s2b expression in the lactating mouse mammary gland
Redundant and non-redundant cytokine-activated enhancers control Csn1s2b expression in the lactating mouse mammary gland
Enhancers and promoters work together to actively regulate gene expression affecting several biological processes. Here, the authors provide molecular insights into the regulation of enhancers and super-enhancers in the Csn1s2b locus during lactation.
- Hye Kyung Lee
- , Michaela Willi
- & Lothar Hennighausen
-
Article
| Open Access Widespread reorganisation of pluripotent factor binding and gene regulatory interactions between human pluripotent states
Widespread reorganisation of pluripotent factor binding and gene regulatory interactions between human pluripotent states
The role of transcriptional enhancers and 3D chromatin organisation in coordinating the transition from naive to primed pluripotency remains poorly understood. Here the authors generate a high-resolution atlas of gene regulatory interactions, chromatin profiles and transcription factor occupancy in naive and primed human pluripotent stem cells to provide insights into these developmental processes.
- Peter Chovanec
- , Amanda J. Collier
- & Peter J. Rugg-Gunn
-
Article
| Open Access Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Glucocorticoid receptors (GR) are thought to bind DNA as dimers or monomers, to regulate different transcription pathways. Here, the authors perform genome-wide studies on GRs with mutations that impair dimerization and provide evidence that monomeric GRs do not play a significant physiologic role.
- Thomas A. Johnson
- , Ville Paakinaho
- & Diego M. Presman
-
Article
| Open Access The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
DNA methylation turnover is an essential epigenetic process during development. Here, the authors look at the changes in DNA methylation during the differentiation of post-mitotic human monocytes (MO), and find that EGR2 interacts with TET2 and is required for DNA demethylation at its binding sites; revealing EGR2 as an epigenetic pioneer factor in human MO.
- Karina Mendes
- , Sandra Schmidhofer
- & Michael Rehli
-
Article
| Open Access Structural basis of ribosomal RNA transcription regulation
Structural basis of ribosomal RNA transcription regulation
Ribosomal RNA (rRNA) expression is regulated at the initiation stage of RNA synthesis. Here, the authors report cryo-EM structures of E. coli RNA polymerase and rRNA promoter complex with DksA/ppGpp on the way to open complex formation, identifying key steps in promoter recognition and opening.
- Yeonoh Shin
- , M. Zuhaib Qayyum
- & Katsuhiko S. Murakami
-
Article
| Open Access GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super-enhancers
GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super-enhancers
Mast cells are critical effectors of allergic inflammation and protection against parasitic infections. Here the authors demonstrate that GATA2 promotes chromatin accessibility at the super-enhancers of mast cell identity genes and primes both typical and super-enhancers at genes that respond to antigenic stimulation.
- Yapeng Li
- , Junfeng Gao
- & Hua Huang
-
Article
| Open Access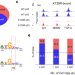 Distinct mechanisms control genome recognition by p53 at its target genes linked to different cell fates
Distinct mechanisms control genome recognition by p53 at its target genes linked to different cell fates
The tumor suppressor p53 is a master regulator of cellular stress response pathways, including cell cycle arrest and apoptosis. Here, the authors identify molecular mechanisms of p53 binding to high- and low-affinity p53 response elements in the genome, linked to cell cycle arrest and pro-apoptotic genes, respectively.
- Marina Farkas
- , Hideharu Hashimoto
- & Steven B. McMahon
-
Article
| Open Access Dissection of the Fgf8 regulatory landscape by in vivo CRISPR-editing reveals extensive intra- and inter-enhancer redundancy
Dissection of the Fgf8 regulatory landscape by in vivo CRISPR-editing reveals extensive intra- and inter-enhancer redundancy
Developmental genes are often regulated by multiple elements, yet their relative contribution to gene expression remains poorly understood. Here the authors apply in vivo CRISPR/Cas9 genome engineering to find two distinct regulatory logics directing Fgf8 in the limb apical ectodermal ridge and the midbrain-hindbrain boundary.
- A. Hörnblad
- , S. Bastide
- & F. Spitz
-
Article
| Open Access BET inhibition disrupts transcription but retains enhancer-promoter contact
BET inhibition disrupts transcription but retains enhancer-promoter contact
The role of BRD4 and Mediator in regulating enhancer-promoter interactions is poorly understood. Here the authors find that treatment with BET inhibitors or pharmacological degradation of BRD4 disrupts transcription while having very little effect on enhancer-promoter interactions.
- Nicholas T. Crump
- , Erica Ballabio
- & Thomas A. Milne
-
Article
| Open Access Order and stochasticity in the folding of individual Drosophila genomes
Order and stochasticity in the folding of individual Drosophila genomes
Genomes are partitioned into topologically associating domains (TADs). Here the authors present single-nucleus Hi-C maps in Drosophila at 10 kb resolution, demonstrating the presence of chromatin compartments in individual nuclei, and partitioning of the genome into non-hierarchical TADs at the scale of 100 kb, which resembles population TAD profiles.
- Sergey V. Ulianov
- , Vlada V. Zakharova
- & Sergey V. Razin
-
Article
| Open Access XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
Hepatocyte 12-hour rhythms have a role in cellular stress and metabolic functions. Here, the authors demonstrate disrupting the 12-hour clock through deletion of XBP1 is associated with the development of NAFLD as well as disruption of phospholipid composition and the maintenance of lipid homeostasis.
- Huan Meng
- , Naomi M. Gonzales
- & Bert W. O’Malley

 Cryptococcal Hsf3 controls intramitochondrial ROS homeostasis by regulating the respiratory process
Cryptococcal Hsf3 controls intramitochondrial ROS homeostasis by regulating the respiratory process