Featured
-
-
Article
| Open Access Uncovering interpretable potential confounders in electronic medical records
Uncovering interpretable potential confounders in electronic medical records
Randomized clinical trials are often plagued by selection bias, and expert-selected covariates may insufficiently adjust for confounding factors. Here, the authors develop a framework based on natural language processing to uncover interpretable potential confounders from text.
- Jiaming Zeng
- , Michael F. Gensheimer
- & Ross D. Shachter
-
Article
| Open Access Cancer patient survival can be parametrized to improve trial precision and reveal time-dependent therapeutic effects
Cancer patient survival can be parametrized to improve trial precision and reveal time-dependent therapeutic effects
Analysis of more than 150 Phase 3 oncology clinical trials supports parametric statistical analysis, significantly increasing the precision of small early-phase trials and relating deviations from the Cox proportional hazards model to trial duration.
- Deborah Plana
- , Geoffrey Fell
- & Peter K. Sorger
-
Article
| Open Access Impacts of rapid mass vaccination against SARS-CoV2 in an early variant of concern hotspot
Impacts of rapid mass vaccination against SARS-CoV2 in an early variant of concern hotspot
Schwaz, Austria, experienced SARS-CoV-2 outbreaks caused by variants of concern in early 2021 and conducted a mass vaccination campaign in response, with 70% of the adult population vaccinated after 5 days. Here, the authors show that this campaign resulted in reduced infections and hospitalisations.
- Jörg Paetzold
- , Janine Kimpel
- & Hannes Winner
-
Article
| Open Access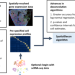 Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data
Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data
The deconvolution of cell types is challenging in spatially-resolved transcriptomics. Here, the authors present SpatialDecon, a method for the deconvolution and quantification of cell types in spatial transcriptomics data, and show how it can be used to analyse immune response heterogeneity in cancer.
- Patrick Danaher
- , Youngmi Kim
- & Joseph M. Beechem
-
Article
| Open Access Microbiome differential abundance methods produce different results across 38 datasets
Microbiome differential abundance methods produce different results across 38 datasets
Many microbiome differential abundance methods are available, but it lacks systematic comparison among them. Here, the authors compare the performance of 14 differential abundance testing methods on 38 16S rRNA gene datasets with two sample groups, and show ALDEx2 and ANCOM-II produce the most consistent results.
- Jacob T. Nearing
- , Gavin M. Douglas
- & Morgan G. I. Langille
-
Article
| Open Access Zero-preserving imputation of single-cell RNA-seq data
Zero-preserving imputation of single-cell RNA-seq data
Missing values in scRNA-seq datasets can bias their analysis. Here, the authors threshold the low rank approximation of the expression matrix, so false zeros can be imputed while true zeros are preserved.
- George C. Linderman
- , Jun Zhao
- & Yuval Kluger
-
Article
| Open Access Simultaneous estimation of bi-directional causal effects and heritable confounding from GWAS summary statistics
Simultaneous estimation of bi-directional causal effects and heritable confounding from GWAS summary statistics
Mendelian Randomization approaches are being increasingly refined, but certain statistical limitations hinder their application to GWAS. Here, the authors propose a new Mendelian Randomization method to estimate bi- directional causal effects and explicitly account for heritable confounding.
- Liza Darrous
- , Ninon Mounier
- & Zoltán Kutalik
-
Article
| Open Access Probabilistic inference of the genetic architecture underlying functional enrichment of complex traits
Probabilistic inference of the genetic architecture underlying functional enrichment of complex traits
Improving inference in large-scale genetic data linked to electronic medical record data requires the development of novel computationally efficient regression methods. Here, the authors develop a Bayesian approach for association analyses to improve SNP-heritability estimation, discovery, fine-mapping and genomic prediction.
- Marion Patxot
- , Daniel Trejo Banos
- & Matthew R. Robinson
-
Article
| Open Access High-throughput mediation analysis of human proteome and metabolome identifies mediators of post-bariatric surgical diabetes control
High-throughput mediation analysis of human proteome and metabolome identifies mediators of post-bariatric surgical diabetes control
Factors underlying the effects of gastric bypass surgery on glucose homeostasis are incompletely understood. Here the authors developed and applied high-throughput mediation analysis to identify proteome/metabolome mediators of improved glucose homeostasis after to gastric bypass surgery, and report that improved glycemia was mediated by the growth hormone receptor.
- Jonathan M. Dreyfuss
- , Yixing Yuchi
- & Mary Elizabeth Patti
-
Article
| Open Access SARS-CoV-2 transmission across age groups in France and implications for control
SARS-CoV-2 transmission across age groups in France and implications for control
In this study, Tran Kiem et al. examine the contribution of different age groups to COVID-19 transmission. Using data from the French epidemic in summer 2020, they report that while individuals aged 80 years and older are more at risk, pandemic control in the absence of vaccines required measures targeted at all age groups.
- Cécile Tran Kiem
- , Paolo Bosetti
- & Simon Cauchemez
-
Article
| Open Access A benchmark study of simulation methods for single-cell RNA sequencing data
A benchmark study of simulation methods for single-cell RNA sequencing data
Simulation is useful for developing and evaluating computational methods. Here, the authors develop a comprehensive evaluation framework, SimBench, to benchmark Single-cell RNA-seq simulation methods through a diverse collection of experimental datasets.
- Yue Cao
- , Pengyi Yang
- & Jean Yee Hwa Yang
-
Article
| Open Access scCODA is a Bayesian model for compositional single-cell data analysis
scCODA is a Bayesian model for compositional single-cell data analysis
Imbalance and loss of cell types is a hallmark in many diseases. Still, quantifying compositional changes in scRNAseq data remains challenging. Here the authors present scCODA, a Bayesian model to assess cell type compositions in scRNA-seq data.
- M. Büttner
- , J. Ostner
- & B. Schubert
-
Article
| Open Access Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Accurate and scalable variant calling from single cell DNA sequencing data with ProSolo
Obtaining accurate variant calls from multiple displacement amplified single cell DNA sequencing data needs dedicated models that account for amplification bias and copy errors. Here, the authors describe ProSolo, a model for calling single nucleotide variants with control over the false discovery rate.
- David Lähnemann
- , Johannes Köster
- & Alexander Schönhuth
-
Article
| Open Access Model-based assessment of Chikungunya and O’nyong-nyong virus circulation in Mali in a serological cross-reactivity context
Model-based assessment of Chikungunya and O’nyong-nyong virus circulation in Mali in a serological cross-reactivity context
O’nyong nyong and Chikungunya virus are arboviruses present in Africa but their prevalence is unknown, partly due to high antibody cross-reactivity with one another. Here, the authors develop a statistical model that accounts for cross-reactivity to characterise circulation of both viruses from seroprevalence surveys.
- Nathanaël Hozé
- , Issa Diarra
- & Simon Cauchemez
-
Article
| Open Access scPower accelerates and optimizes the design of multi-sample single cell transcriptomic studies
scPower accelerates and optimizes the design of multi-sample single cell transcriptomic studies
scRNASeq data is revolutionizing our understanding of biological systems, but is still expensive to generate. Here, the authors present a statistical framework that facilitates informed multi-sample experimental design to reduce unnecessary costs and maximize the utility of the generated data.
- Katharina T. Schmid
- , Barbara Höllbacher
- & Matthias Heinig
-
Article
| Open Access Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Normalisr removes technical bias in single-cell RNA-seq and detects gene differential and coexpression accurately and efficiently. It also infers gene regulatory and co-expression networks from conventional and CRISPR screen single-cell RNA-seq datasets.
- Lingfei Wang
-
Article
| Open Access Using secondary cases to characterize the severity of an emerging or re-emerging infection
Using secondary cases to characterize the severity of an emerging or re-emerging infection
Estimates of the severity of emerging infections did not consider the case ascertainment method, but secondary cases identified by contact tracing of index cases may be more reliable as they are less susceptible to ascertainment bias. Here, the authors perform a systematic review to quantify these differences and model their impacts for COVID-19.
- Tim K. Tsang
- , Can Wang
- & Benjamin J. Cowling
-
Article
| Open Access Quantifying previous SARS-CoV-2 infection through mixture modelling of antibody levels
Quantifying previous SARS-CoV-2 infection through mixture modelling of antibody levels
The proportion of a population that has previously been infected by a pathogen is typically estimated using antibody thresholds adjusted for sensitivity and specificity. Here, the authors present a model-based alternative to threshold methods which accounts for antibody waning and other sources of spectrum bias.
- C. Bottomley
- , M. Otiende
- & J. A. G. Scott
-
Article
| Open Access Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Bayesian log-normal deconvolution for enhanced in silico microdissection of bulk gene expression data
Deconvolution methods reveal individual cell types in complex tissues profiled by bulk methods. Here the authors present a Bayesian deconvolution method that outperforms existing methods when benchmarked on >700 datasets, especially in estimating cell-type-specific gene expression profiles.
- Bárbara Andrade Barbosa
- , Saskia D. van Asten
- & Yongsoo Kim
-
Article
| Open Access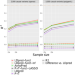 Incorporating functional priors improves polygenic prediction accuracy in UK Biobank and 23andMe data sets
Incorporating functional priors improves polygenic prediction accuracy in UK Biobank and 23andMe data sets
Incorporating functional information has shown promise for improving polygenic risk prediction of complex traits. Here, the authors describe polygenic prediction method LDpred-funct, and demonstrate its utility across 21 heritable traits in the UK Biobank.
- Carla Márquez-Luna
- , Steven Gazal
- & Alkes L. Price
-
Article
| Open Access Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Calibrated rare variant genetic risk scores for complex disease prediction using large exome sequence repositories
Identifying associations of rare variants with disease is challenging due to small effect sizes, technical artefacts and population structure heterogeneity. Here, the authors present RV-EXCALIBER, a method that uses large summary-level exome data to robustly calibrate rare variant burden.
- Ricky Lali
- , Michael Chong
- & Guillaume Paré
-
Article
| Open Access Efficient generative modeling of protein sequences using simple autoregressive models
Efficient generative modeling of protein sequences using simple autoregressive models
Deep learning is a powerful tool for the design of novel protein sequences, yet can be computationally very inefficient. Here the authors propose using simple forecasting models to efficiently generate a large number of novel protein structures.
- Jeanne Trinquier
- , Guido Uguzzoni
- & Martin Weigt
-
Article
| Open Access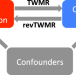 Differentially expressed genes reflect disease-induced rather than disease-causing changes in the transcriptome
Differentially expressed genes reflect disease-induced rather than disease-causing changes in the transcriptome
Identification of gene expression changes between healthy and diseased individuals can reveal mechanistic insights and biomarkers. Here, the authors propose a bi-directional transcriptome-wide Mendelian Randomization approach to assess causal effects between gene expression and complex traits.
- Eleonora Porcu
- , Marie C. Sadler
- & Zoltán Kutalik
-
Article
| Open Access Generalized and scalable trajectory inference in single-cell omics data with VIA
Generalized and scalable trajectory inference in single-cell omics data with VIA
Scalable trajectory inference for multi-omic single cell datasets is challenging in terms of capturing non-tree complex topologies. Here the authors present a method, VIA, that scales to millions of cells across multiple omic modalities using lazy-teleporting random walks.
- Shobana V. Stassen
- , Gwinky G. K. Yip
- & Kevin K. Tsia
-
Article
| Open Access Inferring multilayer interactome networks shaping phenotypic plasticity and evolution
Inferring multilayer interactome networks shaping phenotypic plasticity and evolution
Genetic plasticity drives phenotypic differences. Here, the authors develop a framework to quantify the individual and combinatorial contributions of SNPs on a phenotype of interest and use it to identify SNP-SNP interactions associated with variations in bacteria’s response to external changes.
- Dengcheng Yang
- , Yi Jin
- & Rongling Wu
-
Article
| Open Access Correcting for sparsity and interdependence in glycomics by accounting for glycan biosynthesis
Correcting for sparsity and interdependence in glycomics by accounting for glycan biosynthesis
Glycomics can uncover important molecular changes but measured glycans are highly interconnected and incompatible with common statistical methods, introducing pitfalls during analysis. Here, the authors develop an approach to identify glycan dependencies across samples to facilitate comparative glycomics.
- Bokan Bao
- , Benjamin P. Kellman
- & Nathan E. Lewis
-
Article
| Open Access A hierarchical approach to removal of unwanted variation for large-scale metabolomics data
A hierarchical approach to removal of unwanted variation for large-scale metabolomics data
Mass spectrometry-based metabolomics is a powerful method for profiling large clinical cohorts but batch variations can obscure biologically meaningful differences. Here, the authors develop a computational workflow that removes unwanted data variation while preserving biologically relevant information.
- Taiyun Kim
- , Owen Tang
- & Jean Yee Hwa Yang
-
Article
| Open Access GapClust is a light-weight approach distinguishing rare cells from voluminous single cell expression profiles
GapClust is a light-weight approach distinguishing rare cells from voluminous single cell expression profiles
While rare cell type identification is indispensable in single cell studies, powerful tools with high detection accuracy and computational efficiency are still lacking. Here, the authors propose a light-weight algorithm which can distinguish rare cell types from voluminous single cell expression profiles.
- Botao Fa
- , Ting Wei
- & Zhangsheng Yu
-
Article
| Open Access Sensitive detection of tumor mutations from blood and its application to immunotherapy prognosis
Sensitive detection of tumor mutations from blood and its application to immunotherapy prognosis
It is possible to call single-nucleotide variant (SNV) in cell-free DNA (cfDNA), but the accuracy of detection is often affected by low tumour cfDNA content. Here, the authors develop a method, cfSNV, and show that it can be used even for medium-coverage whole exome sequencing of cfDNA.
- Shuo Li
- , Zorawar S. Noor
- & Xianghong Jasmine Zhou
-
Article
| Open Access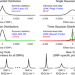 Improved genetic prediction of complex traits from individual-level data or summary statistics
Improved genetic prediction of complex traits from individual-level data or summary statistics
Existing genetic prediction tools typically assume that genetic variants contribute equally towards the phenotype. The authors develop eight prediction tools that allow the user to specify the heritability model, and show that these tools enable substantially improved prediction of complex traits.
- Qianqian Zhang
- , Florian Privé
- & Doug Speed
-
Article
| Open Access The T cell receptor repertoire of tumor infiltrating T cells is predictive and prognostic for cancer survival
The T cell receptor repertoire of tumor infiltrating T cells is predictive and prognostic for cancer survival
Precision medicine needs prognostic markers to select the patients that will benefit more from targeted therapy. Authors show here that high level of baseline T cell receptor diversity is an indicator of favourable prognosis in multiple cancer types, and monoclonal expansion of T-cells correlates with good response to immune checkpoint blockade therapy in metastatic melanoma patients.
- Sara Valpione
- , Piyushkumar A. Mundra
- & Richard Marais
-
Article
| Open Access Insights into household transmission of SARS-CoV-2 from a population-based serological survey
Insights into household transmission of SARS-CoV-2 from a population-based serological survey
Household-based studies can provide insights into SARS-CoV-2 transmission. Here, the authors fit transmission models to serological data from Geneva, Switzerland, and estimate that the risk of infection from single household exposure (17.3%) was higher than for extra-household exposure (5.1%).
- Qifang Bi
- , Justin Lessler
- & Didier Trono
-
Article
| Open Access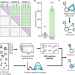 Reliable identification of protein-protein interactions by crosslinking mass spectrometry
Reliable identification of protein-protein interactions by crosslinking mass spectrometry
Cross-linking mass spectrometry (MS) can identify protein-protein interaction (PPI) networks but assessing the reliability of these data remains challenging. To address this issue, the authors develop and validate a method to determine the false-discovery rate of PPIs identified by cross-linking MS.
- Swantje Lenz
- , Ludwig R. Sinn
- & Juri Rappsilber
-
Article
| Open Access Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Pooling participant-level genetic data into a single analysis can result in variance stratification, reducing statistical performance. Here, the authors develop variant-specific inflation factors to assess variance stratification and apply this to pooled individual-level data from whole genome sequencing.
- Tamar Sofer
- , Xiuwen Zheng
- & Kenneth M. Rice
-
Article
| Open Access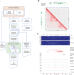 HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP
HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP
The genome-wide investigation of chromatin organization enables insights into global gene expression control. Here, the authors present a computationally efficient method for the analysis of chromatin organization data and use it to recover principles of 3D organization across conditions.
- Merve Sahin
- , Wilfred Wong
- & Christina S. Leslie
-
Article
| Open Access Replicate sequencing libraries are important for quantification of allelic imbalance
Replicate sequencing libraries are important for quantification of allelic imbalance
Allele-specific expression in diploid organisms can be quantified by RNA-seq and it is common practice to rely on a single library. Here, the authors show that the standard approach has variable error rate and present Qllelic as a tool to improve reproducibility of allele-specific RNA-seq analysis.
- Asia Mendelevich
- , Svetlana Vinogradova
- & Alexander A. Gimelbrant
-
Article
| Open Access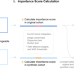 Identification of putative causal loci in whole-genome sequencing data via knockoff statistics
Identification of putative causal loci in whole-genome sequencing data via knockoff statistics
Association analyses that capture rare and noncoding variants in whole genome sequencing data are limited by factors like statistical power. Here, the authors present KnockoffScreen, a statistical method using the knockoff framework to detect, localise and prioritise rare and common risk variants at genome-wide scale.
- Zihuai He
- , Linxi Liu
- & Iuliana Ionita-Laza
-
Article
| Open Access Mining mutation contexts across the cancer genome to map tumor site of origin
Mining mutation contexts across the cancer genome to map tumor site of origin
The vast majority of somatic mutations observed in tumors are rare. Here, the authors show that these large numbers of rare mutations are more predictive of the tissue of origin of a tumor than the information from a few common driver mutations.
- Saptarshi Chakraborty
- , Axel Martin
- & Ronglai Shen
-
Article
| Open Access Detecting and phasing minor single-nucleotide variants from long-read sequencing data
Detecting and phasing minor single-nucleotide variants from long-read sequencing data
Cellular genetic heterogeneity is common across biological conditions, yet application of long-read sequencing to this subject is limited by error rates. Here, the authors present iGDA, a tool for detection and phasing of minor variants from long-read sequencing data, allowing accurate reconstruction of haplotypes.
- Zhixing Feng
- , Jose C. Clemente
- & Eric E. Schadt
-
Article
| Open Access Permutation-based identification of important biomarkers for complex diseases via machine learning models
Permutation-based identification of important biomarkers for complex diseases via machine learning models
Study of human disease remains challenging due to convoluted disease etiologies and complex molecular mechanisms at genetic, genomic, and proteomic levels. Here, the authors propose a computationally efficient Permutation-based Feature Importance Test to assist interpretation and selection of individual features in complex machine learning models for complex disease analysis.
- Xinlei Mi
- , Baiming Zou
- & Jianhua Hu
-
Article
| Open Access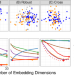 Supervised dimensionality reduction for big data
Supervised dimensionality reduction for big data
Biomedical measurements usually generate high-dimensional data where individual samples are classified in several categories. Vogelstein et al. propose a supervised dimensionality reduction method which estimates the low-dimensional data projection for classification and prediction in big datasets.
- Joshua T. Vogelstein
- , Eric W. Bridgeford
- & Mauro Maggioni
-
Article
| Open Access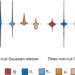 Total genetic contribution assessment across the human genome
Total genetic contribution assessment across the human genome
Quantifying the effects of individual loci on the human phenome is a challenging task. Here, the authors introduce a modelling technique, TGCA, that assesses total genetic contribution per locus and apply this to UK Biobank phenotype domains, revealing top loci and links to tissue-specific gene expression.
- Ting Li
- , Zheng Ning
- & Xia Shen
-
Article
| Open Access Landscape of allele-specific transcription factor binding in the human genome
Landscape of allele-specific transcription factor binding in the human genome
Single-nucleotide variants in enhancers or promoters may affect gene transcription by altering transcription factor binding sites. Here the authors present a meta-analysis empowered by a new statistical method covering thousands of ChIP-Seq experiments resulting in the identification of more than 500 thousand allele-specific binding (ASB) events in the human genome.
- Sergey Abramov
- , Alexandr Boytsov
- & Ivan V. Kulakovskiy
-
Article
| Open Access Estimating COVID-19 mortality in Italy early in the COVID-19 pandemic
Estimating COVID-19 mortality in Italy early in the COVID-19 pandemic
Estimates of COVID-19-related mortality are limited by incomplete testing. Here, the authors perform counterfactual analyses and estimate that there were 59,000–62,000 deaths from COVID-19 in Italy until 9th September 2020, approximately 1.5 times higher than official statistics.
- Chirag Modi
- , Vanessa Böhm
- & Uroš Seljak
-
Article
| Open Access Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Tissue damage and turnover lead to the release of DNA in the blood and can be used to monitor changes in tissue state. Here, the authors developed a tool to accurately estimate the proportion of cell types contributing to cell-free DNA in the blood, with an application to pregnant women and ALS patients.
- Christa Caggiano
- , Barbara Celona
- & Noah Zaitlen
-
Article
| Open Access Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Few genome-wide association studies have explored the genetic architecture of age-of-onset for traits and diseases. Here, the authors develop a Bayesian approach to improve prediction in timing-related phenotypes and perform age-of-onset analyses across complex traits in the UK Biobank.
- Sven E. Ojavee
- , Athanasios Kousathanas
- & Matthew R. Robinson
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access RA3 is a reference-guided approach for epigenetic characterization of single cells
RA3 is a reference-guided approach for epigenetic characterization of single cells
Methods for profiling differences between individual cells are constantly expanding. Here, the authors present a computational framework for the analysis of chromatin accessibility data at the single-cell level that takes into account previous knowledge and data-specific characteristics.
- Shengquan Chen
- , Guanao Yan
- & Zhixiang Lin
-
Article
| Open Access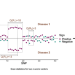 Detecting local genetic correlations with scan statistics
Detecting local genetic correlations with scan statistics
Genetic correlation analyses give insight on complex disease, yet are limited by oversimplification. Here, the authors present LOGODetect, a method using summary statistics from genome-wide association studies to identify genomic regions with correlation signals across multiple phenotypes.
- Hanmin Guo
- , James J. Li
- & Lin Hou

 Predictive models for the selection of thermally tolerant corals based on offspring survival
Predictive models for the selection of thermally tolerant corals based on offspring survival