Article
|
Open Access
Featured
-
-
Article
| Open Access Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Ribosome biogenesis is tightly coordinated with cell-cycle progression. By characterizing the SUMO isopeptidases SENP3/SENP5, Doenig et al. identify a long-sought p53-independent impaired ribosome checkpoint that converges on downregulation of CDK6.
- Judith Dönig
- , Hannah Mende
- & Stefan Müller
-
Article
| Open Access Molecular basis for recognition and deubiquitination of 40S ribosomes by Otu2
Molecular basis for recognition and deubiquitination of 40S ribosomes by Otu2
Otu2-driven deubiquitylation of ribosomal protein eS7 impacts translational efficiency. Here, the authors provide the molecular basis for recognition of monoubiquitinated eS7 on 40S and give mechanistic insights into Otu2’s role in translation reset.
- Ken Ikeuchi
- , Nives Ivic
- & Roland Beckmann
-
Article
| Open Access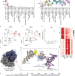 RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L is a paralog of the ribosomal protein RPL3 and specifically expressed in heart and skeletal muscle. Here, the authors show that RPL3L-containing ribosomes regulate translation elongation dynamics especially for the transcripts related to cardiac muscle contraction.
- Chisa Shiraishi
- , Akinobu Matsumoto
- & Keiichi I. Nakayama
-
Article
| Open Access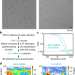 The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Developments in cryo-EM sample preparation and data collection are pivotal for structure determination. Fromm et al. present a 1.55 Å structure of the translating bacterial ribosome that provides new insights on its function and may allow for more precise structure-based drug design.
- Simon A. Fromm
- , Kate M. O’Connor
- & Simone Mattei
-
Article
| Open Access Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Here the authors find that nascent chains enriched in acidic amino acids destabilize the translating ribosome, eventually leading to stochastic premature termination in eukaryotes as in bacteria. Such risk of premature termination influences the amino acid distribution in the proteomes.
- Yosuke Ito
- , Yuhei Chadani
- & Hideki Taguchi
-
Article
| Open Access Modulating co-translational protein folding by rational design and ribosome engineering
Modulating co-translational protein folding by rational design and ribosome engineering
The narrow exit tunnel of the ribosome is important for cotranslational protein folding. Here, authors show that their rationally designed and engineered exit tunnel protein loops modulate the free energy of nascent chain dynamics and folding.
- Minkoo Ahn
- , Tomasz Włodarski
- & John Christodoulou
-
Article
| Open Access Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Initiation factor 2 (IF2) guides the ribosome to the elongation phase of protein synthesis. Here, Basu et al. provide structural insights into how compact IF2-GDP makes way for initiator tRNA accommodation into the peptidyl (P) site of the ribosome.
- Ritwika S. Basu
- , Michael B. Sherman
- & Matthieu G. Gagnon
-
Article
| Open Access Structural basis for late maturation steps of the human mitoribosomal large subunit
Structural basis for late maturation steps of the human mitoribosomal large subunit
Mitochondrial ribosomes (mitoribosomes) are characterized by a distinct architecture and thus biogenesis pathway. Here, cryo-EM structures of mitoribosome large subunit assembly intermediates elucidate final steps of 16 S rRNA folding, methylation and peptidyl transferase centre (PTC) completion, as well as functions of several mitoribosome assembly factors.
- Miriam Cipullo
- , Genís Valentín Gesé
- & Joanna Rorbach
-
Article
| Open Access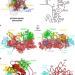 Structural basis of GTPase-mediated mitochondrial ribosome biogenesis and recycling
Structural basis of GTPase-mediated mitochondrial ribosome biogenesis and recycling
Maturation of the ribosomal peptidyl transferase center (PTC) is mediated by universally conserved GTPases. Here, cryo-EM structures of mitochondrial ribosomal large subunit assembly intermediates and of mature ribosomes offer insight into the roles of several assembly factors, including GTPBP6’s role in both ribosome biogenesis and recycling.
- Hauke S. Hillen
- , Elena Lavdovskaia
- & Ricarda Richter-Dennerlein
-
Article
| Open Access Impaired eIF5A function causes a Mendelian disorder that is partially rescued in model systems by spermidine
Impaired eIF5A function causes a Mendelian disorder that is partially rescued in model systems by spermidine
eIF5A is critical for protein synthesis but has not yet been associated with congenital human disease. Here, the authors show that EIF5A variants cause a Mendelian disorder via reduced eIF5A-ribosome interactions and this phenotype is partially corrected by spermidine supplementation in yeast and zebrafish.
- Víctor Faundes
- , Martin D. Jennings
- & Siddharth Banka
-
Article
| Open Access Ribosome-bound Get4/5 facilitates the capture of tail-anchored proteins by Sgt2 in yeast
Ribosome-bound Get4/5 facilitates the capture of tail-anchored proteins by Sgt2 in yeast
The guided entry of tail-anchored proteins (GET) pathway assists in the delivery of such proteins to the ER. Here, the authors reveal that the pathway components Get4/5 probe a region near the ribosomal exit tunnel. Upon emergence of a client protein, Get4/5 recruits Sgt2 and initiates the targeting phase of the pathway.
- Ying Zhang
- , Evelina De Laurentiis
- & Sabine Rospert
-
Article
| Open Access RETRACTED ARTICLE:The Arabidopsis NOT4A E3 ligase promotes PGR3 expression and regulates chloroplast translation
RETRACTED ARTICLE:The Arabidopsis NOT4A E3 ligase promotes PGR3 expression and regulates chloroplast translation
NOT4 proteins associate with ribosomes and are required for co-translational quality control in yeast and animals. Here, Bailey et al. show that Arabidopsis NOT4A positively regulates the expression of the nuclear encoded pentatricopeptide repeat (PPR) protein PGR3 and is required for ribosome biogenesis and mRNA translation in the chloroplast.
- Mark Bailey
- , Aiste Ivanauskaite
- & Daniel J. Gibbs
-
Article
| Open Access ArfB can displace mRNA to rescue stalled ribosomes
ArfB can displace mRNA to rescue stalled ribosomes
Alternative rescue factor B (ArfB) is an enzyme that releases peptides from stalled ribosomes to allow ribosome recycling. Here the authors carry-out cryo-EM analyses of 70S ribosomes complexed with ArfB on either a short or longer mRNA to reveal distinct modes of ArfB function.
- Christine E. Carbone
- , Gabriel Demo
- & Andrei A. Korostelev
-
Article
| Open Access Structural insights into assembly of the ribosomal nascent polypeptide exit tunnel
Structural insights into assembly of the ribosomal nascent polypeptide exit tunnel
The nascent polypeptide exit tunnel (NPET) is a functional center of the large ribosomal subunit through which the nascent polypeptide chains travel from the peptidyltransferase center (PTC). Here the authors provide structural insight into NPET maturation and how it is linked to other aspects of ribosome biogenesis.
- Daniel M. Wilson
- , Yu Li
- & John L. Woolford Jr
-
Article
| Open Access Mechanism of ribosome shutdown by RsfS in Staphylococcus aureus revealed by integrative structural biology approach
Mechanism of ribosome shutdown by RsfS in Staphylococcus aureus revealed by integrative structural biology approach
Upon transition to stationary phase or upon stress, bacteria limit protein synthesis through small inhibitory proteins that bind the ribosome. Here the authors decipher the interaction mode of the bacterial ribosome silencing factor (RsfS) at atomic details to provide an in depth view of how it shutdowns ribosomes.
- Iskander Khusainov
- , Bulat Fatkhullin
- & Marat Yusupov
-
Article
| Open Access Conserved phosphorylation hotspots in eukaryotic protein domain families
Conserved phosphorylation hotspots in eukaryotic protein domain families
Protein phosphorylation has various regulatory functions. Here, the authors map 241 phosphorylation hotspot regions across 40 eukaryotic species, showing that they are enriched at interfaces and near catalytic residues, and enable the discovery of functionally important phospho-sites.
- Marta J. Strumillo
- , Michaela Oplová
- & Pedro Beltrao
-
Article
| Open Access Suppressor mutations in Rpf2–Rrs1 or Rpl5 bypass the Cgr1 function for pre-ribosomal 5S RNP-rotation
Suppressor mutations in Rpf2–Rrs1 or Rpl5 bypass the Cgr1 function for pre-ribosomal 5S RNP-rotation
During biogenesis of the eukaryotic 60S ribosome, a large rotational movement of the 5S RNP is required to achieve its mature position. By analyzing extragenic suppressors of crg1—a key factor required for rotation—the authors provide mechanistic insight into a key step of ribosome biogenesis.
- Matthias Thoms
- , Valentin Mitterer
- & Ed Hurt
-
Article
| Open Access Molecular basis for disassembly of an importin:ribosomal protein complex by the escortin Tsr2
Molecular basis for disassembly of an importin:ribosomal protein complex by the escortin Tsr2
Ribosomal proteins are transported to the nucleus with the help of importins, from which they are released prior to incorporation into the nascent ribosome. Here the authors report the NMR structure of the ribosomal protein eS26 in complex with the escortin Tsr2 and shed light on the mechanism of eS26 release from importin.
- Sabina Schütz
- , Erich Michel
- & Vikram Govind Panse
-
Article
| Open Access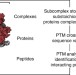 Dissecting ribosomal particles throughout the kingdoms of life using advanced hybrid mass spectrometry methods
Dissecting ribosomal particles throughout the kingdoms of life using advanced hybrid mass spectrometry methods
The authors demonstrate a 3-tier mass spectrometry approach, including bottom-up and top-down proteomics, as well as native mass spectrometry to provide a detailed description of proteoforms, protein processing and post-translational modifications present within ribosomes from bacteria, plant, and human.
- Michiel van de Waterbeemd
- , Sem Tamara
- & Albert J. R. Heck
-
Article
| Open Access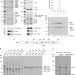 Interdependent action of KH domain proteins Krr1 and Dim2 drive the 40S platform assembly
Interdependent action of KH domain proteins Krr1 and Dim2 drive the 40S platform assembly
The biogenesis of eukaryotic ribosomes involves the coordinated interplay of a large number of assembly factors. Here, the authors detail how the conserved KH domain-containing assembly factors Dim2 and Krr1 function in ordering key events during ribosome maturation.
- Miriam Sturm
- , Jingdong Cheng
- & Ed Hurt
-
Article
| Open Access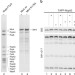 Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Reconstitution of the complete pathway of ITS2 processing at the pre-ribosome
Excision of internal transcribed spacer 2 (ITS2) within eukaryotic pre-ribosomal RNA is essential for ribosome function. Here, the authors reconstitute the entire cycle of ITS2 processing in vitro using purified components, providing insights into the cleavage process and demonstrating that 26S pre-rRNA processing necessarily precedes 7S pre-rRNA processing.
- Lisa Fromm
- , Sebastian Falk
- & Ed Hurt
-
Article
| Open Access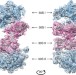 The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
Under conditions of nutrient limitation, bacterial ribosomes undergo dimerization, forming a 100S complex that is translationally inactive. Here the authors present the structural basis for formation of the 100S complexes in Gram-positive bacteria, shedding light on the mechanism of translation suppression by the ribosome-silencing factors.
- Donna Matzov
- , Shintaro Aibara
- & Ada E. Yonath
-
Article
| Open Access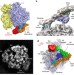 Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Membrane proteins are inserted co-transnationally through the association between ribosome, the signal recognition particle and its receptor, and the membrane-bound translocon. Here the authors present a cryo-EM reconstruction of this quaternary complex in the activated state and propose a model for signal sequence transfer to the translocon.
- Ahmad Jomaa
- , Yu-Hsien Hwang Fu
- & Nenad Ban
-
Article
| Open Access Molecular basis for protection of ribosomal protein L4 from cellular degradation
Molecular basis for protection of ribosomal protein L4 from cellular degradation
Acl4 is a dedicated assembly chaperone for ribosomal protein RpL4 that recognizes RpL4 in the cytoplasm to facilitate its nuclear import. Here the authors reveal the mechanism whereby Acl4 recognizes RpL4 and functions to protect it from Tom1-mediated degradation until RpL4 incorporation into the maturing 60S pre-ribosomal subunit.
- Ferdinand M. Huber
- & André Hoelz
-
Article
| Open Access Nop9 is a PUF-like protein that prevents premature cleavage to correctly process pre-18S rRNA
Nop9 is a PUF-like protein that prevents premature cleavage to correctly process pre-18S rRNA
Nop9 is a conserved small ribosomal subunit biogenesis factor. Here, Zhang et al. show that Nop9, in complex with RNA, adopts a C-shaped fold formed from 11 Pumillo repeats and propose that Nop9 inhibits premature cleavage of 20S pre-rRNA by inhibiting the Nob1 nuclease.
- Jun Zhang
- , Kathleen L. McCann
- & Traci M. Tanaka Hall
-
Article
| Open Access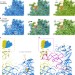 Structure–function insights reveal the human ribosome as a cancer target for antibiotics
Structure–function insights reveal the human ribosome as a cancer target for antibiotics
The ribosome of bacteria and other unicellular pathogens is a common target for antibiotic drugs. Here the authors determine a structure of the human ribosome bound to the translation inhibitor cycloheximide, and provide evidence that targeting the ribosome is a promising avenue for cancer therapy.
- Alexander G. Myasnikov
- , S. Kundhavai Natchiar
- & Bruno P. Klaholz
-
Article
| Open Access Involvement of human ribosomal proteins in nucleolar structure and p53-dependent nucleolar stress
Involvement of human ribosomal proteins in nucleolar structure and p53-dependent nucleolar stress
The nucleolus is a specialized functional domain of the nucleus where ribosome biogenesis is initiated and also implicated in a p53-dependent anti-tumor surveillance. Here the authors use a quantitative imaging approach to detail the role of each ribosomal protein on the structural integrity of the nucleolus and p53 homeostasis.
- Emilien Nicolas
- , Pascaline Parisot
- & Denis L. J. Lafontaine
-
Article
| Open Access Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Tsr1 is an essential ribosome biogenesis factor that has known similarity to GTPases. Here, the authors report the Tsr1 crystal structure and show that it is similar to GTPases but that active site residues are not conserved; modelling of the structure into the pre-40S maps allows inferences on ribosomal maturation to be drawn.
- Urszula M. McCaughan
- , Uma Jayachandran
- & Atlanta G. Cook
-
Article
| Open Access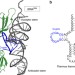 Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Ribosomes tend to stall during the translation of consecutive proline residues, which can be rescued by the co-translational factor EF-P. Here the authors identify a structural element of tRNAProresponsible for specific recognition by EF-P and stimulation of Pro-Pro peptide bond formation.
- Takayuki Katoh
- , Ingo Wohlgemuth
- & Hiroaki Suga
-
Article
| Open Access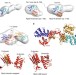 Sequential domain assembly of ribosomal protein S3 drives 40S subunit maturation
Sequential domain assembly of ribosomal protein S3 drives 40S subunit maturation
Ribosome biogenesis involves the hierarchical assembly of several proteins and RNA components. Here the authors describe a mechanism for ribosomal protein S3 incorporation into 40S ribosomal subunits that involves S3 dimerization and stepwise incorporation of two distinct S3 interaction domains coupled to release of ribosomal maturation factors.
- Valentin Mitterer
- , Guillaume Murat
- & Brigitte Pertschy
-
Article
| Open Access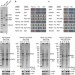 Co-translational capturing of nascent ribosomal proteins by their dedicated chaperones
Co-translational capturing of nascent ribosomal proteins by their dedicated chaperones
The synthesis of ribosomes requires the orderly assembly of many proteins and large RNA molecules, a process that involves several assembly factors. Here the authors show that dedicated chaperones capture the N termini of specific nascent ribosomal proteins to promote folding and assembly into maturing ribosomes.
- Patrick Pausch
- , Ujjwala Singh
- & Dieter Kressler
-
Article
| Open Access Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Insights into the origin of the nuclear localization signals in conserved ribosomal proteins
Eukaryotic ribosomal proteins contain nuclear localization signals (NLSs) that their bacterial counterparts lack. Here the authors compare homologous proteins from bacterial and eukaryotic ribosomes to show how NLSs could emerge in the course of evolution, and use this knowledge to identify novel NLSs.
- Sergey Melnikov
- , Adam Ben-Shem
- & Marat Yusupov
-
Article |
 Organization of the mitochondrial translation machinery studied in situ by cryoelectron tomography
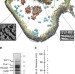
Organization of the mitochondrial translation machinery studied in situ by cryoelectron tomography
While the cytosolic translation machinery is well characterized, the mitochondrial translation system remains largely elusive. Using cryo-electron tomography, Pfeffer et al. describe the ordered organization of mitochondrial polysomes in which each ribosome is tethered to the inner membrane by two defined contacts on the large subunit in situ.
- Stefan Pfeffer
- , Michael W. Woellhaf
- & Friedrich Förster
-
Article |
 Ribosomal protein S1 functions as a termination factor in RNA synthesis by Qβ phage replicase
Ribosomal protein S1 functions as a termination factor in RNA synthesis by Qβ phage replicase
Protein S1, a subunit of the Qß phage RNA-directed RNA polymerase, was thought to only initiate copying of the phage RNA plus strand. Here, the authors show that S1 stimulates replication of any cognate template by promoting release of the newly synthesized product strand.
- Nikita N. Vasilyev
- , Zarina S. Kutlubaeva
- & Alexander B. Chetverin

 Viscosity-dependent control of protein synthesis and degradation
Viscosity-dependent control of protein synthesis and degradation