Article
|
Open Access
Featured
-
-
Article
| Open Access Analysis of SARS-CoV-2 antibodies in COVID-19 convalescent blood using a coronavirus antigen microarray
Analysis of SARS-CoV-2 antibodies in COVID-19 convalescent blood using a coronavirus antigen microarray
COVID-19 diagnosis is commonly performed by PCR testing, however, serologic methods are more accurate and versatile for monitoring disease burden and epidemiology. Here the authors report a protein microarray with antigens from SARS-CoV-2, SARS-CoV, MERS-CoV as well as common human respiratory viruses.
- Rafael R. de Assis
- , Aarti Jain
- & Saahir Khan
-
Article
| Open Access Hydroxamic acid-modified peptide microarrays for profiling isozyme-selective interactions and inhibition of histone deacetylases
Hydroxamic acid-modified peptide microarrays for profiling isozyme-selective interactions and inhibition of histone deacetylases
Current histone microarrays cannot be used to directly study the transient interactions of histone deacetylases (HDACs). Here, the authors show that hydroxamic acid-modified microarrays can capture HDACs, provide insights into their substrate specificity, and serve to develop peptide inhibitors.
- Carlos Moreno-Yruela
- , Michael Bæk
- & Christian A. Olsen
-
Article
| Open Access An epigenetic gene silencing pathway selectively acting on transgenic DNA in the green alga Chlamydomonas
An epigenetic gene silencing pathway selectively acting on transgenic DNA in the green alga Chlamydomonas
Strong transgene suppression has been observed in Chlamydomonas reinhardtii, but the underlying mechanism is unknown. Here, the authors identify a sirtuin-type histone deacetylase that selectively acts on transgenic DNA to repress gene expression by assembling a repressive chromatin structure composed of deacetylated histones.
- Juliane Neupert
- , Sean D. Gallaher
- & Ralph Bock
-
Article
| Open Access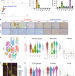 Functional compensation precedes recovery of tissue mass following acute liver injury
Functional compensation precedes recovery of tissue mass following acute liver injury
The liver possesses the ability to regenerate following sudden injury. Here, the authors use single-cell RNA-sequencing and in situ transcriptional analyses to identify a new phase of liver regeneration in mice aimed at maintaining essential functions throughout the regenerative process.
- Chad M. Walesky
- , Kellie E. Kolb
- & Wolfram Goessling
-
Article
| Open Access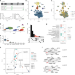 Transcriptomic analysis links diverse hypothalamic cell types to fibroblast growth factor 1-induced sustained diabetes remission
Transcriptomic analysis links diverse hypothalamic cell types to fibroblast growth factor 1-induced sustained diabetes remission
In rodent models of type 2 diabetes, sustained remission of hyperglycemia can be induced by FGF1 action in the mediobasal hypothalamus. Here, the authors show that FGF1-injection is followed by marked changes in glial cell populations and that the sustained glycemic response is dependent on intact melanocortin signaling.
- Marie A. Bentsen
- , Dylan M. Rausch
- & Tune H. Pers
-
Article
| Open Access Representation of features as images with neighborhood dependencies for compatibility with convolutional neural networks
Representation of features as images with neighborhood dependencies for compatibility with convolutional neural networks
Convolutional Neural Networks (CNN) are often unsuitable for predictive modeling involving nonimage based biological features. Here, the authors present a mapping termed REFINED to represent high dimensional vectors as compact images with spatial correlation that makes it compatible with CNN based learning.
- Omid Bazgir
- , Ruibo Zhang
- & Ranadip Pal
-
Article
| Open Access Selective flexible packaging pathways of the segmented genome of influenza A virus
Selective flexible packaging pathways of the segmented genome of influenza A virus
The mechanism underlying packaging of the 8 segments of the influenza virus genome into virions is not well understood. Here, the authors use a multiplexed FISH assay to monitor the 8 segments in parallel in infected cells suggesting bundling routes during the packaging process.
- Ivan Haralampiev
- , Simon Prisner
- & Andreas Herrmann
-
Comment
| Open Access Single cell transcriptomics comes of age
Single cell transcriptomics comes of age
Single cell transcriptomics technologies have vast potential in advancing our understanding of biology and disease. Here, Sarah Aldridge and Sarah Teichmann review the last decade of technological advancements in single-cell transcriptomics and highlight some of the recent discoveries enabled by this technology.
- Sarah Aldridge
- & Sarah A. Teichmann
-
Article
| Open Access Multiplexed single-cell transcriptional response profiling to define cancer vulnerabilities and therapeutic mechanism of action
Multiplexed single-cell transcriptional response profiling to define cancer vulnerabilities and therapeutic mechanism of action
Large-scale screens of chemical and genetic vulnerabilities in cancer are typically limited to simple readouts of cell viability. Here, the authors develop a method for profiling post-perturbation transcriptional responses across large pools of cancer cell lines, enabling deep characterization of shared and context-specific responses.
- James M. McFarland
- , Brenton R. Paolella
- & Aviad Tsherniak
-
Article
| Open Access Multiscale causal networks identify VGF as a key regulator of Alzheimer’s disease
Multiscale causal networks identify VGF as a key regulator of Alzheimer’s disease
To investigate the molecular foundation of sporadic Alzheimer’s disease (AD), Beckmann et al. constructed multiscale causal networks on a large human AD multi-omics dataset, detecting AD-associated networks and their top predicted regulator, VGF, with extensive validation in the 5xFAD mouse model.
- Noam D. Beckmann
- , Wei-Jye Lin
- & Eric E. Schadt
-
Article
| Open Access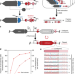 Large-scale DNA-based phenotypic recording and deep learning enable highly accurate sequence-function mapping
Large-scale DNA-based phenotypic recording and deep learning enable highly accurate sequence-function mapping
Current methods to generate sequence-function data at large scale are either technically complex or limited to specific applications. Here the authors introduce DNA-based phenotypic recording to overcome these limitations and enable deep learning for accurate prediction of function from sequence.
- Simon Höllerer
- , Laetitia Papaxanthos
- & Markus Jeschek
-
Article
| Open Access SARS-CoV-2 proteome microarray for global profiling of COVID-19 specific IgG and IgM responses
SARS-CoV-2 proteome microarray for global profiling of COVID-19 specific IgG and IgM responses
Currently very little is known about how our immune system responds to SARS-CoV-2 infection. Here the authors generate a SARS-CoV-2 proteome microarray for profiling of IgG and IgM responses to COVID-19 in patients and find significant responses to ORF9b and NSP5, as well as the S1 and N proteins.
- He-wei Jiang
- , Yang Li
- & Sheng-ce Tao
-
Article
| Open Access Comprehensive characterization of claudin-low breast tumors reflects the impact of the cell-of-origin on cancer evolution
Comprehensive characterization of claudin-low breast tumors reflects the impact of the cell-of-origin on cancer evolution
Claudin-low tumors are a rare aggressive subtype of breast cancers. In this study, the authors use a multiomics approach to demonstrate that these tumors are heterogeneous and comprise three main subgroups that emerge from different evolutionary processes.
- Roxane M. Pommier
- , Amélien Sanlaville
- & Alain Puisieux
-
Article
| Open Access Mapping effector genes at lupus GWAS loci using promoter Capture-C in follicular helper T cells
Mapping effector genes at lupus GWAS loci using promoter Capture-C in follicular helper T cells
T cells are a major cell type involved in systemic lupus erythematosus (SLE). Here, the authors use promoter capture-C and ATAC-seq in human follicular T helper cells to identify SLE genes distant from GWAS loci (via 3D interaction) and validate the function of key regulatory elements and genes in vitro.
- Chun Su
- , Matthew E. Johnson
- & Andrew D. Wells
-
Article
| Open Access Substrate specificity of the TRAMP nuclear surveillance complexes
Substrate specificity of the TRAMP nuclear surveillance complexes
During nuclear surveillance in yeast and human cells, the RNA exosome functions together with the TRAMP complexes. Here, the authors defined the protein composition of the TRAMP complexes and identified specific RNA binding sites for the different TRAMP components.
- Clémentine Delan-Forino
- , Christos Spanos
- & David Tollervey
-
Article
| Open Access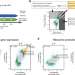 A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites
A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites
The unfolded protein response (UPR) is a stress response pathway implicated in numerous diseases and chemotherapy resistance. Here, the authors define the UPR regulon with a multi-omics strategy, uncovering changes to mitochondrial one-carbon metabolism and concomitant resistance to folate-based therapeutics.
- Stefan Reich
- , Chi D. L. Nguyen
- & Jan Medenbach
-
Article
| Open Access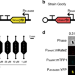 Single-cell bacterial transcription measurements reveal the importance of dimethylsulfoniopropionate (DMSP) hotspots in ocean sulfur cycling
Single-cell bacterial transcription measurements reveal the importance of dimethylsulfoniopropionate (DMSP) hotspots in ocean sulfur cycling
DMSP is a ubiquitous organosulfur compound in the ocean that, once degraded by bacteria, plays key roles in global biogeochemical cycles and climate regulation. Here, the authors use single-cell measurements of transcription to investigate the intricate dynamics of bacterial DMSP degradation.
- Cherry Gao
- , Vicente I. Fernandez
- & Roman Stocker
-
Article
| Open Access Time-lapse single-cell transcriptomics reveals modulation of histone H3 for dormancy breaking in fission yeast
Time-lapse single-cell transcriptomics reveals modulation of histone H3 for dormancy breaking in fission yeast
Asynchronicity in fungal spore germination makes transcriptomic analysis of the process challenging. Here, the authors assay single cell transcriptomes of germinating yeast cells and find that one of the histone H3 genes shows fluctuating expression, disruption of which causes germination defects.
- Hayato Tsuyuzaki
- , Masahito Hosokawa
- & Masamitsu Sato
-
Article
| Open Access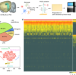 Identification of region-specific astrocyte subtypes at single cell resolution
Identification of region-specific astrocyte subtypes at single cell resolution
Astrocytes are a major cell type in the central nervous system. Using single cell transcriptome sequencing, the authors identify multiple astrocyte subtypes in the adult mouse CNS, which map to distinct spatial locations and show correlations to cell morphology and physiology.
- Mykhailo Y. Batiuk
- , Araks Martirosyan
- & Matthew G. Holt
-
Article
| Open Access Transient genome-wide interactions of the master transcription factor NLP7 initiate a rapid nitrogen-response cascade
Transient genome-wide interactions of the master transcription factor NLP7 initiate a rapid nitrogen-response cascade
Conventional methods cannot reveal transient transcription factors (TFs) and targets interactions. Here, Alvarez et al. capture both stable and transient TF-target interactions by time-series ChIP-seq and/or DamID-seq in a cell-based TF perturbation system and show NLP7 as a master TF to initiate a rapid nitrogen-response cascade.
- José M. Alvarez
- , Anna-Lena Schinke
- & Gloria M. Coruzzi
-
Article
| Open Access Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Small non-coding RNAs contribute to the regulation of aging. Here the authors identify a small nucleolar RNA, the snoRNA jouvence, which extends the lifespan of fruit flies through its function in the gut, and is conserved in humans.
- Stéphanie Soulé
- , Lucille Mellottée
- & Jean-René Martin
-
Article
| Open Access Non-invasive characterization of human bone marrow stimulation and reconstitution by cell-free messenger RNA sequencing
Non-invasive characterization of human bone marrow stimulation and reconstitution by cell-free messenger RNA sequencing
Circulating cell-free mRNA holds great promise as a non-invasive diagnostic biomarker. Here the authors show that cell-free mRNA captures transcripts from the bone marrow and can be used to non-invasively monitor dynamic changes in bone marrow physiology.
- Arkaitz Ibarra
- , Jiali Zhuang
- & Michael Nerenberg
-
Article
| Open Access Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
Single-nuclei RNA-seq on human retinal tissue provides improved transcriptome profiling
The retina is a heterogeneous tissue composed of multiple cell types. Via single-nuclei RNA sequencing on human neural retinal tissue, the authors characterise the transcriptome profile for individual cell types in the human retina.
- Qingnan Liang
- , Rachayata Dharmat
- & Rui Chen
-
Article
| Open Access A Bayesian mixture model for the analysis of allelic expression in single cells
A Bayesian mixture model for the analysis of allelic expression in single cells
Allele-specific expression at single-cell resolution can reveal stochastic and dynamic features of gene expression in greater detail. The authors propose scBASE, a soft zero-and-one inflated model that improves estimation of cellular allelic proportions by pooling information across cells.
- Kwangbom Choi
- , Narayanan Raghupathy
- & Gary A. Churchill
-
Article
| Open Access Microbe-host interplay in atopic dermatitis and psoriasis
Microbe-host interplay in atopic dermatitis and psoriasis
Atopic dermatitis (AD) and psoriasis (PSO) are associated with dysbiosis. Here, by analyses of skin microbiome and host transcriptome of AD and PSO patients, the authors find distinct microbial and disease-related gene transcriptomic signatures that differentiate both diseases.
- Nanna Fyhrquist
- , Gareth Muirhead
- & Harri Alenius
-
Article
| Open Access Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Synthetic RNA-based devices can dynamically control a wide range of processes. Here the authors develop a quantitative and high-throughput mammalian cell-based RNA-seq assay to efficiently engineer ribozyme switches.
- Joy S. Xiang
- , Matias Kaplan
- & Christina D. Smolke
-
Article
| Open Access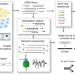 High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes
Single cell RNA sequencing generates short reads from one end of a template, providing incomplete transcript coverage and limiting identification of diverse sequences such as antigen receptors. Here the authors combine long read nanopore sequencing with short read profiling of barcoded libraries to generate full-length antigen receptor sequences.
- Mandeep Singh
- , Ghamdan Al-Eryani
- & Alexander Swarbrick
-
Article
| Open Access Circuit asymmetries underlie functional lateralization in the mouse auditory cortex
Circuit asymmetries underlie functional lateralization in the mouse auditory cortex
The left hemisphere of the brain is especially involved in processing social vocalizations and (in humans) language, but the mechanisms of this lateralization of function are unclear. Here, the authors compared left and right auditory cortex in mice and show lateralized, experience-dependent circuit-motifs.
- Robert B. Levy
- , Tiemo Marquarding
- & Hysell V. Oviedo
-
Article
| Open Access Hydro-Seq enables contamination-free high-throughput single-cell RNA-sequencing for circulating tumor cells
Hydro-Seq enables contamination-free high-throughput single-cell RNA-sequencing for circulating tumor cells
Transcriptome analysis of circulating tumor cells (CTCs) provides insights into monitoring target therapeutics and underlying tumor metastasis. Here the authors present Hydro-Seq, a contamination-free high-throughput hydrodynamic scRNA-seq barcoding technique for rare CTCs.
- Yu-Heng Cheng
- , Yu-Chih Chen
- & Euisik Yoon
-
Article
| Open Access Targeted removal of epigenetic barriers during transcriptional reprogramming
Targeted removal of epigenetic barriers during transcriptional reprogramming
Master transcription factors dominantly direct cell fate and barriers ensuring their tissue specific silencing are not clearly defined. Here, the authors demonstrate that inefficient targeted transactivation of Sox1 in neural progenitor cells is surmountable through targeted promoter demethylation using dCas9-Tet1.
- Valentin Baumann
- , Maximilian Wiesbeck
- & Stefan H. Stricker
-
Article
| Open Access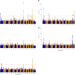 GWAS of bone size yields twelve loci that also affect height, BMD, osteoarthritis or fractures
GWAS of bone size yields twelve loci that also affect height, BMD, osteoarthritis or fractures
Size and shape of bones are important for height and body shape. Here, Styrkarsdottir et al identify 12 loci in a GWAS for bone area derived from DXA scans and show that these loci associate with other bone-related phenotypes including osteoarthritis, height, bone mineral density and risk of hip fracture.
- Unnur Styrkarsdottir
- , Olafur A. Stefansson
- & Kari Stefansson
-
Article
| Open Access Global identification of functional microRNA-mRNA interactions in Drosophila
Global identification of functional microRNA-mRNA interactions in Drosophila
MicroRNAs are mediators of post-transcriptional gene expression silencing. Here authors provide a transcriptome-wide map of miRNA target sites in Drosophila.
- Hans-Hermann Wessels
- , Svetlana Lebedeva
- & Uwe Ohler
-
Article
| Open Access Multiplexed Cas9 targeting reveals genomic location effects and gRNA-based staggered breaks influencing mutation efficiency
Multiplexed Cas9 targeting reveals genomic location effects and gRNA-based staggered breaks influencing mutation efficiency
Designing effective genome engineering strategies requires an understanding of the impact that genomic locus has on CRISPR-Cas9 activity. Here the authors use TRIP integrations to profile editing outcomes genome-wide and observe that gRNA sequence influences the structure of the double strand break.
- Santiago Gisler
- , Joana P. Gonçalves
- & Maarten van Lohuizen
-
Article
| Open Access A tetracycline-dependent ribozyme switch allows conditional induction of gene expression in Caenorhabditis elegans
A tetracycline-dependent ribozyme switch allows conditional induction of gene expression in Caenorhabditis elegans
Tools for conditional induction of gene expression in C. elegans are limited compared to other organisms. Here the authors present a tetracycline-dependent ribozyme that allows conditional control of a gene of interest.
- Lena A. Wurmthaler
- , Monika Sack
- & Martin Gamerdinger
-
Article
| Open Access Expression of novel long noncoding RNAs defines virus-specific effector and memory CD8+ T cells
Expression of novel long noncoding RNAs defines virus-specific effector and memory CD8+ T cells
Long noncoding RNA (lncRNA) genes do not encode protein products yet are emerging as key regulators of cellular processes such as transcription and translation. Here, by examining lncRNA profiles from human and mouse CD8 T cells, the authors show that stages of CD8+ T cell differentiation are defined by expression of lncRNA genes.
- William H. Hudson
- , Nataliya Prokhnevska
- & Haydn T. Kissick
-
Article
| Open Access A computational framework to study sub-cellular RNA localization
A computational framework to study sub-cellular RNA localization
Automated analysis of RNA localisation in smFISH data has been elusive. Here, the authors simulate and use a large dataset of images to design and validate a framework for highly accurate classification of sub-cellular RNA localisation patterns from smFISH experiments.
- Aubin Samacoits
- , Racha Chouaib
- & Florian Mueller
-
Article
| Open Access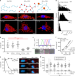 Enhanced mRNA FISH with compact quantum dots
Enhanced mRNA FISH with compact quantum dots
FISH-based techniques to image and count mRNA in single cells can be limited by the photophysical properties of organic dyes. Here the authors develop photostable quantum dot FISH probes for multiplexed imaging.
- Yang Liu
- , Phuong Le
- & Andrew M. Smith
-
Article
| Open Access Single cell molecular alterations reveal target cells and pathways of concussive brain injury
Single cell molecular alterations reveal target cells and pathways of concussive brain injury
Traumatic brain injury (TBI) affects the hippocampus and can lead to neurological and psychiatric disorders. Here, the authors perform single-cell RNA sequencing to reveal molecular pathways across a range of cell types affected during TBI.
- Douglas Arneson
- , Guanglin Zhang
- & Xia Yang
-
Article
| Open Access microCLIP super learning framework uncovers functional transcriptome-wide miRNA interactions
microCLIP super learning framework uncovers functional transcriptome-wide miRNA interactions
AGO-PAR-CLIP is widely used for high-throughput miRNA target characterization. Here, the authors show that the previously neglected non-T-to-C clusters denote functional miRNA binding events, and develop microCLIP, a super learning framework that accurately detects miRNA interactions.
- Maria D. Paraskevopoulou
- , Dimitra Karagkouni
- & Artemis G. Hatzigeorgiou
-
Article
| Open Access Endocrine lineage biases arise in temporally distinct endocrine progenitors during pancreatic morphogenesis
Endocrine lineage biases arise in temporally distinct endocrine progenitors during pancreatic morphogenesis
Endocrine progenitors form early in pancreatic development but the diversity of this cell population is unclear. Here, the authors use single cell RNA sequencing of the mouse pancreas at e14.5 and e16.5 to show that endocrine progenitors are temporally distinct and those formed later are more likely to become beta cells
- Marissa A. Scavuzzo
- , Matthew C. Hill
- & Malgorzata Borowiak
-
Article
| Open Access Genome-wide real-time in vivo transcriptional dynamics during Plasmodium falciparum blood-stage development
Genome-wide real-time in vivo transcriptional dynamics during Plasmodium falciparum blood-stage development
Transcriptomic analysis often doesn’t differentiate between newly synthesized and stabilized mRNAs. Using rapid 4-thiouracil incorporation, Painter et al. here define genome-wide active transcription throughout Plasmodium blood-stage developmental stages and identify associated regulatory DNA sequence motifs.
- Heather J. Painter
- , Neo Christopher Chung
- & Manuel Llinás
-
Article
| Open Access A mixed antagonistic/synergistic miRNA repression model enables accurate predictions of multi-input miRNA sensor activity
A mixed antagonistic/synergistic miRNA repression model enables accurate predictions of multi-input miRNA sensor activity
MicroRNAs (miRNAs) are important post-transcriptional regulators of gene expression but many quantitative aspects of miRNA biology remain to be elucidated. Based on a library of miRNA sensors, the authors quantify miRNA regulation at single cell level and develop a model to predict miRNA target interactions.
- Jeremy J. Gam
- , Jonathan Babb
- & Ron Weiss
-
Article
| Open Access A neuronal basis for fear discrimination in the lateral amygdala
A neuronal basis for fear discrimination in the lateral amygdala
When perceiving new stimuli, organisms need to distinguish between threats versus harmless stimuli. Here, the authors find a set of cells in the lateral amygdala that is required to discriminate or generalize new auditory stimuli based on similarity to previously fear-associate sounds.
- Anna Grosso
- , Giulia Santoni
- & Benedetto Sacchetti
-
Article
| Open Access Transcriptomic signatures of NK cells suggest impaired responsiveness in HIV-1 infection and increased activity post-vaccination
Transcriptomic signatures of NK cells suggest impaired responsiveness in HIV-1 infection and increased activity post-vaccination
Natural killer (NK) cells are important for eliminating cells under stress or infected by virus, and may have a function in anti-HIV immunity. Here the authors show that different NK-activating stimuli induce distinct transcriptional fingerprints in human NK cells that are analogous to changes caused by HIV vaccination or chronic infection.
- Margaret C. Costanzo
- , Dohoon Kim
- & Michael A. Eller
-
Article
| Open Access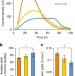 Electrochemically active bacteria sense electrode potentials for regulating catabolic pathways
Electrochemically active bacteria sense electrode potentials for regulating catabolic pathways
Whether electrochemically active bacteria (EAB) can gain energy according to electrode potentials is still unclear. Here, the authors show through transcriptome and deletion mutant analyses that EAB can sense electrode potentials by the Arc system and activate NADH-dependent catabolic pathway to generate ATP.
- Atsumi Hirose
- , Takuya Kasai
- & Atsushi Kouzuma
-
Article
| Open Access Unraveling the determinants of microRNA mediated regulation using a massively parallel reporter assay
Unraveling the determinants of microRNA mediated regulation using a massively parallel reporter assay
MiRNAs are known regulators of gene expression. Here the authors perform a large-scale massively parallel reporter assay to investigate the effect of a large number of designed 3′ UTR sequences on reporter expression and asses how miRNA regulatory elements features affect miRNA mediated repression.
- Ilya Vainberg Slutskin
- , Shira Weingarten-Gabbay
- & Eran Segal
-
Article
| Open Access Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Efficient transgenesis and annotated genome sequence of the regenerative flatworm model Macrostomum lignano
Regeneration capable flatworms have emerged as powerful models for studying stem cell biology and patterning, however their study has been hindered by the lack of transgenesis methods. Here, the authors describe a transgenesis method for Macrostomum lignano, as well as a new annotated genome sequence.
- Jakub Wudarski
- , Daniil Simanov
- & Eugene Berezikov
-
Article
| Open Access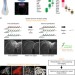 Identification of a neural crest stem cell niche by Spatial Genomic Analysis
Identification of a neural crest stem cell niche by Spatial Genomic Analysis
Neural crest cells arise within the central nervous system, then migrate and contribute to a variety of cell types. Here, the authors use multiplex transcript analysis at single cell resolution to define neural crest and neural subpopulations within the avian neural tube, including a neural crest stem cell niche.
- Antti Lignell
- , Laura Kerosuo
- & Marianne E. Bronner
-
Article
| Open Access RNA localization is a key determinant of neurite-enriched proteome
RNA localization is a key determinant of neurite-enriched proteome
Subcellular localization of RNAs and proteins is important for polarized cells such as neurons. Here the authors differentiate mouse embryonic stem cells into neurons, and analyze the local transcriptome, proteome, and translated transcriptome in their cell bodies and neurites, providing a unique resource for future studies on neuronal polarity.
- Alessandra Zappulo
- , David van den Bruck
- & Marina Chekulaeva

 Tailoring the resolution of single-cell RNA sequencing for primary cytotoxic T cells
Tailoring the resolution of single-cell RNA sequencing for primary cytotoxic T cells