Article
|
Open Access
Featured
-
-
Article
| Open Access Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Leveraging the fact that eukaryotic genomes are organized into gene modules, FISHnCHIPs images multiple co-expressed genes simultaneously for sensitive and high throughput profiling of gene programs and cell types in tissues.
- Xinrui Zhou
- , Wan Yi Seow
- & Kok Hao Chen
-
Article
| Open Access Predictive and robust gene selection for spatial transcriptomics
Predictive and robust gene selection for spatial transcriptomics
Gene selection for spatial transcriptomics is currently not optimal. Here the authors report PERSIST, a flexible deep learning framework that uses existing scRNA-seq data to identify gene targets for spatial transcriptomics; they show this allows you to capture more information with fewer genes.
- Ian Covert
- , Rohan Gala
- & Su-In Lee
-
Article
| Open Access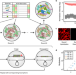 Spatial profiling of microbial communities by sequential FISH with error-robust encoding
Spatial profiling of microbial communities by sequential FISH with error-robust encoding
Spatial analysis of microbiomes at single cell resolution is challenging. Here the authors report a highly multiplexed method for spatial profiling, sequential error-robust fluorescence in situ hybridisation (SEER-FISH), and show that this allows mapping of microbial communities at micron-scale.
- Zhaohui Cao
- , Wenlong Zuo
- & Lei Dai
-
Article
| Open Access Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Methods for jointly analysing the different spatial data modalities in 3D are lacking. Here the authors report the computational framework STACI (Spatial Transcriptomic data using over-parameterized graph-based Autoencoders with Chromatin Imaging data) which they apply to an Alzheimer’s disease mouse model.
- Xinyi Zhang
- , Xiao Wang
- & Caroline Uhler
-
Article
| Open Access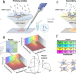 Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis
Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis
Spatial-omics methods with ease-of-use and high multiplexing are in demand. Here the authors report Multi Omic Single-scan Assay with Integrated Combinatorial Analysis (MOSAICA) which uses Spectral and Fluorescence Lifetime Imaging and Microscopy; they apply this to co-detection of mRNA and protein.
- Tam Vu
- , Alexander Vallmitjana
- & Weian Zhao
-
Article
| Open Access Single-nucleus RNA-seq2 reveals functional crosstalk between liver zonation and ploidy
Single-nucleus RNA-seq2 reveals functional crosstalk between liver zonation and ploidy
Single-cell RNA-seq reveals the cellular heterogeneity in development and disease. Here the authors present a single-nucleus RNA-seq2 that allows deep characterization of nuclei isolated from frozen archived tissues, apply it for transcriptional profiling of individual hepatocytes, and determine a functional crosstalk between liver zonation and ploidy.
- M. L. Richter
- , I. K. Deligiannis
- & C. P. Martinez-Jimenez
-
Article
| Open Access Cell segmentation-free inference of cell types from in situ transcriptomics data
Cell segmentation-free inference of cell types from in situ transcriptomics data
Inaccurate cell segmentation has been the major problem for cell-type identification and tissue characterization of the in situ spatially resolved transcriptomics data. Here we show a robust cell segmentation-free computational framework (SSAM), for identifying cell types and tissue domains in 2D and 3D.
- Jeongbin Park
- , Wonyl Choi
- & Naveed Ishaque
-
Article
| Open Access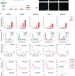 Melanoma subpopulations that rapidly escape MAPK pathway inhibition incur DNA damage and rely on stress signalling
Melanoma subpopulations that rapidly escape MAPK pathway inhibition incur DNA damage and rely on stress signalling
BRAF inhibitors are used to treat late-stage melanoma patients harbouring BRAF mutations. Here the authors track the responses of single melanoma cells to BRAF inhibitors and show that a subset of cells rapidly escapes drug via non-genetic mechanisms and incurs DNA damage.
- Chen Yang
- , Chengzhe Tian
- & Sabrina L. Spencer
-
Article
| Open Access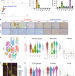 Functional compensation precedes recovery of tissue mass following acute liver injury
Functional compensation precedes recovery of tissue mass following acute liver injury
The liver possesses the ability to regenerate following sudden injury. Here, the authors use single-cell RNA-sequencing and in situ transcriptional analyses to identify a new phase of liver regeneration in mice aimed at maintaining essential functions throughout the regenerative process.
- Chad M. Walesky
- , Kellie E. Kolb
- & Wolfram Goessling
-
Article
| Open Access Selective flexible packaging pathways of the segmented genome of influenza A virus
Selective flexible packaging pathways of the segmented genome of influenza A virus
The mechanism underlying packaging of the 8 segments of the influenza virus genome into virions is not well understood. Here, the authors use a multiplexed FISH assay to monitor the 8 segments in parallel in infected cells suggesting bundling routes during the packaging process.
- Ivan Haralampiev
- , Simon Prisner
- & Andreas Herrmann
-
Article
| Open Access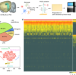 Identification of region-specific astrocyte subtypes at single cell resolution
Identification of region-specific astrocyte subtypes at single cell resolution
Astrocytes are a major cell type in the central nervous system. Using single cell transcriptome sequencing, the authors identify multiple astrocyte subtypes in the adult mouse CNS, which map to distinct spatial locations and show correlations to cell morphology and physiology.
- Mykhailo Y. Batiuk
- , Araks Martirosyan
- & Matthew G. Holt
-
Article
| Open Access A computational framework to study sub-cellular RNA localization
A computational framework to study sub-cellular RNA localization
Automated analysis of RNA localisation in smFISH data has been elusive. Here, the authors simulate and use a large dataset of images to design and validate a framework for highly accurate classification of sub-cellular RNA localisation patterns from smFISH experiments.
- Aubin Samacoits
- , Racha Chouaib
- & Florian Mueller
-
Article
| Open Access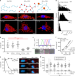 Enhanced mRNA FISH with compact quantum dots
Enhanced mRNA FISH with compact quantum dots
FISH-based techniques to image and count mRNA in single cells can be limited by the photophysical properties of organic dyes. Here the authors develop photostable quantum dot FISH probes for multiplexed imaging.
- Yang Liu
- , Phuong Le
- & Andrew M. Smith
-
Article
| Open Access A neuronal basis for fear discrimination in the lateral amygdala
A neuronal basis for fear discrimination in the lateral amygdala
When perceiving new stimuli, organisms need to distinguish between threats versus harmless stimuli. Here, the authors find a set of cells in the lateral amygdala that is required to discriminate or generalize new auditory stimuli based on similarity to previously fear-associate sounds.
- Anna Grosso
- , Giulia Santoni
- & Benedetto Sacchetti
-
Article
| Open Access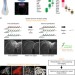 Identification of a neural crest stem cell niche by Spatial Genomic Analysis
Identification of a neural crest stem cell niche by Spatial Genomic Analysis
Neural crest cells arise within the central nervous system, then migrate and contribute to a variety of cell types. Here, the authors use multiplex transcript analysis at single cell resolution to define neural crest and neural subpopulations within the avian neural tube, including a neural crest stem cell niche.
- Antti Lignell
- , Laura Kerosuo
- & Marianne E. Bronner
-
Article
| Open Access Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes
Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes
The spatial organization of the genome within the nucleus impacts many processes. Here the authors combine oligo-based DNA FISH with single-molecule super-resolution microscopy to image single-copy genomic regions and, taking advantage of SNPs, distinguish allelic regions of homologous chromosomes.
- Brian J. Beliveau
- , Alistair N. Boettiger
- & Chao-ting Wu
-
Article |
 High-throughput detection of miRNAs and gene-specific mRNA at the single-cell level by flow cytometry
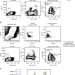
High-throughput detection of miRNAs and gene-specific mRNA at the single-cell level by flow cytometry
Flow cytometry allows high-throughput analysis of multiple proteins in individual cells, but relies on availability of antibodies. Here Porichis et al.report a sensitive method for multi-parameter flow cytometric and imaging detection of proteins together with mRNA or miRNA at the single-cell level.
- Filippos Porichis
- , Meghan G. Hart
- & Daniel E. Kaufmann

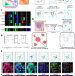 Mapping human tissues with highly multiplexed RNA in situ hybridization
Mapping human tissues with highly multiplexed RNA in situ hybridization