Article
|
Open Access
Featured
-
-
Article |
 Convergence of coronary artery disease genes onto endothelial cell programs
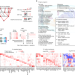
Convergence of coronary artery disease genes onto endothelial cell programs
Variant-to-gene-to-program is a new approach to building maps of genome function to link risk variants to disease genes and to convergent signalling pathways in an unbiased manner; its strength is demonstrated in coronary artery disease.
- Gavin R. Schnitzler
- , Helen Kang
- & Jesse M. Engreitz
-
Article
| Open Access Affinity-optimizing enhancer variants disrupt development
Affinity-optimizing enhancer variants disrupt development
Low-affinity transcription factor binding sites are prevalent across the genome, and single nucleotide changes that increase binding affinity even slightly can cause gain-of-function gene expression and phenotypes (such as polydactyly).
- Fabian Lim
- , Joe J. Solvason
- & Emma K. Farley
-
Article
| Open Access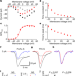 Nav1.7 as a chondrocyte regulator and therapeutic target for osteoarthritis
Nav1.7 as a chondrocyte regulator and therapeutic target for osteoarthritis
The voltage-gated sodium channel Nav1.7 has a dual role in osteoarthritis—in chondrocytes, it promotes joint damage, and in dorsal root ganglia neurons, it increases pain transmission.
- Wenyu Fu
- , Dmytro Vasylyev
- & Chuan-ju Liu
-
Perspective |
 Hold out the genome: a roadmap to solving the cis-regulatory code
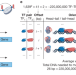
Hold out the genome: a roadmap to solving the cis-regulatory code
A roadmap towards solving the cis-regulatory code using a combination of machine learning and massively parallel assays of exogenous DNA is proposed.
- Carl G. de Boer
- & Jussi Taipale
-
Article
| Open Access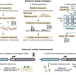 Cell-type-directed design of synthetic enhancers
Cell-type-directed design of synthetic enhancers
Deep learning models were used to design synthetic cell-type-specific enhancers that work in fruit fly brains and human cell lines, an approach that also provides insights into these gene regulatory elements.
- Ibrahim I. Taskiran
- , Katina I. Spanier
- & Stein Aerts
-
Article
| Open Access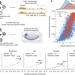 Targeted design of synthetic enhancers for selected tissues in the Drosophila embryo
Targeted design of synthetic enhancers for selected tissues in the Drosophila embryo
Deep learning and transfer learning were used to design tissue-specific enhancers in the Drosophila embryo that were active and specific, validating this approach to achieve tissue-, cell type- and cell state-specific expression control.
- Bernardo P. de Almeida
- , Christoph Schaub
- & Alexander Stark
-
Article
| Open Access The sex-specific factor SOA controls dosage compensation in Anopheles mosquitoes
The sex-specific factor SOA controls dosage compensation in Anopheles mosquitoes
A newly identified gene, sex chromosome activation (SOA), is a master regulator of dosage compensation in Anopheles gambiae.
- Agata Izabela Kalita
- , Eric Marois
- & Claudia Isabelle Keller Valsecchi
-
Article
| Open Access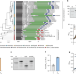 Small protein modules dictate prophage fates during polylysogeny
Small protein modules dictate prophage fates during polylysogeny
Prophage lysogeny-to-lysis transitions are controlled by regulatory modules consisting of transcription factors and partner small proteins that are activated through DNA-damage-independent pathways, including by quorum sensing, and these modules determine inter-prophage competition outcomes.
- Justin E. Silpe
- , Olivia P. Duddy
- & Bonnie L. Bassler
-
Article
| Open Access Organization of the human intestine at single-cell resolution
Organization of the human intestine at single-cell resolution
Intestinal cell types are organized into distinct neighbourhoods and communities within the healthy human intestine, with distinct immunological niches.
- John W. Hickey
- , Winston R. Becker
- & Michael Snyder
-
Article |
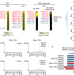 OBOX regulates mouse zygotic genome activation and early development
OBOX regulates mouse zygotic genome activation and early development
OBOX, PRD-like homeobox domain transcription factors (OBOX1–OBOX8), are key regulators of mouse zygotic genome activation and early embryogenesis.
- Shuyan Ji
- , Fengling Chen
- & Wei Xie
-
Article |
 Large-scale mapping and mutagenesis of human transcriptional effector domains
Large-scale mapping and mutagenesis of human transcriptional effector domains
A high throughput recruitment assay testing the transcriptional activity of more than 100,000 protein fragments tiling across most human chromatin regulators and transcription factors maps the locations and strengths of activation, repression and bifunctional domains, and identifies the sequences necessary for these functions.
- Nicole DelRosso
- , Josh Tycko
- & Lacramioara Bintu
-
Article
| Open Access Aberrant phase separation and nucleolar dysfunction in rare genetic diseases
Aberrant phase separation and nucleolar dysfunction in rare genetic diseases
Frameshift mutations that create arginine-rich basic tails in transcription factors and other proteins can lead to altered phase separation in the nucleolus, which in turn leads to syndromes such as brachyphalangy, polydactyly and tibial aplasia.
- Martin A. Mensah
- , Henri Niskanen
- & Denes Hnisz
-
Article |
 A Prox1 enhancer represses haematopoiesis in the lymphatic vasculature
A Prox1 enhancer represses haematopoiesis in the lymphatic vasculature
A transcriptional enhancer element regulates Prox1 expression and lymphatic endothelial cell identity.
- Jan Kazenwadel
- , Parvathy Venugopal
- & Natasha L. Harvey
-
Article |
 Structural variants drive context-dependent oncogene activation in cancer
Structural variants drive context-dependent oncogene activation in cancer
Results are presented that indicate that alterations to gene regulatory three-dimensional architecture are a critical mechanism that enables structural variant-based oncogene activation in cancer genomes and sheds light on the essential elements for such gene activation events.
- Zhichao Xu
- , Dong-Sung Lee
- & Jesse R. Dixon
-
Article
| Open Access A transcriptional switch controls sex determination in Plasmodium falciparum
A transcriptional switch controls sex determination in Plasmodium falciparum
A non-genetic mechanism of sex determination in the human malaria parasite, Plasmodium falciparum, is described, and the male development 1 gene is identified as a potential target for interventions that block malaria transmission.
- A. R. Gomes
- , A. Marin-Menendez
- & A. M. Talman
-
Article |
 The retroelement Lx9 puts a brake on the immune response to virus infection
The retroelement Lx9 puts a brake on the immune response to virus infection
Experiments in mice show that a LINE-1 transposable element, Lx9c11, has a functional role in immunity by negatively regulating the response to viral infection to protect the host from an over-reactive immune response.
- Nenad Bartonicek
- , Romain Rouet
- & Cecile King
-
Article
| Open Access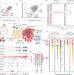 The renal lineage factor PAX8 controls oncogenic signalling in kidney cancer
The renal lineage factor PAX8 controls oncogenic signalling in kidney cancer
The lineage transcription factor PAX8 is shown to play a pivotal part in determining cancer risk in clear cell renal cell carcinoma, providing insights into how genetic mutations lead to specific types of cancer.
- Saroor A. Patel
- , Shoko Hirosue
- & Sakari Vanharanta
-
Article |
 Differential cofactor dependencies define distinct types of human enhancers
Differential cofactor dependencies define distinct types of human enhancers
The systematic categorization of human enhancers by their cofactor dependencies provides a conceptual framework to understand the sequence and chromatin diversity of enhancers and their roles in different gene-regulatory programmes.
- Christoph Neumayr
- , Vanja Haberle
- & Alexander Stark
-
Article |
 Compatibility rules of human enhancer and promoter sequences
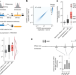
Compatibility rules of human enhancer and promoter sequences
A new high-throughput assay applied to 1,000 enhancers and 1,000 promoters in human cells reveals how different classes of enhancers and promoters control RNA expression.
- Drew T. Bergman
- , Thouis R. Jones
- & Jesse M. Engreitz
-
Article |
 OCA-T1 and OCA-T2 are coactivators of POU2F3 in the tuft cell lineage
OCA-T1 and OCA-T2 are coactivators of POU2F3 in the tuft cell lineage
The POU2F3–OCA-T complex is the master regulator of tuft cell identity and a prominent molecular vulnerability of tuft-cell-like small-cell lung cancer.
- Xiaoli S. Wu
- , Xue-Yan He
- & Christopher R. Vakoc
-
Article |
 Single-cell eQTL models reveal dynamic T cell state dependence of disease loci
Single-cell eQTL models reveal dynamic T cell state dependence of disease loci
A single-cell Poisson model is used to analyse eQTLs in memory T cells across continuous, dynamic cell states, revealing that the cell context is critical to understanding variation in eQTLs and their association with disease.
- Aparna Nathan
- , Samira Asgari
- & Soumya Raychaudhuri
-
Article |
 Transcriptional coupling of distant regulatory genes in living embryos
Transcriptional coupling of distant regulatory genes in living embryos
In Drosophila, there are extensive physical and functional associations of distant paralogous genes, including co-regulation by shared enhancers and co-transcriptional initiation over distances of nearly 250 kilobases.
- Michal Levo
- , João Raimundo
- & Michael S. Levine
-
Article |
 Decoding gene regulation in the fly brain
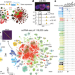
Decoding gene regulation in the fly brain
A chromatin accessibility atlas of 240,919 cells in the adult and developing Drosophila brain reveals 95,000 enhancers, which are integrated in cell-type specific enhancer gene regulatory networks and decoded into combinations of functional transcription factor binding sites using deep learning.
- Jasper Janssens
- , Sara Aibar
- & Stein Aerts
-
Article
| Open Access Cell-type specialization is encoded by specific chromatin topologies
Cell-type specialization is encoded by specific chromatin topologies
A new technique called immunoGAM, which combines genome architecture mapping (GAM) with immunoselection, enabled the discovery of specialized chromatin conformations linked to gene expression in specific cell populations from mouse brain tissues.
- Warren Winick-Ng
- , Alexander Kukalev
- & Ana Pombo
-
Article |
 BANP opens chromatin and activates CpG-island-regulated genes
BANP opens chromatin and activates CpG-island-regulated genes
BANP is identified as the transcription factor that binds the CGCG element in a DNA-methylation-dependent manner, opens chromatin and activates a class of essential CpG-island-regulated genes.
- Ralph S. Grand
- , Lukas Burger
- & Dirk Schübeler
-
Article |
 Defining genome architecture at base-pair resolution
Defining genome architecture at base-pair resolution
Micro Capture-C allows physical contacts to be determined at base-pair resolution, revealing that transcription factors have an important role in the maintenance of the contacts between enhancers and promoters.
- Peng Hua
- , Mohsin Badat
- & James O. J. Davies
-
Article |
 Enhancer release and retargeting activates disease-susceptibility genes
Enhancer release and retargeting activates disease-susceptibility genes
Disruption of a promoter can release its partner enhancer to activate other promoters in the same contact domain, and this process, named ‘enhancer release and retargeting’, can often lead to gene alterations that cause disease.
- Soohwan Oh
- , Jiaofang Shao
- & Michael G. Rosenfeld
-
Article |
 Interpreting type 1 diabetes risk with genetics and single-cell epigenomics
Interpreting type 1 diabetes risk with genetics and single-cell epigenomics
A genome-wide association study combined with single-cell epigenomics identifies risk loci for type 1 diabetes.
- Joshua Chiou
- , Ryan J. Geusz
- & Kyle J. Gaulton
-
Article |
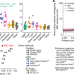 Genome-wide enhancer maps link risk variants to disease genes
Genome-wide enhancer maps link risk variants to disease genes
Mapping enhancer regulation across human cell types and tissues illuminates genome function and provides a resource to connect risk variants for common diseases to their molecular and cellular functions.
- Joseph Nasser
- , Drew T. Bergman
- & Jesse M. Engreitz
-
Article |
 Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator
Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator
The long non-coding RNA locus Maenli controls mouse limb development by regulating En1 activity, and the absence of the homolgous MAENLI locus is associated with severe congenital limb defects in humans.
- Lila Allou
- , Sara Balzano
- & Andrea Superti-Furga
-
Article
| Open Access Regulatory genomic circuitry of human disease loci by integrative epigenomics
Regulatory genomic circuitry of human disease loci by integrative epigenomics
The authors present EpiMap, a compendium that comprises 10,000 epigenomic maps across more than 800 biosamples for the annotation of genome-wide association study circuitry.
- Carles A. Boix
- , Benjamin T. James
- & Manolis Kellis
-
Article |
 m6A RNA methylation regulates the fate of endogenous retroviruses
m6A RNA methylation regulates the fate of endogenous retroviruses
A CRISPR screen in mouse embryonic stem cells shows that transcripts derived from endogenous retroviruses are destabilized by m6A RNA methylation.
- Tomasz Chelmicki
- , Emeline Roger
- & Deborah Bourc’his
-
Article |
 The landscape of RNA Pol II binding reveals a stepwise transition during ZGA
The landscape of RNA Pol II binding reveals a stepwise transition during ZGA
Binding of RNA polymerase II during zygotic genome activation in mouse embryos is determined using the newly developed method Stacc–seq.
- Bofeng Liu
- , Qianhua Xu
- & Wei Xie
-
Article |
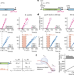 Functionally uncoupled transcription–translation in Bacillus subtilis
Functionally uncoupled transcription–translation in Bacillus subtilis
In Bacillus subtilis, unlike in Escherichia coli, transcription and translation of genes are not tightly coupled, and pioneering ribosomes lag substantially behind RNA polymerases.
- Grace E. Johnson
- , Jean-Benoît Lalanne
- & Gene-Wei Li
-
Article
| Open Access Spatiotemporal DNA methylome dynamics of the developing mouse fetus
Spatiotemporal DNA methylome dynamics of the developing mouse fetus
Analysis of 168 methylomes from 12 mouse tissues at 9 developmental stages sheds light on the epigenetic and regulatory landscape during mammalian fetal development.
- Yupeng He
- , Manoj Hariharan
- & Joseph R. Ecker
-
Article
| Open Access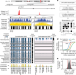 Global reference mapping of human transcription factor footprints
Global reference mapping of human transcription factor footprints
A high-density DNase I cleavage map from 243 human cell and tissue types provides a genome-wide, nucleotide-resolution map of human transcription factor footprints.
- Jeff Vierstra
- , John Lazar
- & John A. Stamatoyannopoulos
-
Article
| Open Access Occupancy maps of 208 chromatin-associated proteins in one human cell type
Occupancy maps of 208 chromatin-associated proteins in one human cell type
ChIP–seq and CETCh–seq data are used to analyse binding maps for 208 transcription factors and other chromatin-associated proteins in a single human cell type, providing a comprehensive catalogue of the transcription factor landscape and gene regulatory networks in these cells.
- E. Christopher Partridge
- , Surya B. Chhetri
- & Eric M. Mendenhall
-
Article |
 Structural cells are key regulators of organ-specific immune responses
Structural cells are key regulators of organ-specific immune responses
Structural cells implement a broad range of immune-regulatory functions beyond their roles as barriers and connective tissues, and they utilize an epigenetically encoded potential for immune gene activation in their rapid response to viral infection.
- Thomas Krausgruber
- , Nikolaus Fortelny
- & Christoph Bock
-
Article |
 Wapl repression by Pax5 promotes V gene recombination by Igh loop extrusion
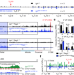
Wapl repression by Pax5 promotes V gene recombination by Igh loop extrusion
Pax5 regulates contraction of the immunoglobulin heavy chain (Igh) locus—an essential step in V(D)J recombination—by promoting chromatin loop extrusion via repression of Wapl expression.
- Louisa Hill
- , Anja Ebert
- & Meinrad Busslinger
-
Article |
 CRISPR screen in regulatory T cells reveals modulators of Foxp3
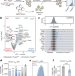
CRISPR screen in regulatory T cells reveals modulators of Foxp3
A CRISPR-based screening platform was used to identify previously uncharacterized genes that regulate the regulatory T cell-specific master transcription factor Foxp3, indicating that this screening method may be broadly applicable for the discovery of other genes involved in autoimmunity and immune responses to cancer.
- Jessica T. Cortez
- , Elena Montauti
- & Deyu Fang
-
Article |
 A satellite repeat-derived piRNA controls embryonic development of Aedes
A satellite repeat-derived piRNA controls embryonic development of Aedes
A conserved satellite repeat in the mosquito Aedes aegypti encodes PIWI-interacting RNAs that promote sequence-specific gene silencing in trans and have an essential role in embryonic development.
- Rebecca Halbach
- , Pascal Miesen
- & Ronald P. van Rij
-
Review Article |
 The promise and challenge of therapeutic genome editing
The promise and challenge of therapeutic genome editing
The scientific, technical and ethical aspects of using CRISPR technology for therapeutic applications in humans are discussed, highlighting both opportunities and challenges of this technology to treat, cure and prevent genetic disease.
- Jennifer A. Doudna
-
Article |
 SPEN integrates transcriptional and epigenetic control of X-inactivation
SPEN integrates transcriptional and epigenetic control of X-inactivation
The transcriptional repressor SPEN bridges the non-coding RNA Xist to transcription machinery, histone deacetylases and chromatin remodelling factors to initiate X-chromosome inactivation.
- François Dossin
- , Inês Pinheiro
- & Edith Heard
-
Article |
 Altered chromosomal topology drives oncogenic programs in SDH-deficient GISTs
Altered chromosomal topology drives oncogenic programs in SDH-deficient GISTs
Gastrointestinal stromal tumours can be initiated by gain-of-function mutations of the KIT or PDGFRA oncogenes but also by loss of the metabolic complex succinate dehydrogenase (SDH), which leads to DNA hypermethylation; this study shows that in SDH-deficient tumours, displacement of CTCF insulators by DNA methylation activates oncogene expression, illustrating how epigenetic alterations can drive oncogenic signalling in the absence of kinase mutations.
- William A. Flavahan
- , Yotam Drier
- & Bradley E. Bernstein
-
Article |
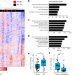 Regulation of lifespan by neural excitation and REST
Regulation of lifespan by neural excitation and REST
Studies of humans, mice and nematodes reveal a conserved role of neural activity and the transcription factor REST in extended longevity.
- Joseph M. Zullo
- , Derek Drake
- & Bruce A. Yankner
-
Letter |
 The RIPK4–IRF6 signalling axis safeguards epidermal differentiation and barrier function
The RIPK4–IRF6 signalling axis safeguards epidermal differentiation and barrier function
Signalling between the transcription factor IRF6 and the kinase RIPK4, in particular the phosphorylation of IRF6 by RIPK4, regulates epidermal differentiation and lipid metabolism, thereby maintaining the function of the epidermal barrier.
- Nina Oberbeck
- , Victoria C. Pham
- & Vishva M. Dixit
-
Letter |
 The fundamental role of chromatin loop extrusion in physiological V(D)J recombination
The fundamental role of chromatin loop extrusion in physiological V(D)J recombination
V(D)J recombination in B cells involves cohesin-mediated extrusion of chromatin loops to present DNA targets for cleavage and joining.
- Yu Zhang
- , Xuefei Zhang
- & Frederick W. Alt
-
Letter |
 Pol II phosphorylation regulates a switch between transcriptional and splicing condensates
Pol II phosphorylation regulates a switch between transcriptional and splicing condensates
RNA polymerase II with a hypophosphorylated C-terminal domain preferentially incorporates into mediator condensates, and with a hyperphosphorylated C-terminal domain into splicing-factor condensates, revealing phosphorylation as a regulatory mechanism in condensate preference.
- Yang Eric Guo
- , John C. Manteiga
- & Richard A. Young
-
Letter |
 Transcriptional cofactors display specificity for distinct types of core promoters
Transcriptional cofactors display specificity for distinct types of core promoters
A screen of 23 transcriptional cofactors for their ability to activate 72,000 candidate core promoters in Drosophila melanogaster identified distinct compatibility groups, providing insight into mechanisms that underlie the selective activation of transcriptional programs.
- Vanja Haberle
- , Cosmas D. Arnold
- & Alexander Stark

 Selfish conflict underlies RNA-mediated parent-of-origin effects
Selfish conflict underlies RNA-mediated parent-of-origin effects