Featured
-
-
Article
| Open Access Structural basis of ribosomal RNA transcription regulation
Structural basis of ribosomal RNA transcription regulation
Ribosomal RNA (rRNA) expression is regulated at the initiation stage of RNA synthesis. Here, the authors report cryo-EM structures of E. coli RNA polymerase and rRNA promoter complex with DksA/ppGpp on the way to open complex formation, identifying key steps in promoter recognition and opening.
- Yeonoh Shin
- , M. Zuhaib Qayyum
- & Katsuhiko S. Murakami
-
Article
| Open Access Structural basis of RNA cap modification by SARS-CoV-2
Structural basis of RNA cap modification by SARS-CoV-2
Specific non-structural proteins (nsp) of SARS coronaviruses are involved in methylation of virally encoded mRNAs to mimic cellular mRNAs for protection against host innate immune restriction. Here, the authors present a high resolution structure of SARS-CoV-2 nsp16/nsp10 ternary complex in the presence of cognate RNA substrate analogue and methyl donor, S-adenosyl methionine, revealing unique ligand-binding sites that may represent alternative targets for antiviral development.
- Thiruselvam Viswanathan
- , Shailee Arya
- & Yogesh K. Gupta
-
Article
| Open Access Cryo-EM structure of a licensed DNA replication origin
Cryo-EM structure of a licensed DNA replication origin
Origins of replication are licensed by loading of MCM onto DNA, and origin firing depends on interaction with Cdc45 and GINS to form two CMG holo-helicases. Here, authors determine the cryo-EM structures of DNA-bound MCM and visualise a phospho-dependent MCM element important for Cdc45 recruitment.
- Ferdos Abid Ali
- , Max E. Douglas
- & Alessandro Costa
-
Article
| Open Access Structural basis of human PCNA sliding on DNA
Structural basis of human PCNA sliding on DNA
DNA sliding clamps are ring-shaped proteins that encircle DNA and harbour polymerases and other factors that promote processive DNA replication. Here the authors use X-ray crystallography, NMR and MD simulations to propose a model for a PCNA sliding mechanism that relies on short-lived polar interactions.
- Matteo De March
- , Nekane Merino
- & Alfredo De Biasio
-
Article
| Open Access Structure and substrate selectivity of the 750-kDa α6β6 holoenzyme of geranyl-CoA carboxylase
Structure and substrate selectivity of the 750-kDa α6β6 holoenzyme of geranyl-CoA carboxylase
The structures of the biotin-dependent carboxylases have revealed details of their function. Here, the authors describe the first structure of Pseudomonasgeranyl-CoA carboxylase, and compare it with the previously characterised homologous 3-methylcrotonyl-CoA carboxylase.
- Ashley R. Jurado
- , Christine S. Huang
- & Liang Tong
-
Article
| Open Access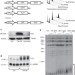 Proteomics of yeast telomerase identified Cdc48-Npl4-Ufd1 and Ufd4 as regulators of Est1 and telomere length
Proteomics of yeast telomerase identified Cdc48-Npl4-Ufd1 and Ufd4 as regulators of Est1 and telomere length
Regulating telomere length and telomerase activity are critical biological processes implicated in ageing and cancer. Here the authors use mass spectrometry to identify the Cdc48-Npl4-Ufd1 complex, which targets proteins for degradation, as a novel regulator of the yeast telomerase Est1.
- Kah-Wai Lin
- , Karin R. McDonald
- & Virginia A. Zakian

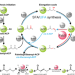 Helicobacter pylori FabX contains a [4Fe-4S] cluster essential for unsaturated fatty acid synthesis
Helicobacter pylori FabX contains a [4Fe-4S] cluster essential for unsaturated fatty acid synthesis