Featured
-
-
Article
| Open Access Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Normalisr removes technical bias in single-cell RNA-seq and detects gene differential and coexpression accurately and efficiently. It also infers gene regulatory and co-expression networks from conventional and CRISPR screen single-cell RNA-seq datasets.
- Lingfei Wang
-
Article
| Open Access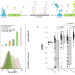 Improvement of a synthetic live bacterial therapeutic for phenylketonuria with biosensor-enabled enzyme engineering
Improvement of a synthetic live bacterial therapeutic for phenylketonuria with biosensor-enabled enzyme engineering
PKU patients have elevated phenylalanine levels which can result in neurological impairment. Here the authors utilize biosensor-based ultra-high-throughput screening to optimize PAL activity in a synthetic biotic platform for improved in vivo performance.
- Kristin J. Adolfsen
- , Isolde Callihan
- & Vincent M. Isabella
-
Article
| Open Access Engineering digitizer circuits for chemical and genetic screens in human cells
Engineering digitizer circuits for chemical and genetic screens in human cells
Cell-based transcriptional reporters are an invaluable part of highthroughput screening, but many such reporters have weak or transient responses. Here, the authors describe a digitizer circuit for amplifying reporter activity, increasing sensitivity, and retaining memory of pathway activation.
- Nicole M. Wong
- , Elizabeth Frias
- & Wilson W. Wong
-
Article
| Open Access GAK and PRKCD are positive regulators of PRKN-independent mitophagy
GAK and PRKCD are positive regulators of PRKN-independent mitophagy
The mechanisms involved in programmed or damage-induced removal of mitochondria by mitophagy remain elusive. Here the authors use an siRNA library to screen lipid-binding proteins, and identify the kinases GAK and PRKCD as positive regulators of PRKN-independent mitophagy.
- Michael J. Munson
- , Benan J. Mathai
- & Anne Simonsen
-
Article
| Open Access A microfluidic platform for highly parallel bite by bite profiling of mosquito-borne pathogen transmission
A microfluidic platform for highly parallel bite by bite profiling of mosquito-borne pathogen transmission
High-throughput molecular surveillance of mosquitoes carrying dangerous pathogens is currently challenging. Here the authors present Vectorchip, a low-cost microfluidic platform enabling multiplexed detection of mosquito DNA, viral RNA and infectious viral particles at single bite resolution.
- Shailabh Kumar
- , Felix J. H. Hol
- & Manu Prakash
-
Article
| Open Access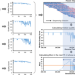 Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing
Current methods to identify G-quadruplex structures in DNA require specialized protocols and multiple rounds of sequencing. Here, the authors develop a method to detect G-quadruplex structures in DNA based on fluctuations in sequencing quality in a standard sequencing experiment.
- Jing Tu
- , Mengqin Duan
- & Zuhong Lu
-
Article
| Open Access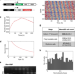 Circa-SCOPE: high-throughput live single-cell imaging method for analysis of circadian clock resetting
Circa-SCOPE: high-throughput live single-cell imaging method for analysis of circadian clock resetting
Phase Transition Curves (PTCs) describe phase shifts of circadian oscillations due to a stimulus and they are important for studying circadian clock resetting. Here, the authors present a method for high-throughput reconstruction of PTCs using fluorescent live imaging and single-cell analysis.
- Gal Manella
- , Dan Aizik
- & Gad Asher
-
Article
| Open Access High-throughput RNA sequencing of paraformaldehyde-fixed single cells
High-throughput RNA sequencing of paraformaldehyde-fixed single cells
Current high-throughput single-cell transcriptomic methods are incompatible with paraformaldehyde, a common cell fixation technique. Here the authors present FD-seq, a method for droplet-based RNA sequencing of paraformaldehyde-fixed, stained and sorted single cells.
- Hoang Van Phan
- , Michiel van Gent
- & Savaş Tay
-
Article
| Open Access Super-enhancer-based identification of a BATF3/IL-2R−module reveals vulnerabilities in anaplastic large cell lymphoma
Super-enhancer-based identification of a BATF3/IL-2R−module reveals vulnerabilities in anaplastic large cell lymphoma
Anaplastic large cell lymphoma (ALCL) is an aggressive T-cell lymphoma often with poor prognosis. To identify genes defining ALCL cell state and dependencies, the authors here characterize ALCL-specific super-enhancers and describe the BATF3/IL-2R−module as a therapeutic opportunity for ALCL.
- Huan-Chang Liang
- , Mariantonia Costanza
- & Olaf Merkel
-
Article
| Open Access Small-molecule suppression of calpastatin degradation reduces neuropathology in models of Huntington’s disease
Small-molecule suppression of calpastatin degradation reduces neuropathology in models of Huntington’s disease
Mitochondrial dysfunction is a common hallmark of neurological disorders. Here, the authors identify CHIR99021 as a potent enhancer of mitochondrial function, which improved mitochondrial phenotypes in Huntington’s disease models. CHIR99021 was shown to stabilize calpastatin, which suppressed calpain activation and Drp1-induced mitochondrial fragmentation.
- Di Hu
- , Xiaoyan Sun
- & Xin Qi
-
Article
| Open Access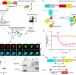 Integration of FRET and sequencing to engineer kinase biosensors from mammalian cell libraries
Integration of FRET and sequencing to engineer kinase biosensors from mammalian cell libraries
Existing Förster Resonance Energy Transfer (FRET) biosensors are often limited in their sensitivity. Here the authors report FRET-seq which they use to identify Fyn and ZAP70 kinase biosensors with enhanced performance, and use them to image T-cell activation and screen drugs.
- Longwei Liu
- , Praopim Limsakul
- & Yingxiao Wang
-
Article
| Open Access Genome-scale target identification in Escherichia coli for high-titer production of free fatty acids
Genome-scale target identification in Escherichia coli for high-titer production of free fatty acids
Identification of gene targets is one of the major challenges to construct superior microbial cell factory for chemical synthesis. Here, the authors employ CRISPRi and omics analyses for genome-scale target genes identification for high-titer production of free fatty acids in E. coli.
- Lixia Fang
- , Jie Fan
- & Hao Song
-
Article
| Open Access Integrative oncogene-dependency mapping identifies RIT1 vulnerabilities and synergies in lung cancer
Integrative oncogene-dependency mapping identifies RIT1 vulnerabilities and synergies in lung cancer
RIT1 mutations are mutually exclusive with other lung cancer drivers and lack targeted therapies. Here the authors examine genetic dependencies of mutant RIT1 with genome-wide CRISPR screens, revealing synergy between RIT1 and YAP1, and increased sensitivity to Aurora kinase inhibitors.
- Athea Vichas
- , Amanda K. Riley
- & Alice H. Berger
-
Article
| Open Access High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies
High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies
The engineering of 5′ UTRs that modulate protein expression remains a great challenge. Here we leverage synthetic biology and computational design to develop a high-throughput strategy to design, screen, and optimize 5′ UTRs that enhance protein expression for non-viral gene therapies.
- Jicong Cao
- , Eva Maria Novoa
- & Timothy K. Lu
-
Article
| Open Access High-resolution characterization of gene function using single-cell CRISPR tiling screen
High-resolution characterization of gene function using single-cell CRISPR tiling screen
Identifying functional domains and genetic regulatory mechanisms is essential for developing new therapies. Here the authors present sc-Tiling, single-cell high-density CRISPR tiling screening for functional domain characterization.
- Lu Yang
- , Anthony K. N. Chan
- & Chun-Wei Chen
-
Article
| Open Access A high-throughput screen for TMPRSS2 expression identifies FDA-approved compounds that can limit SARS-CoV-2 entry
A high-throughput screen for TMPRSS2 expression identifies FDA-approved compounds that can limit SARS-CoV-2 entry
The serine protease TMPRSS2 primes SARS-CoV-2 glycoprotein for cell entry. Here, the authors perform a screen to identify drugs that reduce TMPRSS2 expression and find that halofuginone modulates proteasome-mediated degradation of TMPRSS2 and reduces entry of SARS-CoV-2.
- Yanwen Chen
- , Travis B. Lear
- & Bill B. Chen
-
Article
| Open Access Drug repurposing screens identify chemical entities for the development of COVID-19 interventions
Drug repurposing screens identify chemical entities for the development of COVID-19 interventions
Here, the authors perform repurposing screens of the ReFRAME drug library in two cell lines and identify inhibitors of SARS-CoV-2 infection. Antiviral activity of prodrug MK-4482 is confirmed in hamsters.
- Malina A. Bakowski
- , Nathan Beutler
- & Thomas F. Rogers
-
Article
| Open Access High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer
High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer
Transcription factor binding dynamics can drive epigenetic states, enabling a diversity of phenotypes. Here the authors present Spear-ATAC to quantify and map perturbations to chromatin accessibility in single cells at high throughput.
- Sarah E. Pierce
- , Jeffrey M. Granja
- & William J. Greenleaf
-
Article
| Open Access Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Highly endangered species like the Sumatran rhinoceros are at risk from inbreeding. Five historical and 16 modern genomes from across the species range show mutational load, but little evidence for local adaptation, suggesting that future inbreeding depression could be mitigated by assisted gene flow among populations.
- Johanna von Seth
- , Nicolas Dussex
- & Love Dalén
-
Article
| Open Access Machine learning guided aptamer refinement and discovery
Machine learning guided aptamer refinement and discovery
Current aptamer discovery approaches are unable to probe the complete space of possible sequences. Here, the authors use machine learning to facilitate the development of DNA aptamers with improved binding affinities, and truncate them without significantly compromising binding affinity.
- Ali Bashir
- , Qin Yang
- & B. Scott Ferguson
-
Article
| Open Access Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen
Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen
Characterizing proteases in their native environment is still challenging. Here, the authors develop a proteomics workflow for analyzing protease-specific peptides from cell lysates in 96-well format, providing mechanistic insights into blood proteases and enabling the prediction of protease substrates.
- Federico Uliana
- , Matej Vizovišek
- & Ruedi Aebersold
-
Article
| Open Access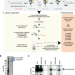 psiCLIP reveals dynamic RNA binding by DEAH-box helicases before and after exon ligation
psiCLIP reveals dynamic RNA binding by DEAH-box helicases before and after exon ligation
ATP-dependent helicases remodel the spliceosome and proofread splice site recognition. A new method – Purified Spliceosome iCLIP (psiCLIP) – probes protein-RNA interactions in defined spliceosome complexes to reveal how the helicases Prp16 and Prp22 promote correct mRNA synthesis through dynamic binding on their RNA substrates.
- Lisa M. Strittmatter
- , Charlotte Capitanchik
- & Kiyoshi Nagai
-
Article
| Open Access A multiplexed, automated evolution pipeline enables scalable discovery and characterization of biosensors
A multiplexed, automated evolution pipeline enables scalable discovery and characterization of biosensors
Biosensors are key to engineered biological systems. Here the authors demonstrate rapid de novo in vitro evolution of RNA biosensors of small molecules in a fully automated system.
- Brent Townshend
- , Joy S. Xiang
- & Christina D. Smolke
-
Article
| Open Access A multiplexed, next generation sequencing platform for high-throughput detection of SARS-CoV-2
A multiplexed, next generation sequencing platform for high-throughput detection of SARS-CoV-2
Wide-spread outbreaks of pathogens require high intensity testing to manage. Here, the authors present C19-SPAR-Seq, a scalable and automated platform to analyse tens of thousands of SARS-CoV-2 patient samples in a single run.
- Marie-Ming Aynaud
- , J. Javier Hernandez
- & Jeffrey L. Wrana
-
Article
| Open Access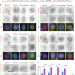 A choreography of centrosomal mRNAs reveals a conserved localization mechanism involving active polysome transport
A choreography of centrosomal mRNAs reveals a conserved localization mechanism involving active polysome transport
Centrosomes function as microtubule organizing centers where several mRNAs accumulate. By employing high-throughput single molecule FISH screening, the authors discover that 8 human mRNAs localize to centrosomes with unique cell cycle dependent patterns using an active polysome targeting mechanism.
- Adham Safieddine
- , Emeline Coleno
- & Edouard Bertrand
-
Article
| Open Access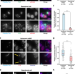 Exon junction complex dependent mRNA localization is linked to centrosome organization during ciliogenesis
Exon junction complex dependent mRNA localization is linked to centrosome organization during ciliogenesis
Exon junction complexes (EJCs) that mark untranslated mRNA are involved in transport, translation and nonsense-mediated mRNA decay. Here the authors show centrosomal localization of EJCs which appears to be required for both the localization of NIN mRNA around centrosomes and ciliogenesis.
- Oh Sung Kwon
- , Rahul Mishra
- & Hervé Le Hir
-
Article
| Open Access Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Given the severity of the SARS-CoV-2 pandemic, a major challenge is to rapidly repurpose existing approved drugs for clinical interventions. Here, the authors identify robust druggable protein targets within a principled causal framework that makes use of multiple data modalities and integrates aging signatures.
- Anastasiya Belyaeva
- , Louis Cammarata
- & Caroline Uhler
-
Article
| Open Access An all-to-all approach to the identification of sequence-specific readers for epigenetic DNA modifications on cytosine
An all-to-all approach to the identification of sequence-specific readers for epigenetic DNA modifications on cytosine
Identifying readers of epigenetic marks is a critical step for understanding the role of epigenetic marks in biology. Here, the authors applied DAPPL, an all-to-all approach to profile the interactions between TFs and epigenetic modified DNA libraries.
- Guang Song
- , Guohua Wang
- & Heng Zhu
-
Article
| Open Access High-throughput phenotypic screen and transcriptional analysis identify new compounds and targets for macrophage reprogramming
High-throughput phenotypic screen and transcriptional analysis identify new compounds and targets for macrophage reprogramming
Macrophages may polarize into different states with distinct regulatory functions for inflammation. Here the authors perform high-throughput in vitro screening of a library of ~4000 compounds to identify those with specific effects on human macrophage polarization, while RNAseq helps uncover the targets and pathways mediating these effects.
- Guangan Hu
- , Yang Su
- & Jianzhu Chen
-
Article
| Open Access RNA structure-wide discovery of functional interactions with multiplexed RNA motif library
RNA structure-wide discovery of functional interactions with multiplexed RNA motif library
Structured RNA motifs can be obtained by structure probing, duplex capture, and motif prediction. Here the authors develop a multiplexed affinity assay system to identify functional protein interactors from an RNA structure library with validated or predicted RNA motifs.
- Kaoru R. Komatsu
- , Toshiki Taya
- & Hirohide Saito
-
Article
| Open Access Leveraging multi-way interactions for systematic prediction of pre-clinical drug combination effects
Leveraging multi-way interactions for systematic prediction of pre-clinical drug combination effects
Combinatorial treatments have become a standard of care for various complex diseases including cancers. Here, the authors show that combinatorial responses of two anticancer drugs can be accurately predicted using factorization machines trained on large-scale pharmacogenomic data for guiding precision oncology studies.
- Heli Julkunen
- , Anna Cichonska
- & Juho Rousu
-
Article
| Open Access UMI-linked consensus sequencing enables phylogenetic analysis of directed evolution
UMI-linked consensus sequencing enables phylogenetic analysis of directed evolution
The success of protein evolution is dependent on the sequence context mutations are introduced into. Here the authors present UMIC-seq that allows consensus generation for closely related genes by using unique molecular identifiers linked to gene variants.
- Paul Jannis Zurek
- , Philipp Knyphausen
- & Florian Hollfelder
-
Article
| Open Access Plant hairy roots enable high throughput identification of antimicrobials against Candidatus Liberibacter spp.
Plant hairy roots enable high throughput identification of antimicrobials against Candidatus Liberibacter spp.
The putative causal agent of citrus greening Candidatus Liberibacter asiaticus (CLas) cannot be cultured, which hampers finding new therapies to control this devastating disease. Here, the authors show that hairy roots support CLas propagation and enable high throughput antimicrobial screening.
- Sonia Irigoyen
- , Manikandan Ramasamy
- & Kranthi K. Mandadi
-
Article
| Open Access Pharmacological targeting of MCL-1 promotes mitophagy and improves disease pathologies in an Alzheimer’s disease mouse model
Pharmacological targeting of MCL-1 promotes mitophagy and improves disease pathologies in an Alzheimer’s disease mouse model
Previous work suggests that mitophagy in neurons is could be therapeutic in Alzheimer’s disease (AD). Here, the authors screen a library of drugs and identify UMI-77, a mitophagy inducer with beneficial effects in an AD mouse model, by binding MCL-1, which they identify as a mitophagy receptor.
- Xufeng Cen
- , Yanying Chen
- & Hongguang Xia
-
Article
| Open Access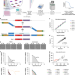 Quantitative and multiplexed chemical-genetic phenotyping in mammalian cells with QMAP-Seq
Quantitative and multiplexed chemical-genetic phenotyping in mammalian cells with QMAP-Seq
Identifying chemical-genetic interactions in mammalian cells is limited to low-throughput or computational methods. Here, the authors present QMAP-Seq, a broadly accessible and scalable approach that uses NGS for pooled high-throughput chemical-genetic profiling in mammalian cells.
- Sonia Brockway
- , Geng Wang
- & Marc L. Mendillo
-
Article
| Open Access A combined high-throughput and high-content platform for unified on-chip synthesis, characterization and biological screening
A combined high-throughput and high-content platform for unified on-chip synthesis, characterization and biological screening
On-chip synthesis and screening has been used to automate drug discovery but on-chip analysis still remains a major limitation. Here, the authors report on a dendrimer-based surface patterning method to create nanodroplet arrays on materials which allow for on-chip high-throughput analysis.
- Maximilian Benz
- , Arndt Asperger
- & Pavel A. Levkin
-
Article
| Open Access A nanoluciferase SARS-CoV-2 for rapid neutralization testing and screening of anti-infective drugs for COVID-19
A nanoluciferase SARS-CoV-2 for rapid neutralization testing and screening of anti-infective drugs for COVID-19
A high-throughput platform would greatly facilitate coronavirus disease 2019 (COVID-19) serological testing and antiviral screening. To address this, Shi and colleagues present a high-throughput nanoluciferase severe respiratory syndrome coronavirus 2 (SARS-CoV2-Nluc), and show that it has potential for large-scale vaccine evaluation and neutralizing antibody testing.
- Xuping Xie
- , Antonio E. Muruato
- & Pei-Yong Shi
-
Article
| Open Access CRISPR screening of porcine sgRNA library identifies host factors associated with Japanese encephalitis virus replication
CRISPR screening of porcine sgRNA library identifies host factors associated with Japanese encephalitis virus replication
Here the authors report the construction of a genome-scale porcine CRISPR/Cas9 library, called PigGeCKO, for screening and analyses of host resistance genes and factors associated with Japanese encephalitis virus replication.
- Changzhi Zhao
- , Hailong Liu
- & Shuhong Zhao
-
Article
| Open Access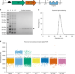 A role for Biofoundries in rapid development and validation of automated SARS-CoV-2 clinical diagnostics
A role for Biofoundries in rapid development and validation of automated SARS-CoV-2 clinical diagnostics
The SARS-CoV-2 pandemic has created large demand on global testing capability. Here the authors use the London Biofoundry, an automated synthetic biology platform, and develop an open-source virus-like particle to implement high-throughput diagnostics.
- Michael A. Crone
- , Miles Priestman
- & Paul S. Freemont
-
Article
| Open Access Both Boceprevir and GC376 efficaciously inhibit SARS-CoV-2 by targeting its main protease
Both Boceprevir and GC376 efficaciously inhibit SARS-CoV-2 by targeting its main protease
Coronavirus main protease is essential for viral polyprotein processing and maturation. Here Fu et al. report efficient inhibition of SARS-CoV-2 replication using two inhibitors - Boceprevir and GC376 - targeting the active site of the main viral protease.
- Lifeng Fu
- , Fei Ye
- & George Fu Gao
-
Article
| Open Access Engineering regulatory networks for complex phenotypes in E. coli
Engineering regulatory networks for complex phenotypes in E. coli
Regulatory networks respond to environmental and genetic perturbations by reprogramming metabolism. Here the authors screen a library of 82 regulators with 110,120 mutations to map the regulatory network of 4000 genes.
- Rongming Liu
- , Liya Liang
- & Ryan T. Gill
-
Article
| Open Access In vitro Cas9-assisted editing of modular polyketide synthase genes to produce desired natural product derivatives
In vitro Cas9-assisted editing of modular polyketide synthase genes to produce desired natural product derivatives
Several different genetic strategies have been reported for the modification of polyketide synthases but the highly repetitive modular structure makes this difficult. Here the authors report on an adapted Cas9 reaction and Gibson assembly to edit a target region of the polyketide synthases gene in vitro.
- Kei Kudo
- , Takuya Hashimoto
- & Kazuo Shin-ya
-
Article
| Open Access Strategies to enable large-scale proteomics for reproducible research
Strategies to enable large-scale proteomics for reproducible research
Clinical proteomics critically depends on the ability to acquire highly reproducible data over an extended period of time. Here, the authors assess reproducibility over four months across different mass spectrometers and develop a computational approach to mitigate variation among instruments over time.
- Rebecca C. Poulos
- , Peter G. Hains
- & Qing Zhong
-
Article
| Open Access Raman image-activated cell sorting
Raman image-activated cell sorting
Most current cell sorting methods are based on fluorescence detection with no imaging capability. Here the authors generate and use Raman image-activated cell sorting with a throughput of around 100 events per second, providing molecular images with no need for labeling.
- Nao Nitta
- , Takanori Iino
- & Keisuke Goda
-
Article
| Open Access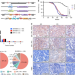 NOTCH1 activation compensates BRCA1 deficiency and promotes triple-negative breast cancer formation
NOTCH1 activation compensates BRCA1 deficiency and promotes triple-negative breast cancer formation
BRCA1 mutation carriers have higher chances of developing triple-negative breast cancer (TNBC). Here, the authors use the Sleeping Beauty mutagenesis system in Brca1 deficient mice and identify 169 putative driver genes, of which NOTCH1 accelerates TNBC formation through promoting epithelial-mesenchymal transition and cell cycle progression.
- Kai Miao
- , Josh Haipeng Lei
- & Chu-Xia Deng
-
Article
| Open Access Fast and sensitive flow-injection mass spectrometry metabolomics by analyzing sample-specific ion distributions
Fast and sensitive flow-injection mass spectrometry metabolomics by analyzing sample-specific ion distributions
Flow-injection mass spectrometry (FI-MS) enables high-throughput metabolomic profiling, but ion overload typically limits its sensitivity. Here, the authors show rapid and highly sensitive FI-MS overcoming an overload of the Orbitrap by analyzing sample-specific ion distributions.
- Boris Sarvin
- , Shoval Lagziel
- & Tomer Shlomi
-
Article
| Open Access High-throughput interrogation of programmed ribosomal frameshifting in human cells
High-throughput interrogation of programmed ribosomal frameshifting in human cells
Programmed ribosomal frameshifting—the slippage of the ribosome to an alternative frame — is critical for viral replication and cellular processes. Here the authors present an approach that can assess the frameshifting potential of a sequence and elucidate the rules governing ribosomal frameshifting.
- Martin Mikl
- , Yitzhak Pilpel
- & Eran Segal
-
Article
| Open Access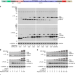 Split Intein-Mediated Protein Ligation for detecting protein-protein interactions and their inhibition
Split Intein-Mediated Protein Ligation for detecting protein-protein interactions and their inhibition
Protein-protein interactions are fundamental to the regulation of protein activity and cellular phyisology. Here the authors present Split Intein-Mediated Protein Ligation, which uses bait and prey proteins fused to intein fragments to generate single intact proteins upon interaction.
- Zhong Yao
- , Farzaneh Aboualizadeh
- & Igor Stagljar
-
Article
| Open Access High-throughput cell and spheroid mechanics in virtual fluidic channels
High-throughput cell and spheroid mechanics in virtual fluidic channels
High-throughput rheological measurements of cells and cell clusters by microfluidics is limited by fixed channel dimensions. Here the authors create virtual fluidic channels inside the cuvette of commercial flow cytometers to dynamically tune channel cross section to enable rheological measurements from cells and cell clusters.
- Muzaffar H. Panhwar
- , Fabian Czerwinski
- & Oliver Otto

 Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit
Massively parallel interrogation of protein fragment secretability using SECRiFY reveals features influencing secretory system transit