Article
|
Open Access
Featured
-
-
Article
| Open Access Machine learning-driven multifunctional peptide engineering for sustained ocular drug delivery
Machine learning-driven multifunctional peptide engineering for sustained ocular drug delivery
Sustained drug delivery is critical for patient adherence to chronic disease treatments. Here the authors apply machine learning to engineer multifunctional peptides with high melanin binding, high cell-penetration, and low cytotoxicity, enhancing the duration and efficacy of peptide-drug conjugates for sustained ocular delivery.
- Henry T. Hsueh
- , Renee Ti Chou
- & Laura M. Ensign
-
Article
| Open Access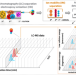 PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
Alternative algorithms exploiting advantages of multidimensional mass spectrometry in untargeted metabolomics are needed. Here, the authors develop and demonstrate PeakDecoder for confident and accurate metabolite profiling in 116 microbial sample runs and using a library built from 64 standards.
- Aivett Bilbao
- , Nathalie Munoz
- & Kristin E. Burnum-Johnson
-
Article
| Open Access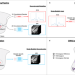 Cross-modal autoencoder framework learns holistic representations of cardiovascular state
Cross-modal autoencoder framework learns holistic representations of cardiovascular state
A challenge in diagnostics is integrating different data modalities to characterize physiological state. Here, the authors show, using the heart as a model system, that cross-modal autoencoders can integrate and translate modalities to improve diagnostics and identify associated genetic variants.
- Adityanarayanan Radhakrishnan
- , Sam F. Friedman
- & Caroline Uhler
-
Article
| Open Access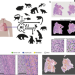 Bridging clinic and wildlife care with AI-powered pan-species computational pathology
Bridging clinic and wildlife care with AI-powered pan-species computational pathology
Artificial Intelligence (AI) has the potential of assisting the study and diagnosis of veterinary cancers. Here, the authors build a cancer digital pathology atlas encompassing multiple animal species and demonstrate an AI approach for comparative pathology, which yields insights about immune response and morphological similarities.
- Khalid AbdulJabbar
- , Simon P. Castillo
- & Yinyin Yuan
-
Article
| Open Access A machine learning model identifies patients in need of autoimmune disease testing using electronic health records
A machine learning model identifies patients in need of autoimmune disease testing using electronic health records
Early diagnosis can significantly improve treatment options and prevent severe organ damage in individuals with autoimmune diseases. Here, the authors develop a machine learning model that uses electronic health records to identify patients with clinical suspicion of autoimmune diseases.
- Iain S. Forrest
- , Ben O. Petrazzini
- & Ron Do
-
Article
| Open Access Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Prediction of antibody structures is critical for understanding and designing novel therapeutic and diagnostic molecules. Here, the authors present IgFold: a fast, accurate method for antibody structure prediction using an end-to-end deep learning model.
- Jeffrey A. Ruffolo
- , Lee-Shin Chu
- & Jeffrey J. Gray
-
Article
| Open Access Teasing out missing reactions in genome-scale metabolic networks through hypergraph learning
Teasing out missing reactions in genome-scale metabolic networks through hypergraph learning
A computational method for rapid and accurate gap-filling of metabolic networks without using phenotypic data is unavailable. Here, the authors address this problem by developing a deep learning based method that can predict missing reactions using topological features of the metabolic networks.
- Can Chen
- , Chen Liao
- & Yang-Yu Liu
-
Article
| Open Access Sample-to-answer platform for the clinical evaluation of COVID-19 using a deep learning-assisted smartphone-based assay
Sample-to-answer platform for the clinical evaluation of COVID-19 using a deep learning-assisted smartphone-based assay
The lateral flow assay (LFA) has been considered a rapid test tool but with low sensitivity hampering the precise diagnosis. Here, the authors report bioengineered enrichment tools for LFAs with enhanced sensitivity and specificity that can reinforce LFA’s clinical performance.
- Seungmin Lee
- , Sunmok Kim
- & Jeong Hoon Lee
-
Article
| Open Access Latent generative landscapes as maps of functional diversity in protein sequence space
Latent generative landscapes as maps of functional diversity in protein sequence space
In this work, the authors study protein families’ VAE latent manifolds and coevolutionary Hamiltonians. These Latent Generative Landscapes predict phylogenetic groupings, fitness & functional properties for several systems with clear protein engineering/design potential.
- Cheyenne Ziegler
- , Jonathan Martin
- & Faruck Morcos
-
Article
| Open Access DeMAG predicts the effects of variants in clinically actionable genes by integrating structural and evolutionary epistatic features
DeMAG predicts the effects of variants in clinically actionable genes by integrating structural and evolutionary epistatic features
Interpretation of rare genetic variants remains challenging. Here, the authors develop a supervised variant effect predictor for use in clinically actionable genes which incorporates evolutionary and structural relationships between residues and has balanced specificity and sensitivity.
- Federica Luppino
- , Ivan A. Adzhubei
- & Agnes Toth-Petroczy
-
Article
| Open Access Histopathology images predict multi-omics aberrations and prognoses in colorectal cancer patients
Histopathology images predict multi-omics aberrations and prognoses in colorectal cancer patients
Histopathological analysis is an essential tool in diagnosing colorectal cancer, but is limited in predicting prognosis and molecular profiles. Here, the authors designed a machine learning-based platform to predict multi-omics profiles and prognosis from pathology images.
- Pei-Chen Tsai
- , Tsung-Hua Lee
- & Kun-Hsing Yu
-
Article
| Open Access Predictive and robust gene selection for spatial transcriptomics
Predictive and robust gene selection for spatial transcriptomics
Gene selection for spatial transcriptomics is currently not optimal. Here the authors report PERSIST, a flexible deep learning framework that uses existing scRNA-seq data to identify gene targets for spatial transcriptomics; they show this allows you to capture more information with fewer genes.
- Ian Covert
- , Rohan Gala
- & Su-In Lee
-
Article
| Open Access Integrative modeling of tumor genomes and epigenomes for enhanced cancer diagnosis by cell-free DNA
Integrative modeling of tumor genomes and epigenomes for enhanced cancer diagnosis by cell-free DNA
Despite advances in ctDNA cancer detection, early detection remains difficult. Here, the authors utilise whole genome sequencing of 2,125 patient samples to create a model for early cancer and tissue of origin detection.
- Mingyun Bae
- , Gyuhee Kim
- & Jung Kyoon Choi
-
Article
| Open Access Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Prostate cancer is known to have a variable response to androgen receptor signalling inhibitors. Here, the authors use machine learning to predict response to therapy from genomic, transcriptomic and clinical data.
- Anouk C. de Jong
- , Alexandra Danyi
- & Martijn P. Lolkema
-
Article
| Open Access Improving the generalizability of protein-ligand binding predictions with AI-Bind
Improving the generalizability of protein-ligand binding predictions with AI-Bind
State-of-the-art machine learning models in drug discovery fail to reliably predict the binding properties of poorly annotated proteins and small molecules. Here, the authors present AI-Bind, a machine learning pipeline to improve generalizability and interpretability of binding predictions.
- Ayan Chatterjee
- , Robin Walters
- & Giulia Menichetti
-
Article
| Open Access Predicting compound activity from phenotypic profiles and chemical structures
Predicting compound activity from phenotypic profiles and chemical structures
Experimental assays are used to determine if compounds cause a desired activity in cells. Here the authors demonstrate that computational methods can predict compound bioactivity given their chemical structure, imaging and gene expression data from historic screening libraries.
- Nikita Moshkov
- , Tim Becker
- & Juan C. Caicedo
-
Article
| Open Access Cellcano: supervised cell type identification for single cell ATAC-seq data
Cellcano: supervised cell type identification for single cell ATAC-seq data
Accurately annotating cell types is a fundamental step in single-cell omics data analysis. Here, the authors develop a computational method called Cellcano based on a two-round supervised learning algorithm to identify cell types for scATAC-seq data and perform benchmarking to demonstrate its accuracy, robustness and computational efficiency.
- Wenjing Ma
- , Jiaying Lu
- & Hao Wu
-
Article
| Open Access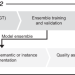 Deep learning-enabled segmentation of ambiguous bioimages with deepflash2
Deep learning-enabled segmentation of ambiguous bioimages with deepflash2
The signal-to-noise ratio in bioimages is often low, which is problematic for segmentation. Here the authors report a deep learning method, deepflash2, to facilitate the segmentation of ambiguous bioimages through multi-expert annotations and integrated quality assurance.
- Matthias Griebel
- , Dennis Segebarth
- & Christoph M. Flath
-
Article
| Open Access Interoperable slide microscopy viewer and annotation tool for imaging data science and computational pathology
Interoperable slide microscopy viewer and annotation tool for imaging data science and computational pathology
There is a lack of standardisation in slide microscopy imaging data. Here the authors report Slim, an open-source, web-based slide microscopy viewer implementing the Digital Imaging and Communications in Medicine (DICOM) standard to achieve interoperability with a range of existing medical imaging systems.
- Chris Gorman
- , Davide Punzo
- & Markus D. Herrmann
-
Article
| Open Access A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
This study comprehensively benchmarks 18 state-of-the-art methods for cellular deconvolution of spatial transcriptomics and provide decision-tree-style guidelines and recommendations for method selection.
- Haoyang Li
- , Juexiao Zhou
- & Xin Gao
-
Article
| Open Access Clinical and genetic associations of deep learning-derived cardiac magnetic resonance-based left ventricular mass
Clinical and genetic associations of deep learning-derived cardiac magnetic resonance-based left ventricular mass
A genome-wide association study of cardiac magnetic resonance-derived left ventricular mass index including 43,000 UK Biobank participants reveals 12 associations (11 novel), implicating genes involved in cardiac contractility and cardiomyopathy.
- Shaan Khurshid
- , Julieta Lazarte
- & Steven A. Lubitz
-
Article
| Open Access Discovering highly potent antimicrobial peptides with deep generative model HydrAMP
Discovering highly potent antimicrobial peptides with deep generative model HydrAMP
Antimicrobial peptides emerge as compounds that can alleviate the global health hazard of antimicrobial resistance. Here, the authors propose HydrAMP, an extended conditional variational autoencoder. HydrAMP generated antimicrobial peptides with high activity against bacteria, including multidrug-resistant species.
- Paulina Szymczak
- , Marcin Możejko
- & Ewa Szczurek
-
Article
| Open Access Interpretable and context-free deconvolution of multi-scale whole transcriptomic data with UniCell deconvolve
Interpretable and context-free deconvolution of multi-scale whole transcriptomic data with UniCell deconvolve
There is interest in measuring the influence of spatial cellular organization on pathophysiology, which is being accomplished through spatial transcriptomics. There the authors present UniCell Deconvolve, a pre-trained deep learning model that predicts cell identity and deconvolves cell type fractions using a 28 M cell database.
- Daniel Charytonowicz
- , Rachel Brody
- & Robert Sebra
-
Article
| Open Access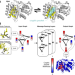 Predicting locations of cryptic pockets from single protein structures using the PocketMiner graph neural network
Predicting locations of cryptic pockets from single protein structures using the PocketMiner graph neural network
Cryptic pockets enable targeting of proteins currently considered undruggable because they lack pockets in their ground state structures. Here, the authors develop a graph neural network that accurately predicts cryptic pockets in static structures by training using molecular simulation data alone.
- Artur Meller
- , Michael Ward
- & Gregory R. Bowman
-
Article
| Open Access Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with GraphST
Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with GraphST
Advances in spatial transcriptomics technologies have enabled the gene expression profiling of tissues while retaining spatial context. Here the authors present GraphST, a graph self-supervised contrastive learning method that learns informative and discriminative spot representations from spatial transcriptomics data.
- Yahui Long
- , Kok Siong Ang
- & Jinmiao Chen
-
Article
| Open Access Hierarchical graph learning for protein–protein interaction
Hierarchical graph learning for protein–protein interaction
Despite recent progress, machine learning methods remain inadequate in modeling the natural protein-protein interaction (PPI) hierarchy for PPI prediction. Here, the authors present a double-viewed hierarchical graph learning model, HIGH-PPI, to predict PPIs and extrapolate the molecular details involved.
- Ziqi Gao
- , Chenran Jiang
- & Jia Li
-
Article
| Open Access Evaluating native-like structures of RNA-protein complexes through the deep learning method
Evaluating native-like structures of RNA-protein complexes through the deep learning method
RNA-protein docking is a very challenging area. Here, the authors develop a deep-learning based method, DRPScore, to evaluate RNA-protein complexes. DRPScore is robust and consistently performs better than existing methods on representative testing sets.
- Chengwei Zeng
- , Yiren Jian
- & Yunjie Zhao
-
Article
| Open Access Batch alignment of single-cell transcriptomics data using deep metric learning
Batch alignment of single-cell transcriptomics data using deep metric learning
The increasing scale of single-cell RNA-seq studies presents new challenge for integrating datasets from different batches. Here, the authors develop scDML, a tool that simultaneously removes batch effects, improves clustering performance, recovers true cell types, and scales well to large datasets.
- Xiaokang Yu
- , Xinyi Xu
- & Xiangjie Li
-
Article
| Open Access Single-cell biological network inference using a heterogeneous graph transformer
Single-cell biological network inference using a heterogeneous graph transformer
Single-cell multi-omics and deep learning could lead to the inference of biological networks across specific cell types. Here, the authors develop DeepMAPS, a deep learning, graph-based approach for cell-type specific network inference from single-cell multi-omics data that is tested on healthy and tumour tissue datasets.
- Anjun Ma
- , Xiaoying Wang
- & Qin Ma
-
Article
| Open Access Virtual elastography ultrasound via generative adversarial network for breast cancer diagnosis
Virtual elastography ultrasound via generative adversarial network for breast cancer diagnosis
The current use of elastography ultrasound faces challenges, including vulnerability to subjective manipulation, echo signal attenuation, unknown risks of elastic pressure and high imaging hardware cost. Here, the author shows a virtual elastography to empower low-end ultrasound devices with state-of-art elastography function.
- Zhao Yao
- , Ting Luo
- & JianQiao Zhou
-
Article
| Open Access Direct generation of protein conformational ensembles via machine learning
Direct generation of protein conformational ensembles via machine learning
Computational methods to study protein structural dynamics are a powerful tool in life sciences but are computationally expensive. Here, the authors show that machine learning can be used to efficiently generate protein conformational ensembles and test their method on intrinsically disordered peptides.
- Giacomo Janson
- , Gilberto Valdes-Garcia
- & Michael Feig
-
Article
| Open Access Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches
Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches
Application of CRISPR-Cas13d is limited by the inability to predict on- and off-targets. Here the authors perform CRISPR-Cas13d proliferation screens followed by modeling of Cas13d on- and off-targets; they design a deep learning model, DeepCas13, to predict the on-target activity of a gRNA.
- Xiaolong Cheng
- , Zexu Li
- & Wei Li
-
Article
| Open Access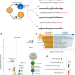 Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia
Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia
The PML-RARA gene fusion is the characteristic driver of Acute Promyelocytic Leukaemia (APL) and is known to bind to the genome. Here, the authors characterise the impact of PML-RARA on gene regulation in APL cell lines and patient samples using transcriptomics, epigenomics, and machine learning.
- William Villiers
- , Audrey Kelly
- & Cameron S. Osborne
-
Article
| Open Access Mapping lesion-specific response and progression dynamics and inter-organ variability in metastatic colorectal cancer
Mapping lesion-specific response and progression dynamics and inter-organ variability in metastatic colorectal cancer
Understanding the heterogeneity of growth, response to therapy and progression dynamics in metastatic colorectal cancer (mCRC) remains critical. Here, the authors analyse lesion-specific response heterogeneity in 4,308 mCRC patients and find that organ-level progression sequence is associated with long-term survival.
- Jiawei Zhou
- , Amber Cipriani
- & Yanguang Cao
-
Article
| Open Access Decision level integration of unimodal and multimodal single cell data with scTriangulate
Decision level integration of unimodal and multimodal single cell data with scTriangulate
Single-cell genomics has expanded to measure diverse molecular modalities within the same cell. Here the authors provide a computational framework called scTriangulate to integrate cluster annotations from diverse independent sources, algorithms, and modalities to define statistically stable populations.
- Guangyuan Li
- , Baobao Song
- & Nathan Salomonis
-
Article
| Open Access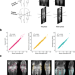 BMI-adjusted adipose tissue volumes exhibit depot-specific and divergent associations with cardiometabolic diseases
BMI-adjusted adipose tissue volumes exhibit depot-specific and divergent associations with cardiometabolic diseases
Different location of adipose tissue may have different consequences to cardiometabolic risk. Here the authors report that deep learning enabled accurate prediction of specific adipose tissue volumes, and that after adjustment for BMI, visceral adiposity was associated with increased risk of cardiometabolic disease, while gluteofemoral adiposity was associated with reduced risk.
- Saaket Agrawal
- , Marcus D. R. Klarqvist
- & Amit V. Khera
-
Article
| Open Access Transformer for one stop interpretable cell type annotation
Transformer for one stop interpretable cell type annotation
Developing computational tools for interpretable cell type annotation in scRNA-seq data remains challenging. Here the authors propose a Transformer-based model for interpretable annotation transfer using biologically understandable entities, and demonstrate its performance on large or atlas datasets.
- Jiawei Chen
- , Hao Xu
- & Jing-Dong J. Han
-
Article
| Open Access Prediction of designer-recombinases for DNA editing with generative deep learning
Prediction of designer-recombinases for DNA editing with generative deep learning
Design of recombinases with new target sites is usually achieved through cycles of directed molecular evolution. Here the authors report Recombinase Generator, RecGen, an algorithm for generation of designer-recombinases; they perform experimental validation to show that this can predict recombinase sequences.
- Lukas Theo Schmitt
- , Maciej Paszkowski-Rogacz
- & Frank Buchholz
-
Article
| Open Access Accuracy and data efficiency in deep learning models of protein expression
Accuracy and data efficiency in deep learning models of protein expression
Synthetic biology often involves engineering microbial strains to express high-value proteins. Here the authors build deep learning predictors of protein expression from sequence that deliver accurate models with fewer data than previously assumed, helping to lower costs of model-driven strain design.
- Evangelos-Marios Nikolados
- , Arin Wongprommoon
- & Diego A. Oyarzún
-
Article
| Open Access Estimating diagnostic uncertainty in artificial intelligence assisted pathology using conformal prediction
Estimating diagnostic uncertainty in artificial intelligence assisted pathology using conformal prediction
Artificial intelligence prediction accuracy can be reduced with new data. Here, the authors utilise conformal prediction to reduce incorrect predictions in histopathological analysis of prostate cancer biopsies.
- Henrik Olsson
- , Kimmo Kartasalo
- & Martin Eklund
-
Article
| Open Access A unifying Bayesian framework for merging X-ray diffraction data
A unifying Bayesian framework for merging X-ray diffraction data
Observation of the chemical and conformational dynamics of biomolecules by diffraction methods is impeded by several physical artifacts. The authors present an extensible framework for accurate correction of such data that can keep pace with rapid developments in diffraction methods.
- Kevin M. Dalton
- , Jack B. Greisman
- & Doeke R. Hekstra
-
Article
| Open Access Clustering of single-cell multi-omics data with a multimodal deep learning method
Clustering of single-cell multi-omics data with a multimodal deep learning method
Single-cell multimodal sequencing technologies are developed to simultaneously profile different modalities of data in the same cell. Here the authors develops a multimodal deep clustering method for the analysis of single-cell multi-omics data that supports clustering different types of multi-omics data and multi-batch data, as well as downstream differential expression analysis.
- Xiang Lin
- , Tian Tian
- & Hakon Hakonarson
-
Article
| Open Access Tumor fractions deciphered from circulating cell-free DNA methylation for cancer early diagnosis
Tumor fractions deciphered from circulating cell-free DNA methylation for cancer early diagnosis
‘Circulating cell-free DNA can be used to predict cancer, but it is more challenging to assess in early stage cancer. Here, the authors created a diagnostic model using tumor fractions deciphered from circulating cfDNA methylation signatures, which exhibited an 86% sensitivity in detecting early-stage cancer.
- Xiao Zhou
- , Zhen Cheng
- & Weibin Cheng
-
Article
| Open Access Deep transfer learning enables lesion tracing of circulating tumor cells
Deep transfer learning enables lesion tracing of circulating tumor cells
Liquid biopsy offers great promise for noninvasive cancer diagnostics, while the lack of adequate target characterization and analysis hinders its wide application. Here, the authors design a transfer learning-based algorithm to transfer lesion labels from the primary cancer cell atlas to circulating tumor cells.
- Xiaoxu Guo
- , Fanghe Lin
- & Jia Song
-
Article
| Open Access An in silico method to assess antibody fragment polyreactivity
An in silico method to assess antibody fragment polyreactivity
Off-target binding hinders the development of therapeutic antibodies and reproducibility in basic research settings. Here the authors develop a method to quantify and reduce the polyreactivity of antibody fragments based on protein sequence alone.
- Edward P. Harvey
- , Jung-Eun Shin
- & Andrew C. Kruse
-
Article
| Open Access A framework for clinical cancer subtyping from nucleosome profiling of cell-free DNA
A framework for clinical cancer subtyping from nucleosome profiling of cell-free DNA
Nucleosome profiling from cell-free DNA (cfDNA) represents a potential approach for cancer detection and classification. Here, the authors develop Griffin, a computational framework for tumour subtype classification based on cfDNA nucleosome profiling that can work with ultra-low pass sequencing data.
- Anna-Lisa Doebley
- , Minjeong Ko
- & Gavin Ha
-
Article
| Open Access Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Methods for jointly analysing the different spatial data modalities in 3D are lacking. Here the authors report the computational framework STACI (Spatial Transcriptomic data using over-parameterized graph-based Autoencoders with Chromatin Imaging data) which they apply to an Alzheimer’s disease mouse model.
- Xinyi Zhang
- , Xiao Wang
- & Caroline Uhler
-
Article
| Open Access Reference panel guided topological structure annotation of Hi-C data
Reference panel guided topological structure annotation of Hi-C data
Predicting topological structures from Hi-C data provides insight into comprehending gene expression and regulation. Here, the authors present RefHiC, an attention-based deep learning framework that leverages a reference panel of Hi-C datasets to assist topological structure annotation from a given study sample.
- Yanlin Zhang
- & Mathieu Blanchette
-
Article
| Open Access A unified computational framework for single-cell data integration with optimal transport
A unified computational framework for single-cell data integration with optimal transport
Integrating heterogeneous single-cell multi-omics as well as spatially resolved transcriptomic data remains a major challenge. Here the authors report a unified single-cell data integration framework using an unbalanced optimal transport-based deep network.
- Kai Cao
- , Qiyu Gong
- & Lin Wan

 Explainable multi-task learning for multi-modality biological data analysis
Explainable multi-task learning for multi-modality biological data analysis