Article
|
Open Access
Featured
-
-
Article
| Open Access Order and stochasticity in the folding of individual Drosophila genomes
Order and stochasticity in the folding of individual Drosophila genomes
Genomes are partitioned into topologically associating domains (TADs). Here the authors present single-nucleus Hi-C maps in Drosophila at 10 kb resolution, demonstrating the presence of chromatin compartments in individual nuclei, and partitioning of the genome into non-hierarchical TADs at the scale of 100 kb, which resembles population TAD profiles.
- Sergey V. Ulianov
- , Vlada V. Zakharova
- & Sergey V. Razin
-
Article
| Open Access XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
XBP1 links the 12-hour clock to NAFLD and regulation of membrane fluidity and lipid homeostasis
Hepatocyte 12-hour rhythms have a role in cellular stress and metabolic functions. Here, the authors demonstrate disrupting the 12-hour clock through deletion of XBP1 is associated with the development of NAFLD as well as disruption of phospholipid composition and the maintenance of lipid homeostasis.
- Huan Meng
- , Naomi M. Gonzales
- & Bert W. O’Malley
-
Article
| Open Access Coupling of H3K27me3 recognition with transcriptional repression through the BAH-PHD-CPL2 complex in Arabidopsis
Coupling of H3K27me3 recognition with transcriptional repression through the BAH-PHD-CPL2 complex in Arabidopsis
Histone 3 Lys 27 trimethylation (H3K27me3) mediates epigenetic silencing of gene expression. Here, Zhang et al. show that in Arabidopsis, the BAH-domain H3K27me3-reader protein AIPP3 forms a complex with PHD proteins and CPL2, a plant-specific Pol II phosphatase, to inhibit Pol II activity by dephosphorylation.
- Yi-Zhe Zhang
- , Jianlong Yuan
- & Cheng-Guo Duan
-
Article
| Open Access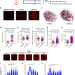 TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
The TGFβ signaling pathway has been shown to regulate transcription by regulating enhancer activity. Here, the authors perform a comprehensive analysis of enhancers in normal mammary epithelial gland cells to elucidate how TGFβ-dependent enhancers control gene transcription in these cells.
- Jose A. Guerrero-Martínez
- , María Ceballos-Chávez
- & Jose C. Reyes
-
Article
| Open Access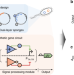 Synthetic protein-binding DNA sponge as a tool to tune gene expression and mitigate protein toxicity
Synthetic protein-binding DNA sponge as a tool to tune gene expression and mitigate protein toxicity
Decoy binding sites are natural regulators of gene expression. Here the authors design synthetic DNA sponges that fine tune the performance of synthetic gene circuits in a simple yet systematic manner, expanding the synthetic biology toolkit for gene regulation.
- Xinyi Wan
- , Filipe Pinto
- & Baojun Wang
-
Article
| Open Access Population-scale study of eRNA transcription reveals bipartite functional enhancer architecture
Population-scale study of eRNA transcription reveals bipartite functional enhancer architecture
Enhancer RNAs are transcribed bidirectionally from core transcription initiation regions. Here, by employing nascent RNA sequencing, the authors identify quantitative trait loci (QTLs) associated with enhancer RNA level and directionality, revealing the bipartite architecture of enhancers.
- Katla Kristjánsdóttir
- , Alexis Dziubek
- & Hojoong Kwak
-
Article
| Open Access Enhancer remodeling promotes tumor-initiating activity in NRF2-activated non-small cell lung cancers
Enhancer remodeling promotes tumor-initiating activity in NRF2-activated non-small cell lung cancers
Aberrant activation of NRF2 in cancer cells contributes to tumorigenicity and therapeutic resistance. Here, the authors show that NRF2 cooperates with CEBPB and remodels enhancers to confer tumor-initiating activity on NRF2- activated non-small cell lung cancers.
- Keito Okazaki
- , Hayato Anzawa
- & Hozumi Motohashi
-
Article
| Open Access Co-regulation of the transcription controlling ATF2 phosphoswitch by JNK and p38
Co-regulation of the transcription controlling ATF2 phosphoswitch by JNK and p38
The ATF2 transcription factor is phosphorylated by different mitogen-activated protein (MAP) kinases. Here, the authors show that the functionally distinct MAP kinases JNK and p38 control ATF2 through different binding sites and differential phosphorylation, thereby modulating ATF2’s sensitivity to the JNK and p38 pathways.
- Klára Kirsch
- , András Zeke
- & Attila Reményi
-
Article
| Open Access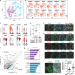 Reconstitution of prospermatogonial specification in vitro from human induced pluripotent stem cells
Reconstitution of prospermatogonial specification in vitro from human induced pluripotent stem cells
Spermatogonia establishment in the fetal and postnatal period is essential for spermatozoa production. Here the authors present a protocol for in vitro reconstitution of human prospermatogonial specification and perform single cell RNA-sequencing to delineate lineage trajectories.
- Young Sun Hwang
- , Shinnosuke Suzuki
- & Kotaro Sasaki
-
Article
| Open Access TASOR is a pseudo-PARP that directs HUSH complex assembly and epigenetic transposon control
TASOR is a pseudo-PARP that directs HUSH complex assembly and epigenetic transposon control
The HUSH complex plays a key role in controlling transcription of viruses and transposable elements. Here, the authors define the biochemical basis of HUSH assembly and show that the TASOR subunit contains a pseudo-PARP domain critical for HUSH-dependent transgene repression and H3K9me3 deposition over targets genome wide.
- Christopher H. Douse
- , Iva A. Tchasovnikarova
- & Yorgo Modis
-
Article
| Open Access Structure of the TFIIIC subcomplex τA provides insights into RNA polymerase III pre-initiation complex formation
Structure of the TFIIIC subcomplex τA provides insights into RNA polymerase III pre-initiation complex formation
Transcription factor TFIIIC plays roles in Pol III transcription and in chromatin organization. CryoEM structure of the yeast TFIIIC subcomplex τA, a negative stain reconstruction of τA bound to the TFIIIB subunits Brf1 and TBP and accompanying biochemistry suggest how τA achieves positioning of TFIIIB upstream of the TSS and remodeling of the TFIIIC complex during assembly of TFIIIB.
- Matthias K. Vorländer
- , Anna Jungblut
- & Christoph W. Müller
-
Article
| Open Access Hyperacetylated chromatin domains mark cell type-specific genes and suggest distinct modes of enhancer function
Hyperacetylated chromatin domains mark cell type-specific genes and suggest distinct modes of enhancer function
Super-enhancer are usually defined by high levels of chromatin modification and associate with cell-specific gene expression. Here, the authors define hyperacetylated chromatin domains (HCDs) by using histone hyperacetylation peak breadth information and show that HCDs associated more closely with cell identity than super-enhancers.
- Sierra Fox
- , Jacquelyn A. Myers
- & Michael Bulger
-
Article
| Open Access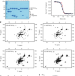 Taf14 recognizes a common motif in transcriptional machineries and facilitates their clustering by phase separation
Taf14 recognizes a common motif in transcriptional machineries and facilitates their clustering by phase separation
S. cerevisiae TBP associated factor 14 (Taf14) is a transcriptional regulator that interacts with multiple nuclear complexes. Here, the authors report that the extra-terminal domain of Taf14 (Taf14ET) recognizes a common motif in various transcriptional coactivator proteins and they solve the NMR structure of Taf14ET bound the ET-binding motif of Sth1, the catalytic subunit of the RSC (Remodel the Structure of Chromatin) complex, and furthermore show that Taf14ET undergoes liquid-liquid phase separation, which is enhanced by Taf14 interaction partners.
- Guochao Chen
- , Duo Wang
- & Yong Chen
-
Article
| Open Access Interaction of the pioneer transcription factor GATA3 with nucleosomes
Interaction of the pioneer transcription factor GATA3 with nucleosomes
GATA 3 functions as a pioneer factor during cellular reprogramming. Here the authors delineate nucleosome positioning relative to GATA3 binding motifs and describe the structure of a GATA3–nucleosome complex; providing insight into how a pioneer factor interacts with nucleosomes and catalyze their local remodelling to produce an accessible enhancer.
- Hiroki Tanaka
- , Yoshimasa Takizawa
- & Hitoshi Kurumizaka
-
Article
| Open Access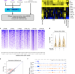 Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia
Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia
Transposable elements are a potential source of transcriptional regulators, but how these sequences contribute to oncogenesis remains poorly understood. Here, the authors identify endogenous retroviruses (ERVs) with acute myeloid leukemia (AML)-associated enhancer chromatin signatures, and provide evidence that ERV activation provides an additional layer of gene regulation in AML.
- Özgen Deniz
- , Mamataz Ahmed
- & Miguel R. Branco
-
Article
| Open Access Enhancer and super-enhancer dynamics in repair after ischemic acute kidney injury
Enhancer and super-enhancer dynamics in repair after ischemic acute kidney injury
Acute kidney injury is a major health problem amongst hospitalized patients. Here the authors provide a comprehensive characterization of enhancer and super-enhancer elements, and the transcription factor motifs associated with these elements in response to kidney injury in vivo; providing insight into the regulation of kidney repair.
- Julia Wilflingseder
- , Michaela Willi
- & Joseph V. Bonventre
-
Article
| Open Access Intragenic recruitment of NF-κB drives splicing modifications upon activation by the oncogene Tax of HTLV-1
Intragenic recruitment of NF-κB drives splicing modifications upon activation by the oncogene Tax of HTLV-1
The nuclear factors κB (NF-κB) is a transcription factor involved in immune functions, inflammation, and cancer. Here, the authors show that the NF-κB factor RELA regulates splicing of target genes by recruiting DDX17 on chromatin upon expression of the viral oncogene Tax.
- Lamya Ben Ameur
- , Paul Marie
- & Franck Mortreux
-
Article
| Open Access Regulated repression governs the cell fate promoter controlling yeast meiosis
Regulated repression governs the cell fate promoter controlling yeast meiosis
In budding yeast, the decision to enter meiosis is defined by nutrient and mating-type signals regulating expression of the master transcription factor for meiotic entry, IME1. Here, the authors characterize the steps involved in regulating the entering in meiosis via the co-repressor complex Tup1-Cyc8.
- Janis Tam
- & Folkert J. van Werven
-
Article
| Open Access The scaffold protein p62 regulates adaptive thermogenesis through ATF2 nuclear target activation
The scaffold protein p62 regulates adaptive thermogenesis through ATF2 nuclear target activation
Beta-adrenergic stimulation of brown adipose tissue leads to thermogenesis via the activating transcription factor 2 (ATF2) mediated expression of the thermogenic genes Ucp1 and Pgc-1α. Here, the authors show that the scaffold protein p62 regulates brown adipose tissue function through modifying ATF2 genomic binding and subsequent Ucp1 and Pgc-1α induction.
- Katrin Fischer
- , Anna Fenzl
- & Timo D. Müller
-
Article
| Open Access Candidate silencer elements for the human and mouse genomes
Candidate silencer elements for the human and mouse genomes
Identification of silencer elements by computational or experimental approaches in a genome-wide manner is still challenging. Here authors define uncharacterized cis-regulatory elements (CREs) in human and mouse genomes likely containing silencer elements, and test them in cells using massively parallel reporter assays to identify silencer elements that showed silencer activity.
- Naresh Doni Jayavelu
- , Ajay Jajodia
- & R. David Hawkins
-
Article
| Open Access Role of cell-type specific nucleosome positioning in inducible activation of mammalian promoters
Role of cell-type specific nucleosome positioning in inducible activation of mammalian promoters
Nucleosome organisation plays important roles in regulating functional genomic elements. Here, the authors use high-resolution profiling to analyse dynamic nucleosome positioning at inducible and cell-type-specific promoters, providing a global view of chromatin architecture at inducible promoters.
- Agata Oruba
- , Simona Saccani
- & Dominic van Essen
-
Article
| Open Access C/EBPɑ is crucial determinant of epithelial maintenance by preventing epithelial-to-mesenchymal transition
C/EBPɑ is crucial determinant of epithelial maintenance by preventing epithelial-to-mesenchymal transition
In breast cancer TGF-β regulates C/EBPα and can induce epithelial-to-mesenchymal transition (EMT). Here, the authors show that C/EBPα maintains epithelial homeostasis and prevents EMT-mediated tumorigenesis.
- Ana Rita Lourenço
- , M. Guy Roukens
- & Paul J. Coffer
-
Article
| Open Access The COMET toolkit for composing customizable genetic programs in mammalian cells
The COMET toolkit for composing customizable genetic programs in mammalian cells
Engineering mammalian cellular functions requires a toolkit of orthogonal and well-characterized genetic components. Here the authors develop COMET: an ensemble of transcription factors, promoters, and accompanying models for the design and construction of genetic programs.
- Patrick S. Donahue
- , Joseph W. Draut
- & Joshua N. Leonard
-
Article
| Open Access Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Tissues-specific gene expression requires coordinated cis-regulatory elements. Here the authors use dCas9-based enhancer targeting to remodel local epigenetic landscapes and activate or inactive transcription.
- Kailong Li
- , Yuxuan Liu
- & Jian Xu
-
Article
| Open Access Dual-initiation promoters with intertwined canonical and TCT/TOP transcription start sites diversify transcript processing
Dual-initiation promoters with intertwined canonical and TCT/TOP transcription start sites diversify transcript processing
The functional significance of start site choice in promoter architectures is little understood. Here the authors identify in zebrafish development and mammalian cells a class of dual-initiation promoters, in which non-canonical YC dinucleotides reflecting 5’-TOP/TCT initiation are intertwined with canonical YR-initiation.
- Chirag Nepal
- , Yavor Hadzhiev
- & Ferenc Müller
-
Article
| Open Access Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Although transcription factor (TF) cooperativity is widespread, a global mechanistic understanding of the role of TF cooperativity is still lacking. Here the authors introduce a statistical learning framework that provides structural insight into TF cooperativity and its functional consequences based on next generation sequencing data and provide mechanistic insights into TF cooperativity and its impact on protein-phenotype interactions.
- Ignacio L. Ibarra
- , Nele M. Hollmann
- & Judith B. Zaugg
-
Article
| Open Access BEN-solo factors partition active chromatin to ensure proper gene activation in Drosophila
BEN-solo factors partition active chromatin to ensure proper gene activation in Drosophila
The BEN-solo proteins—including Insensitive (Insv), Elba1 and Elba2—function in both transcriptional repression and chromatin insulation. Here, the authors investigate the role of these proteins in Drosophila embryos, finding that ELBA and Insv function as general insulators and partition active chromatin to ensure proper gene activation in Drosophila.
- Malin Ueberschär
- , Huazhen Wang
- & Qi Dai
-
Article
| Open Access Molecular insight into RNA polymerase I promoter recognition and promoter melting
Molecular insight into RNA polymerase I promoter recognition and promoter melting
RNA polymerase I (Pol I) catalyses the transcription of pre-ribosomal RNA and for transcription initiation Pol I assembles with core factor and Rrn3 on the rDNA core promoter. Here the authors provide insights into the molecular mechanism of promoter opening by Pol I by determining the cryo-EM structures of two closed, two intermediate and two open Pol I pre-initiation complexes.
- Yashar Sadian
- , Florence Baudin
- & Christoph W. Müller
-
Article
| Open Access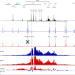 A revised model for promoter competition based on multi-way chromatin interactions at the α-globin locus
A revised model for promoter competition based on multi-way chromatin interactions at the α-globin locus
The coordination of interactions between multiple regulatory elements and genes within a chromatin domain remains poorly understood. Here, the authors use a method to detect multi-way chromatin interactions in a mouse model in which the α-globin domain is extended to include several additional genes, finding that the promoters do not form mutually exclusive interactions with the enhancers, but all interact simultaneously in a single complex.
- A. Marieke Oudelaar
- , Caroline L. Harrold
- & Jim R. Hughes
-
Article
| Open Access An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
Mutations associated with SCN5A, which encodes the major cardiac sodium channel, influence impulse conduction and are associated with arrhythmia disorders. Here, the authors identify an evolutionary conserved, super enhancer-like, regulatory cluster downstream of SCN5A and show that it controls the chromatin architecture of the locus and Scn5a expression.
- Joyce C. K. Man
- , Rajiv A. Mohan
- & Vincent M. Christoffels
-
Article
| Open Access Identification of atrial fibrillation associated genes and functional non-coding variants
Identification of atrial fibrillation associated genes and functional non-coding variants
The majority of disease-associated genetic variants lie in non-coding regions. Here the authors generated and compiled human transcriptomic, epigenomic and chromatin conformation datasets, to identify genes associated with atrial fibrillation and functional non-coding variants.
- Antoinette F. van Ouwerkerk
- , Fernanda M. Bosada
- & Vincent M. Christoffels
-
Article
| Open Access Second messenger Ap4A polymerizes target protein HINT1 to transduce signals in FcεRI-activated mast cells
Second messenger Ap4A polymerizes target protein HINT1 to transduce signals in FcεRI-activated mast cells
The second messenger Ap4A contributes to mast cell activation by interacting with HINT1 but the molecular mechanism is not fully understood. Here, the authors show that Ap4A regulates downstream signaling by polymerizing HINT1 and provide insights into how specificity over other second messengers is achieved.
- Jing Yu
- , Zaizhou Liu
- & Jing Wang
-
Article
| Open Access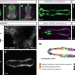 A conserved regulatory program initiates lateral plate mesoderm emergence across chordates
A conserved regulatory program initiates lateral plate mesoderm emergence across chordates
Numerous tissues are derived from the lateral plate mesoderm (LPM) but how this is specified is unclear. Here, the authors identify a pan-LPM reporter activity found in the zebrafish draculin (drl) gene that also shows transgenic activity in LPM-corresponding territories of several chordates, including chicken, axolotl, lamprey, Ciona, and amphioxus.
- Karin D. Prummel
- , Christopher Hess
- & Christian Mosimann
-
Article
| Open Access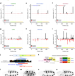 In vivo Hox binding specificity revealed by systematic changes to a single cis regulatory module
In vivo Hox binding specificity revealed by systematic changes to a single cis regulatory module
Hox proteins are expressed in partially overlapping regions to inform development along the embryo’s head-tail axis. Here the authors analyse a cis regulatory module directly regulated by seven different Drosophila Hox proteins to uncover how different Hox class proteins differentially control its expression.
- Carlos Sánchez-Higueras
- , Chaitanya Rastogi
- & James C.-G. Hombría
-
Article
| Open Access Functional genetic variants can mediate their regulatory effects through alteration of transcription factor binding
Functional genetic variants can mediate their regulatory effects through alteration of transcription factor binding
Functional variants have been proposed to alter transcription factor binding. Here, the authors provide direct evidence that functional variants within the TBC1D4 gene, encoding an NFκB binding site, can alter transcription factor binding, and use CRISPR-Cas9 to reveal localization of the transcription factor to be the regulator of chromatin accessibility and p65 binding and ultimately TBC1D4 expression.
- Andrew D. Johnston
- , Claudia A. Simões-Pires
- & John M. Greally
-
Article
| Open Access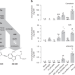 Expansion of a core regulon by transposable elements promotes Arabidopsis chemical diversity and pathogen defense
Expansion of a core regulon by transposable elements promotes Arabidopsis chemical diversity and pathogen defense
Arabidopsis plants can produce 4-hydroxyindole-3-carbonitrile (4OH-ICN) upon pathogen infection. Here, the authors show that EPCOT3, a retrotransposonderived enhancer, mediates WRKY33-binding, pathogen-responsive transcription of CYP82C2, and synthesis of 4OH-ICN.
- Brenden Barco
- , Yoseph Kim
- & Nicole K. Clay
-
Article
| Open Access Regulatory pathways governing murine coronary vessel formation are dysregulated in the injured adult heart
Regulatory pathways governing murine coronary vessel formation are dysregulated in the injured adult heart
How coronary vessels develop and respond to injury is not fully understood. Here, the authors use murine enhancer:reporter models to identify three transcriptional pathways active in different parts of coronary vasculature. These also contribute to neovascularization in the injured neonatal, but not adult, heart.
- Sophie Payne
- , Mala Gunadasa-Rohling
- & Sarah De Val
-
Article
| Open Access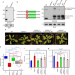 The repressive role of Arabidopsis H2A.Z in transcriptional regulation depends on AtBMI1 activity
The repressive role of Arabidopsis H2A.Z in transcriptional regulation depends on AtBMI1 activity
Arabidopsis H2A.Z plays an important role in regulating gene expression in response to stressors; however, the underlying mechanism is still puzzling. Here, the authors show that monoubiquitination of H2A.Z by AtBMI1 is required for H2A.Z-mediated transcriptional repression.
- Ángeles Gómez-Zambrano
- , Wiam Merini
- & Myriam Calonje
-
Article
| Open Access DOT1L inhibition reveals a distinct subset of enhancers dependent on H3K79 methylation
DOT1L inhibition reveals a distinct subset of enhancers dependent on H3K79 methylation
Histone 3 lysine 79 is mono (me1), di (me2), or tri (me3) methylated by the methyltransferase DOT1L. Here the authors reveal a group of enhancers defined by H3K79me2/3 which regulates enhancer-promoter interactions and other key enhancer features in MLL-AF4 leukemia cells.
- Laura Godfrey
- , Nicholas T. Crump
- & Thomas A. Milne
-
Article
| Open Access Chromatin interaction maps reveal genetic regulation for quantitative traits in maize
Chromatin interaction maps reveal genetic regulation for quantitative traits in maize
Spatial organization of regulatory elements and its impact on gene expression in plants remain unclear. Here, the authors construct maize chromatin interaction maps using chromatin interaction analysis by paired-end tag sequencing (ChIA-PET) and show their associations with gene expression and agronomic traits.
- Yong Peng
- , Dan Xiong
- & Xingwang Li
-
Article
| Open Access Long-range interactions between proximal and distal regulatory regions in maize
Long-range interactions between proximal and distal regulatory regions in maize
Chromatin interaction analysis by paired-end tag sequencing (ChIA-PET) can discover specific protein-centered chromatin interactions in high resolution. Here, the authors use ChIA-PET to reveal the complex and dynamic interactions between proximal and distal regulatory regions of genes in maize.
- En Li
- , Han Liu
- & Jinsheng Lai
-
Article
| Open Access Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
The type II nuclear receptors (NRs) and the retinoid X receptor (RXR) form heterodimeric transcription factors to regulate development, metabolism, and inflammation. Here the authors employ protein-binding microarrays to comprehensively analyze the DNA binding of 12 NR:RXRα heterodimers, and report promiscuous NR-DNA binding.
- Ashley Penvose
- , Jessica L. Keenan
- & Trevor Siggers
-
Article
| Open Access ZFP30 promotes adipogenesis through the KAP1-mediated activation of a retrotransposon-derived Pparg2 enhancer
ZFP30 promotes adipogenesis through the KAP1-mediated activation of a retrotransposon-derived Pparg2 enhancer
Although Krüppel-associated box zinc finger proteins (KZFPs) were found to mostly repress transposable elements, recent studies found that KFZPs also play other roles in cells. Here, the authors provide evidence that the KZFP ZFP30 promotes adipogenesis by targeting and activating a retrotransposon-derived Pparg2 enhancer in cooperation with co-regulator KRAB-associated protein 1 (KAP1).
- Wanze Chen
- , Petra C. Schwalie
- & Bart Deplancke
-
Article
| Open Access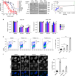 CDK12 loss in cancer cells affects DNA damage response genes through premature cleavage and polyadenylation
CDK12 loss in cancer cells affects DNA damage response genes through premature cleavage and polyadenylation
Cdk12 is primarily involved in the regulation of DNA damage response (DDR) gene transcription as well as mRNA processing. Here, the authors demonstrate that CDK12 suppresses intronic polyadenylation, and that inhibition of this kinase primarily affects the expression of long genes with higher numbers of polyA sites, features common to many DDR genes.
- Malgorzata Krajewska
- , Ruben Dries
- & Rani E. George
-
Article
| Open Access MYB96 recruits the HDA15 protein to suppress negative regulators of ABA signaling in Arabidopsis
MYB96 recruits the HDA15 protein to suppress negative regulators of ABA signaling in Arabidopsis
MYB96 can regulate both positive and negative regulators of ABA signaling to maximize plant drought tolerance. Here, the authors show that MYB96 represses expression of ABA negative regulators in Arabidopsis by interacting with HDA15 and promoting histone deacetylation at the cognate regions.
- Hong Gil Lee
- & Pil Joon Seo
-
Article
| Open Access An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
CTCF plays key roles in gene regulation, chromatin insulation and organizing the higher-order chromatin architecture of mammalian genomes. Here the authors investigate the function an alternatively spliced shorter CTCF isoform, finding that this isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis.
- Jiao Li
- , Kaimeng Huang
- & Hongjie Yao
-
Article
| Open Access A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
A TetR-family transcription factor regulates fatty acid metabolism in the archaeal model organism Sulfolobus acidocaldarius
Certain archaea appear to metabolize fatty acids, but the regulation of these pathways is unclear. Here, Wang et al. provide genetic, functional and structural evidence supporting that a TetR-family transcriptional regulator is involved in regulation of fatty acid metabolism in Sulfolobus acidocaldarius.
- Kun Wang
- , David Sybers
- & Eveline Peeters
-
Article
| Open Access The anti-cancer drugs curaxins target spatial genome organization
The anti-cancer drugs curaxins target spatial genome organization
Curaxins are a recently discovered class of anti-cancer agents that disturbs DNA/histone interactions within. Here the authors provide evidence that curaxins affect the spatial genome organization and compromise enhancer-promoter communication necessary for expression of several oncogenes, including MYC.
- Omar L. Kantidze
- , Artem V. Luzhin
- & Sergey V. Razin
-
Article
| Open Access Functional genomics reveal gene regulatory mechanisms underlying schizophrenia risk
Functional genomics reveal gene regulatory mechanisms underlying schizophrenia risk
We know a large number of risk SNPs for schizophrenia, but little about how these SNPs contribute to the disorder. Here, the authors use functional genomics to identify risk SNPs that disrupt transcription factor binding and validate the regulatory effects of the transcription factor binding-disrupting SNPs.
- Yongxia Huo
- , Shiwu Li
- & Xiong-Jian Luo

 BET inhibition disrupts transcription but retains enhancer-promoter contact
BET inhibition disrupts transcription but retains enhancer-promoter contact