Featured
-
-
Article
| Open Access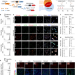 Genetic fate-mapping reveals surface accumulation but not deep organ invasion of pleural and peritoneal cavity macrophages following injury
Genetic fate-mapping reveals surface accumulation but not deep organ invasion of pleural and peritoneal cavity macrophages following injury
Body cavity macrophages reside on the serous surfaces of organs and believed to participate in organ repair following injury. Here the authors show with a fate-mapping reporter system that these cells, although accumulate at the surfaces of injured liver or lung, don’t penetrate deeply into the tissue.
- Hengwei Jin
- , Kuo Liu
- & Bin Zhou
-
Article
| Open Access Cryopreservation method for Drosophila melanogaster embryos
Cryopreservation method for Drosophila melanogaster embryos
The development of a widely adopted cryopreservation method remains a major challenge in Drosophila melanogaster research. Here the authors report a robust cryopreservation protocol of Drosophila embryos and showcase its implementation in 25 distinct strains from different sources.
- Li Zhan
- , Min-gang Li
- & John Bischof
-
Article
| Open Access Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Circulating tumour DNA (ctDNA) represents a non-invasive option to monitor cancer progression. Here, the authors perform deep sequencing of plasma cell-free DNA, and find that nucleosome-dependent cfDNA degradation at 6 specific regulatory regions is predictive of ctDNA burden.
- Guanhua Zhu
- , Yu A. Guo
- & Anders J. Skanderup
-
Article
| Open Access Universal toxin-based selection for precise genome engineering in human cells
Universal toxin-based selection for precise genome engineering in human cells
Genome engineering in cell lines or human stem cells often has poor efficiency, limiting the development of research and therapeutic applications. Here, the authors use a toxin-based selection system for precise bi-allelic engineering in cells and in vivo.
- Songyuan Li
- , Nina Akrap
- & Marcello Maresca
-
Article
| Open Access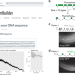 Engineering rules that minimize germline silencing of transgenes in simple extrachromosomal arrays in C. elegans
Engineering rules that minimize germline silencing of transgenes in simple extrachromosomal arrays in C. elegans
C. elegans has strong repressive mechanisms that silence transgenes in the germline. Here the authors elucidate the design rules for efficient germline expression.
- Mohammed D. Aljohani
- , Sonia El Mouridi
- & Christian Frøkjær-Jensen
-
Article
| Open Access A Cas-embedding strategy for minimizing off-target effects of DNA base editors
A Cas-embedding strategy for minimizing off-target effects of DNA base editors
DNA base editors can display off-target effects on DNA and RNA. Here the authors embed the base editing enzymes in the middle of nCas9 to reduce these without impacting on-target editing.
- Yajing Liu
- , Changyang Zhou
- & Tian Chi
-
Article
| Open Access MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
Cas9 fused to DNA damage repair proteins can improve rates of gene knock-in but the chimeric protein is often large. Here the authors fuse Cas9 to a minimal Brex27 motif from BRCA2 consisting of thirty-six amino acids to enhance the efficacy and safety of gene editing.
- Linyuan Ma
- , Jinxue Ruan
- & Jie Xu
-
Article
| Open Access An enhanced isothermal amplification assay for viral detection
An enhanced isothermal amplification assay for viral detection
Current state-of-the-art diagnostics for infectious diseases are sensitive but require extensive equipment. Here the authors develop an enhanced recombinase polymerase amplification reaction for SARS-CoV-2 that allows for inexpensive and rapid testing with minimal equipment.
- Jason Qian
- , Sarah A. Boswell
- & Michael Springer
-
Article
| Open Access Docking sites inside Cas9 for adenine base editing diversification and RNA off-target elimination
Docking sites inside Cas9 for adenine base editing diversification and RNA off-target elimination
Current SpCas9 adenine base editors that minimise RNA off-target activities have constrained editing windows. Here the authors use domain insertion of TadA into Cas9 to narrow, expand or shift the editing window with RNA off-target minimization simultaneously
- Shuo Li
- , Bo Yuan
- & Tian-Lin Cheng
-
Article
| Open Access The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice
The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice
Auxin-inducible degron systems can be leaky and require high doses of auxin. Here the authors establish AID2 which uses an OsTIR1 mutant and the ligand 5-Ph-IAA to overcome these problems and establish AID-mediated target depletion in mice.
- Aisha Yesbolatova
- , Yuichiro Saito
- & Masato T. Kanemaki
-
Article
| Open Access Synthesizing AND gate minigene circuits based on CRISPReader for identification of bladder cancer cells
Synthesizing AND gate minigene circuits based on CRISPReader for identification of bladder cancer cells
Synthetic biology logic gates can be used to distinguish healthy cells from cancer cells. Here the authors design minigene circuits that show more robust identification of cancer cells compared to traditional genetic circuits.
- Yuchen Liu
- , Weiren Huang
- & Zhiming Cai
-
Article
| Open Access Prime editing for functional repair in patient-derived disease models
Prime editing for functional repair in patient-derived disease models
Prime editing uses Cas9 nickase fused to a reverse transcriptase to edit genetic information. Here, the authors prime edit primary adult stem cells in 3D organoid cultures to show functional correction of pathogenic mutations without genome-wide off-target effects.
- Imre F. Schene
- , Indi P. Joore
- & Sabine A. Fuchs
-
Article
| Open Access Enhancement of trans-cleavage activity of Cas12a with engineered crRNA enables amplified nucleic acid detection
Enhancement of trans-cleavage activity of Cas12a with engineered crRNA enables amplified nucleic acid detection
CRISPR-Cas12a based detection systems can be sensitive down to the picomolar range. Here the authors modify the 3′- and 5′-ends of the crRNA and show this enhances trans-cleavage for improved sensitivity.
- Long T. Nguyen
- , Brianna M. Smith
- & Piyush K. Jain
-
Article
| Open Access Analysis of chromatin organization and gene expression in T cells identifies functional genes for rheumatoid arthritis
Analysis of chromatin organization and gene expression in T cells identifies functional genes for rheumatoid arthritis
Although genome-wide association studies have identified genetic variation contributing to disease risk, assigning causal genes is challenging. Here, the authors generate ATAC-seq, Hi-C, Capture Hi-C and RNA-seq data in stimulated CD4+ T cells to identify functional enhancers and demonstrate interactions of expression quantitative trait loci with target genes in rheumatoid arthritis.
- Jing Yang
- , Amanda McGovern
- & Stephen Eyre
-
Article
| Open Access A scalable CRISPR/Cas9-based fluorescent reporter assay to study DNA double-strand break repair choice
A scalable CRISPR/Cas9-based fluorescent reporter assay to study DNA double-strand break repair choice
Cells employ different repair pathways to repair DNA double strand breaks. Here, the authors develop a CRISPR/Cas9-dependent method to study choices in DNA repair called the Color Assay Tracing-Repair (CAT-R) which simultaneously measure outcomes of DSB repair via end-protection and end-resection pathways.
- Paris Roidos
- , Stephanie Sungalee
- & Balca R. Mardin
-
Article
| Open Access Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
The complexity of structural variation (SV) and short tandem repeats (STRs) makes it necessary to apply different calling and filtering strategies to sequencing datasets. Here, Jakubosky et al. report a comprehensive SV and STR callset from whole-genome sequencing of 477 individuals from iPSCORE and HipSci using five algorithms.
- David Jakubosky
- , Erin N. Smith
- & Kelly A. Frazer
-
Article
| Open Access Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Numerous strategies exist to limit the off-target activity of CRISPR-Cas9 nucleases. Here the authors co-administer truncated gRNAs that block both Cas9 and high-fidelity Cas9 variants from cleaving at off-target sites.
- John C. Rose
- , Nicholas A. Popp
- & Douglas M. Fowler
-
Article
| Open Access A critical role of PRDM14 in human primordial germ cell fate revealed by inducible degrons
A critical role of PRDM14 in human primordial germ cell fate revealed by inducible degrons
PRDM14 is a critical transcription factor for mouse primordial germ cell specification, but its role in human remains unclear. Here, PRDM14 protein depletion using auxin-inducible degron uncovers a critical role for human germ cell specification, but regulation of a different set of target genes than in mouse.
- Anastasiya Sybirna
- , Walfred W. C. Tang
- & M. Azim Surani
-
Article
| Open Access Synthetic chimeric nucleases function for efficient genome editing
Synthetic chimeric nucleases function for efficient genome editing
CRISPR-Cas systems have well characterized, modular structures. Here the authors use that architecture to design a Cas12a library of 560 synthetic chimeras, with altered PAM preferences and specificities.
- R. M. Liu
- , L. L. Liang
- & R. T. Gill
-
Article
| Open Access Functional profiling of single CRISPR/Cas9-edited human long-term hematopoietic stem cells
Functional profiling of single CRISPR/Cas9-edited human long-term hematopoietic stem cells
Previous gene editing in haematopoietic stem cells (HSCs) has focussed on a heterogeneous CD34+ population. Here, the authors demonstrate high efficiency CRISPR/Cas9-based editing of purified long-term HSCs using non-homologous end joining and homology-directed repair, by directing isoform-specific expression of GATA1.
- Elvin Wagenblast
- , Maria Azkanaz
- & Eric R. Lechman
-
Article
| Open Access Dendronized fluorosurfactant for highly stable water-in-fluorinated oil emulsions with minimal inter-droplet transfer of small molecules
Dendronized fluorosurfactant for highly stable water-in-fluorinated oil emulsions with minimal inter-droplet transfer of small molecules
Microdroplets are used as chemical and biological reactors; however, stability and inter-droplet transfer are major issues. Here, the authors report on the development of dendritic glycerol-based surfactants for the creation of stable microdroplets and demonstrate application for PCR, minimal emulsion, and cell encapsulation.
- Mohammad Suman Chowdhury
- , Wenshan Zheng
- & Rainer Haag
-
Article
| Open Access Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Meiotic crossovers (COs) generate genetic variation and ensure proper chromosome segregation. Here, the authors develop a method for identifying COs at kilobase resolution in pooled recombinants using linked-read sequencing data, and apply it to investigate genome-wide CO landscapes of Arabidopsis thaliana.
- Hequan Sun
- , Beth A. Rowan
- & Korbinian Schneeberger
-
Article
| Open Access Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Application of highly specific Cas9 variants can be restricted by the design of the guide RNA. Here the authors present DeepHF, a gRNA activity prediction tool built from genome-scale screens of 50,000 guides covering 20,000 genes.
- Daqi Wang
- , Chengdong Zhang
- & Yongming Wang
-
Article
| Open Access Expanding C–T base editing toolkit with diversified cytidine deaminases
Expanding C–T base editing toolkit with diversified cytidine deaminases
Cytosine base editors are limited by editing scope and potential off-target effects. Here the authors screen diversified lamprey cytidine deaminases along with different protein fusion architectures and present base editors with improved fidelity.
- Tian-Lin Cheng
- , Shuo Li
- & Zilong Qiu
-
Article
| Open Access Development of hRad51–Cas9 nickase fusions that mediate HDR without double-stranded breaks
Development of hRad51–Cas9 nickase fusions that mediate HDR without double-stranded breaks
Here the authors fuse hRad51 and variants thereof to Cas9 nickase to facilitate homology-directed repair without generating double strand breaks, minimizing indel formation and off-target editing. This tool represents progress towards the goal of performing HDR without an excess of undesired side products.
- Holly A. Rees
- , Wei-Hsi Yeh
- & David R. Liu
-
Article
| Open Access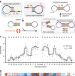 High-resolution specificity profiling and off-target prediction for site-specific DNA recombinases
High-resolution specificity profiling and off-target prediction for site-specific DNA recombinases
The development of site-specific recombinases as genome editing tools is limited by the difficulty of altering their DNA sequence specificity. Here the authors present Rec-seq, a method for identifying specificity determinants and off-target substrates of recombinases in an unbiased manner.
- Jeffrey L. Bessen
- , Lena K. Afeyan
- & David R. Liu
-
Article
| Open Access Multiplexed Cas9 targeting reveals genomic location effects and gRNA-based staggered breaks influencing mutation efficiency
Multiplexed Cas9 targeting reveals genomic location effects and gRNA-based staggered breaks influencing mutation efficiency
Designing effective genome engineering strategies requires an understanding of the impact that genomic locus has on CRISPR-Cas9 activity. Here the authors use TRIP integrations to profile editing outcomes genome-wide and observe that gRNA sequence influences the structure of the double strand break.
- Santiago Gisler
- , Joana P. Gonçalves
- & Maarten van Lohuizen
-
Article
| Open Access Diversifying the structure of zinc finger nucleases for high-precision genome editing
Diversifying the structure of zinc finger nucleases for high-precision genome editing
Genome editing often requires cleavage within a narrow sequence window. Here the authors develop an expanded set of zinc finger nuclease architectures that increase the available configurations by a factor of 64 and can target almost every base at loci of therapeutic significance.
- David E. Paschon
- , Stephanie Lussier
- & Edward J. Rebar
-
Article
| Open Access Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer
Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer
Triple negative breast cancers (TNBC) disseminate and metastasise, but the clonal relationship of metastases to primary tumours is poorly understood. Here, the authors use cellular barcoding of TNBC patient-derived xenografts and track the fate of barcoded clones in primary tumours and their metastases, including after resection or chemotherapy.
- D. Merino
- , T. S. Weber
- & S. H. Naik
-
Article
| Open Access Engineering of high-precision base editors for site-specific single nucleotide replacement
Engineering of high-precision base editors for site-specific single nucleotide replacement
Base editors can target multiple bases within a window around the target site, reducing their specificity. Here the authors engineer the connection between the deaminase and Cas domain to narrow the window of activity.
- Junjie Tan
- , Fei Zhang
- & Ralph Bock
-
Article
| Open Access An efficient and multiple target transgenic RNAi technique with low toxicity in Drosophila
An efficient and multiple target transgenic RNAi technique with low toxicity in Drosophila
Drosophila transgenic RNAi can have drawbacks such as false positives and negative results. Here the authors develop a next generation RNAi system with reduced leakiness of expression and simultaneous knockdown.
- Huan-Huan Qiao
- , Fang Wang
- & Jian-Quan Ni
-
Article
| Open Access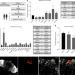 Discovering human diabetes-risk gene function with genetics and physiological assays
Discovering human diabetes-risk gene function with genetics and physiological assays
The function of genes linked to type 2 diabetes is poorly characterized. Here the authors combine Drosophila genetics and physiology with human islet biology to identify new regulators of insulin secretion including BCL11A.
- Heshan Peiris
- , Sangbin Park
- & Seung K. Kim
-
Article
| Open Access Highly efficient genome editing by CRISPR-Cpf1 using CRISPR RNA with a uridinylate-rich 3′-overhang
Highly efficient genome editing by CRISPR-Cpf1 using CRISPR RNA with a uridinylate-rich 3′-overhang
Cpf1 is a promising alternative to Cas9 though indel generation efficiency is target dependent. Here the authors show that the addition of a polyU 3′ overhang can improve the efficiency of low efficiency guide RNAs.
- Su Bin Moon
- , Jeong Mi Lee
- & Yong-Sam Kim
-
Article
| Open Access In vivo base editing of post-mitotic sensory cells
In vivo base editing of post-mitotic sensory cells
Base editing allows the precise introduction of point mutations into cellular DNA without requiring double-stranded DNA breaks or homology-directed repair, which is inefficient in postmitotic cells. Here the authors demonstrate in vivo base editing of post-mitotic somatic cells in the postnatal mouse inner ear with physiological outcomes.
- Wei-Hsi Yeh
- , Hao Chiang
- & David R. Liu
-
Article
| Open Access GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality
GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality
Rice grain shape or size is an important trait associated with both yield and appearance quality. Here, the authors identify GS9 as a negative transcription regulator of slender grain and show it can improve grain shape and appearance independently from other previously identified grain size genes.
- Dong-Sheng Zhao
- , Qian-Feng Li
- & Qiao-Quan Liu
-
Article
| Open Access Incomplete prophage tolerance by type III-A CRISPR-Cas systems reduces the fitness of lysogenic hosts
Incomplete prophage tolerance by type III-A CRISPR-Cas systems reduces the fitness of lysogenic hosts
CRISPR-Cas systems, such as type III-A CRISPR-Cas, provide an immune mechanism for prokaryotic hosts to resist parasites, including phages. Here, the authors show that maintenance of conditionally tolerant type III-A systems can affect the fitness of Staphylococcus aureus lysogens.
- Gregory W. Goldberg
- , Elizabeth A. McMillan
- & Luciano A. Marraffini
-
Article
| Open Access Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Regeneration capable flatworms have emerged as powerful models for studying stem cell biology and patterning, however their study has been hindered by the lack of transgenesis methods. Here, the authors describe a transgenesis method for Macrostomum lignano, as well as a new annotated genome sequence.
- Jakub Wudarski
- , Daniil Simanov
- & Eugene Berezikov
-
Article
| Open Access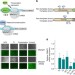 Engineering cell signaling using tunable CRISPR–Cpf1-based transcription factors
Engineering cell signaling using tunable CRISPR–Cpf1-based transcription factors
Cpf1 has been repurposed as a transcriptional repressor in bacteria and plants. Here, the authors construct activators and repressors in human cells using Cpf1 coupled to riboswitches and GPCRs.
- Yuchen Liu
- , Jinghong Han
- & Guohui Nie
-
Article
| Open Access Robust RNA-based in situ mutation detection delineates colorectal cancer subclonal evolution
Robust RNA-based in situ mutation detection delineates colorectal cancer subclonal evolution
Methods that analyze intra-tumor genetic heterogeneity often do not preserve the spatial context of tumor subclones. Here, the authors present BaseScope, a mutation-specific RNA in situ hybridization assay and spatially map colorectal cancer and adenoma KRAS, BRAF and PIK3CA driver gene mutant subclones.
- Ann-Marie Baker
- , Weini Huang
- & Trevor A. Graham
-
Article
| Open Access Fusion guide RNAs for orthogonal gene manipulation with Cas9 and Cpf1
Fusion guide RNAs for orthogonal gene manipulation with Cas9 and Cpf1
Cas9 and Cpf1 have both been adapted for genome engineering, editing and gene expression regulation. Here the authors design a fusion guide RNA that can interact with both proteins for multiple and orthogonal genome manipulation.
- Jiyeon Kweon
- , An-Hee Jang
- & Yongsub Kim
-
Article
| Open Access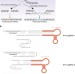 Assembly of CRISPR ribonucleoproteins with biotinylated oligonucleotides via an RNA aptamer for precise gene editing
Assembly of CRISPR ribonucleoproteins with biotinylated oligonucleotides via an RNA aptamer for precise gene editing
Using CRISPR to write specific genetic sequences can sometimes be difficult due to the preference of mammalian cells to repair breaks using NHEJ. Here the authors form nanoparticles to localize the template sequence to the nuclease, shifting repair in favor of HDR.
- Jared Carlson-Stevermer
- , Amr A. Abdeen
- & Krishanu Saha
-
Article
| Open Access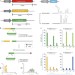 In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni
In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni
The amount of genetic material that can be packaged in AAV vectors used for genome editing is limited. Here the authors show that the smallest known Cas9 orthologue, cjCas9, can be packaged in a single AAV vector along with sgRNA and a marker gene, and demonstrate efficient gene editing in mice.
- Eunji Kim
- , Taeyoung Koo
- & Jin-Soo Kim
-
Article
| Open Access Ligand-binding domains of nuclear receptors facilitate tight control of split CRISPR activity
Ligand-binding domains of nuclear receptors facilitate tight control of split CRISPR activity
CRISPR-Cas9 has been widely used to manipulate genomes, however control over activity is necessary to explore transcriptional or epigenetic regulation. Here the authors use a split Cas9 fused to nuclear receptor ligand-binding domains to achieve tuneable Cas9 activity.
- Duy P. Nguyen
- , Yuichiro Miyaoka
- & James A. Wells
-
Article
| Open Access Genetic dissection of mammalian ERAD through comparative haploid and CRISPR forward genetic screens
Genetic dissection of mammalian ERAD through comparative haploid and CRISPR forward genetic screens
CRISPR/Cas9-mediated forward genetic screens and gene-trap mutagenesis screens in haploid cells are both powerful techniques to examine gene function. Here, the authors show the two approaches have high concordance and identify an uncharacterized gene, TXNDC11, which is involved in endoplasmic reticulum-associated degradation.
- Richard T. Timms
- , Sam A. Menzies
- & Paul J. Lehner
-
Article
| Open Access Circulating tumour DNA profiling reveals heterogeneity of EGFR inhibitor resistance mechanisms in lung cancer patients
Circulating tumour DNA profiling reveals heterogeneity of EGFR inhibitor resistance mechanisms in lung cancer patients
EGFR-mutant non-small cell lung cancer is routinely treated with EGFR inhibitors, although resistance inevitably develops. Here, the authors sequence circulating tumour DNA and show that resistance to the third-generation inhibitor rociletinib is heterogeneous and recurrently involves somatic alterations of MET, EGFR, PIK3CA, ERRB2, and KRAS.
- Jacob J. Chabon
- , Andrew D. Simmons
- & Maximilian Diehn
-
Article
| Open Access Internal guide RNA interactions interfere with Cas9-mediated cleavage
Internal guide RNA interactions interfere with Cas9-mediated cleavage
While the CRISPR-Cas9 system has revolutionised molecular biology, it is still a mystery why not every guide RNA elicits target DNA cleavage. Here the authors show that genomic context and internal gRNA interactions can inhibit cleavage.
- Summer B. Thyme
- , Laila Akhmetova
- & Alexander F. Schier
-
Article
| Open Access Design of a bioactive small molecule that targets r(AUUCU) repeats in spinocerebellar ataxia 10
Design of a bioactive small molecule that targets r(AUUCU) repeats in spinocerebellar ataxia 10
Expanded RNA repeats in non-coding region of a gene represent a hallmark of several diseases. Here, the authors identify two small molecules that selectively bind AU repeats and use them to design a compound that targets the pathogenic RNA associated with spinocerebellar ataxia type 10.
- Wang-Yong Yang
- , Rui Gao
- & Matthew D. Disney
-
Article
| Open Access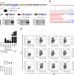 Editing of mouse and human immunoglobulin genes by CRISPR-Cas9 system
Editing of mouse and human immunoglobulin genes by CRISPR-Cas9 system
CRISPR-Cas9 has been used to generate a range of genetic modifications including gene knock-outs and chromosomal rearrangements. Here the authors target the immunogloblin genes and demonstrate the induction of class switch recombination, opening up possibilities for research and antibody production.
- Taek-Chin Cheong
- , Mara Compagno
- & Roberto Chiarle
-
Article
| Open Access Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice
Multiplexed pancreatic genome engineering and cancer induction by transfection-based CRISPR/Cas9 delivery in mice
CRISPR/Cas9 technology has been used for genome engineering in vivo. Here, the authors use a transfection technique to deliver multiple guide RNAs to the pancreas of adult mice, allowing genetic screening and chromosome engineering in pancreatic cancer.
- Roman Maresch
- , Sebastian Mueller
- & Roland Rad

 Multiplexed detection of SARS-CoV-2 and other respiratory infections in high throughput by SARSeq
Multiplexed detection of SARS-CoV-2 and other respiratory infections in high throughput by SARSeq