Featured
-
-
Article
| Open Access DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy
DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy
5-hydroxymethylation of cysteine (5-hmC) plays a role in epigenetic regulation. Here the authors analyse the hydroxymethylome in embryonic, neonatal, adult and hypertrophic mouse cardiomyocytes and show that the dynamic modulation of hydroxymethylated DNA is important for cardiomyocyte gene expression programming in heart development and failure.
- Carolina M. Greco
- , Paolo Kunderfranco
- & Gianluigi Condorelli
-
Article
| Open Access Extra-coding RNAs regulate neuronal DNA methylation dynamics
Extra-coding RNAs regulate neuronal DNA methylation dynamics
DNA methylation in the brain is a dynamic process, but gene-specific regulation of this process is poorly understood. Here, Day and colleagues show that extra-coding RNAs interact with DNA methyltransferases and regulate neuronal DNA methylation to control gene expression in locus-specific manner in neurons.
- Katherine E. Savell
- , Nancy V. N. Gallus
- & Jeremy J. Day
-
Article
| Open Access Hypoxia causes transgenerational impairments in reproduction of fish
Hypoxia causes transgenerational impairments in reproduction of fish
Hypoxia has diverse effects on aquatic life. Wang et al.show that reproductive defects resulting from hypoxia are epigenetically heritable in Japanese rice fish, and that this intergenerational inheritance is accompanied by differential methylation and gene expression in sperm.
- Simon Yuan Wang
- , Karen Lau
- & Rudolf Shiu-Sun Wu
-
Article
| Open Access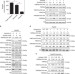 Acetate functions as an epigenetic metabolite to promote lipid synthesis under hypoxia
Acetate functions as an epigenetic metabolite to promote lipid synthesis under hypoxia
Cancer cells under stress use acetate to maintain the acetyl-CoA pool and fuel lipid biosynthesis. Here, the authors show that acetate also promotes de novo lipid synthesis by increasing histone acetylation at the promoters of lipogenic enzymes ACACA and FASN, thus inducing their expression.
- Xue Gao
- , Shu-Hai Lin
- & Qun-Ying Lei
-
Article
| Open Access Structural basis for the recognition of guide RNA and target DNA heteroduplex by Argonaute
Structural basis for the recognition of guide RNA and target DNA heteroduplex by Argonaute
Argonaute proteins are important in the silencing machinery with some regulatory RNAs. Here, the authors solve the structure of an argonaute protein in complex with both the guide RNA and target DNA and propose a mechanism for their recognition.
- Tomohiro Miyoshi
- , Kosuke Ito
- & Toshio Uchiumi
-
Article
| Open Access MTHFD1 controls DNA methylation in Arabidopsis
MTHFD1 controls DNA methylation in Arabidopsis
DNA methylation contributes to transcriptional silencing. Here, Groth et al.show that mutant plants defective in MTHFD1, an enzyme involved in folate metabolism, have a DNA hypomethylation phenotype highlighting the link between one-carbon metabolism and DNA methylation, which is mediated by SAM as a common methyl donor.
- Martin Groth
- , Guillaume Moissiard
- & Steven E. Jacobsen
-
Article
| Open Access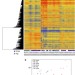 Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Rheumatoid arthritis is an inflammatory disease that selectively affects different joints. Here the authors show that gene expression and DNA methylation patterns of fibroblast-like synoviocytes differ between hip and knee joints in patients with RA, thus providing epigenetic and transcriptomic evidence for this anatomic selectivity of inflammation.
- Rizi Ai
- , Deepa Hammaker
- & Wei Wang
-
Article
| Open Access The Arabidopsis acetylated histone-binding protein BRAT1 forms a complex with BRP1 and prevents transcriptional silencing
The Arabidopsis acetylated histone-binding protein BRAT1 forms a complex with BRP1 and prevents transcriptional silencing
Transposons and repetitive sequences are typically subject to transcription silencing. Here, Zhang et al. find that the bromodomain-containing protein BRAT1 forms a complex with BRP1, recognizes histone acetylation and acts to prevent transcriptional silencing in Arabidopsis.
- Cui-Jun Zhang
- , Xiao-Mei Hou
- & Xin-Jian He
-
Article
| Open Access CRISPR-dCas9 and sgRNA scaffolds enable dual-colour live imaging of satellite sequences and repeat-enriched individual loci
CRISPR-dCas9 and sgRNA scaffolds enable dual-colour live imaging of satellite sequences and repeat-enriched individual loci
The ability to target Cas9 to specific genomic loci offers the potential for applications beyond genome editing. Here the authors use deactivated Cas9 and chimeric guide RNAs to visualise individual repetitive loci in living cells.
- Yi Fu
- , Pedro P. Rocha
- & Jane A. Skok
-
Article
| Open Access Epigenetic reprogramming of fallopian tube fimbriae in BRCA mutation carriers defines early ovarian cancer evolution
Epigenetic reprogramming of fallopian tube fimbriae in BRCA mutation carriers defines early ovarian cancer evolution
Women with germline variants in BRCA genes are predisposed to ovarian cancer. In this study, the authors demonstrate that fimbrial tissue from the ovary, the site of ovarian cancer, in BRCAmutant carriers contains marked DNA methylation changes compared with the proximal region of the ovary.
- Thomas E. Bartlett
- , Kantaraja Chindera
- & Martin Widschwendter
-
Article
| Open Access Caloric restriction blocks neuropathology and motor deficits in Machado–Joseph disease mouse models through SIRT1 pathway
Caloric restriction blocks neuropathology and motor deficits in Machado–Joseph disease mouse models through SIRT1 pathway
SIRTs have been reported to provide neuroprotective actions in polyglutamine diseases, and are linked to the beneficial effects of caloric restrictive diets. Here, the authors show caloric restriction improves behavioural and neuropathological deficits in MJD model mice, an effect dependent on SIRT1 activity.
- Janete Cunha-Santos
- , Joana Duarte-Neves
- & Cláudia Cavadas
-
Article
| Open Access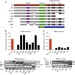 DNMT3B isoforms without catalytic activity stimulate gene body methylation as accessory proteins in somatic cells
DNMT3B isoforms without catalytic activity stimulate gene body methylation as accessory proteins in somatic cells
De novoDNA methylation is carried out by DNA methyltransferase DNMT3A/B, although DNMT3B isoforms without active catalytic domains are widely expressed. Here, the authors show that DNMT3B isoforms stimulate gene body methylation and re-methylation independently of the isoforms' catalytic activity.
- Christopher E. Duymich
- , Jessica Charlet
- & Gangning Liang
-
Article
| Open Access Epigenetic profiles signify cell fate plasticity in unipotent spermatogonial stem and progenitor cells
Epigenetic profiles signify cell fate plasticity in unipotent spermatogonial stem and progenitor cells
Spermatogonial stem cells (SSCs) spontaneously convert to multipotent adult spermatogonial-derived stem cells (MASCs). Here, the authors reveal the dynamics of bivalent histone H3-lysine4 and -lysine27 methylation signatures at somatic gene promoters in SSCs and ESC-like promoter chromatin states in MASCs.
- Ying Liu
- , Eugenia G. Giannopoulou
- & Marco Seandel
-
Article
| Open Access Genetic and environmental influences interact with age and sex in shaping the human methylome
Genetic and environmental influences interact with age and sex in shaping the human methylome
Differential impact of genetic and environmental influences on DNA methylation may result in sex- and age-related physiological variation and disease susceptibility. By analysing DNA methylome of 2,603 individuals from twin families, here, the authors establish a catalogue of between-individual variation in DNA methylation.
- Jenny van Dongen
- , Michel G. Nivard
- & Dorret I. Boomsma
-
Article
| Open Access Hemi-methylated DNA opens a closed conformation of UHRF1 to facilitate its histone recognition
Hemi-methylated DNA opens a closed conformation of UHRF1 to facilitate its histone recognition
UHRF1 is involved in the maintenance of DNA methylation, but the regulatory mechanism of this epigenetic regulator is unclear. Here, the authors show that it has a closed conformation and are able to make conclusions about the mechanism of recognition of epigenetic marks.
- Jian Fang
- , Jingdong Cheng
- & Yanhui Xu
-
Article
| Open Access Blood-based biomarkers of age-associated epigenetic changes in human islets associate with insulin secretion and diabetes
Blood-based biomarkers of age-associated epigenetic changes in human islets associate with insulin secretion and diabetes
Aging is associated with impaired pancreatic islet function, increased risk of type 2 diabetes, and changes in DNA methylation. Here the authors find blood-based biomarkers that reflect age-associated DNA methylation changes in human pancreatic islets associated with insulin secretion and diabetes.
- Karl Bacos
- , Linn Gillberg
- & Charlotte Ling
-
Article
| Open Access Sequence features accurately predict genome-wide MeCP2 binding in vivo
Sequence features accurately predict genome-wide MeCP2 binding in vivo
MeCP2 is critical for proper brain development, and mutations in the gene encoding MeCP2 are responsible for several neurological disorders. Here, the authors show that the previously reported genome-wide preference of MeCP2 to methylated CpGs is in part due to MeCP2's affinity to GC-rich chromatin.
- H. Tomas Rube
- , Wooje Lee
- & Qizhi Gong
-
Article
| Open Access Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation
Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation
Theoretical studies suggest that homologous DNA duplexes can preferentially associate with one another in the absence of proteins. Here, the authors show that GC-rich DNA with methylated cytosine and AT-rich DNA duplexes associate more strongly than GC-rich duplexes regardless of the sequence homology.
- Jejoong Yoo
- , Hajin Kim
- & Taekjip Ha
-
Article
| Open Access Genome-wide DNA methylation levels and altered cortisol stress reactivity following childhood trauma in humans
Genome-wide DNA methylation levels and altered cortisol stress reactivity following childhood trauma in humans
Exposure to childhood trauma is a major risk factor for the development of almost all psychiatric disorders. By epigenome-wide studies, here, Houtepen et al. show that DNA methylation at a locus in the Kit ligand gene (KITLG) mediates the relationship between childhood trauma and cortisol stress reactivity.
- Lotte C. Houtepen
- , Christiaan H. Vinkers
- & Marco P. M. Boks
-
Article
| Open Access Epigenetic regulation of diacylglycerol kinase alpha promotes radiation-induced fibrosis
Epigenetic regulation of diacylglycerol kinase alpha promotes radiation-induced fibrosis
Radiotherapy can induce fibrosis in cancer patients, limiting its use in clinical settings. Here, the authors identify a differentially methylated enhancer of the lipid kinase DGKA in fibroblasts from breast cancer patients developing fibrosis after radiotherapy and they show that DGKA inhibition affects lipid homeostasis and reduces pro-fibrotic fibroblast activation.
- Christoph Weigel
- , Marlon R. Veldwijk
- & Odilia Popanda
-
Article
| Open Access Biochemical reconstitution of TET1–TDG–BER-dependent active DNA demethylation reveals a highly coordinated mechanism
Biochemical reconstitution of TET1–TDG–BER-dependent active DNA demethylation reveals a highly coordinated mechanism
Cytosine methylation is a dynamic DNA modification with the involvement of the base excision repair pathway suspected to be involved in demethylation. Here the authors show that TET1 and TDG interact to target modified bases and coordinate BER to avoid double strand breaks.
- Alain R. Weber
- , Claudia Krawczyk
- & Primo Schär
-
Article
| Open Access Mutation allele burden remains unchanged in chronic myelomonocytic leukaemia responding to hypomethylating agents
Mutation allele burden remains unchanged in chronic myelomonocytic leukaemia responding to hypomethylating agents
Chronic myelomonocytic leukaemia is treated with agents that modify DNA methylation but whether they have direct cytotoxic effects is unclear. Here, the authors show that cells from treated patients show marked methylation changes without altered somatic mutation burden, suggesting that cytotoxicity is not a major factor in therapeutic efficacy.
- Jane Merlevede
- , Nathalie Droin
- & Eric Solary
-
Article
| Open Access Maternal plasma folate impacts differential DNA methylation in an epigenome-wide meta-analysis of newborns
Maternal plasma folate impacts differential DNA methylation in an epigenome-wide meta-analysis of newborns
Folic acid is routinely recommended for women trying to conceive to ensure proper fetal development. Here, the authors perform a large epigenomics study to examine which fetal epigenetic changes are associated with varied maternal plasma folate levels.
- Bonnie R. Joubert
- , Herman T. den Dekker
- & Stephanie J. London
-
Article
| Open Access Epigenetic re-expression of HIF-2α suppresses soft tissue sarcoma growth
Epigenetic re-expression of HIF-2α suppresses soft tissue sarcoma growth
Hypoxia is a common feature of soft tissue sarcomas, resulting in the activation of HIF-1α, which promotes metastasis. Here, Nakazawa et al. show that paradoxically HIF-2α is epigenetically silenced during the progression of multiple sarcoma subtypes, and when reexpressed, blocks tumour growth in vivo.
- Michael S. Nakazawa
- , T. S. Karin Eisinger-Mathason
- & M. Celeste Simon
-
Article
| Open Access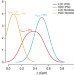 Non-CG DNA methylation is a biomarker for assessing endodermal differentiation capacity in pluripotent stem cells
Non-CG DNA methylation is a biomarker for assessing endodermal differentiation capacity in pluripotent stem cells
The methylation of non-CpG residues is a poorly understood marker of pluripotent cells, gradually lost as cells differentiate. Here the authors show non-CG methylation can be used as a marker of differentiation potential.
- Lee M. Butcher
- , Mitsuteru Ito
- & Stephan Beck
-
Article
| Open Access DNA methylation outliers in normal breast tissue identify field defects that are enriched in cancer
DNA methylation outliers in normal breast tissue identify field defects that are enriched in cancer
Altered epigenetics is a feature of cancer but whether these changes occur early in tumour development is unclear. Here, the authors analyse methylation events in breast cancer and adjacent normal pairs, and show that methylation changes in the normal tissue are also found in the tumour, suggesting that some of these events occur early in cancer.
- Andrew E Teschendorff
- , Yang Gao
- & Martin Widschwendter
-
Article
| Open Access The histone demethylase JMJD2A/KDM4A links ribosomal RNA transcription to nutrients and growth factors availability
The histone demethylase JMJD2A/KDM4A links ribosomal RNA transcription to nutrients and growth factors availability
Histone methylation regulates gene expression. Here, Salifou et al.show that KDM4A, a histone lysine demethylase, regulates RNA polymerase I-driven transcription in a PI3K dependent manner, therefore linking nutrients and growth factors availability to ribosomal RNA transcription.
- Kader Salifou
- , Swagat Ray
- & Marie Vandromme
-
Article
| Open Access NSD1 mutations generate a genome-wide DNA methylation signature
NSD1 mutations generate a genome-wide DNA methylation signature
Sotos syndrome is an growth syndrome characterized by advanced growth in childhood, characteristic facial appearance and intellectual disability. Here the authors identify a genome-wide DNA methylation signature that accurately diagnoses Sotos Syndrome and distinguishes it from similar conditions.
- S. Choufani
- , C. Cytrynbaum
- & R. Weksberg
-
Article
| Open Access H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase
H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase
DNA methylation is an important regulatory process and is essential for correct development and physiology. Here the authors show the long non-coding RNA H19regulates methylation by binding and inhibiting S-adenosylhomocysteine hydrolase.
- Jichun Zhou
- , Lihua Yang
- & Yingqun Huang
-
Article
| Open Access Embryonic transcription is controlled by maternally defined chromatin state
Embryonic transcription is controlled by maternally defined chromatin state
Histone modifying enzymes are required for cell differentiation and lineage commitment during embryonic development. By a comprehensive set of epigenome reference maps of Xenopusembryos, the authors show that H3K4me3 and H3K27me3 exert an extended maternal control well into post-gastrulation development.
- Saartje Hontelez
- , Ila van Kruijsbergen
- & Gert Jan C. Veenstra
-
Article
| Open Access Abasic pivot substitution harnesses target specificity of RNA interference
Abasic pivot substitution harnesses target specificity of RNA interference
RNA interference inadvertently represses off-target transcripts. Here, Lee et al.report that substituting nucleotide in position 6 of the seed region of the small interfering RNAs with abasic spacers can significantly decrease miRNA-like off-target repression while preserving on-target activity.
- Hye-Sook Lee
- , Heeyoung Seok
- & Sung Wook Chi
-
Article
| Open Access Epigenetic regulation of puberty via Zinc finger protein-mediated transcriptional repression
Epigenetic regulation of puberty via Zinc finger protein-mediated transcriptional repression
Zinc finger (ZNF) genes are implicated in timing human puberty. Here, the authors show that GATAD1, a ZNF protein, represses transcription of key puberty-activating genes by recruiting histone demethylase KDM1A to their promoters, suggesting GATAD1 epitomizes a subset of ZNFs involved in repression of primate puberty.
- Alejandro Lomniczi
- , Hollis Wright
- & Sergio R. Ojeda
-
Article
| Open Access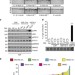 Epigenetic switch drives the conversion of fibroblasts into proinvasive cancer-associated fibroblasts
Epigenetic switch drives the conversion of fibroblasts into proinvasive cancer-associated fibroblasts
Carcinoma-associated fibroblasts are key components of solid tumours and associated with poor clinical outcome. Here the authors show that the cytokine LIF initiates an epigenetic switch which results in the sustained invasive activity of the tumour cells.
- Jean Albrengues
- , Thomas Bertero
- & Cedric Gaggioli
-
Article
| Open Access Hypomethylation of smoking-related genes is associated with future lung cancer in four prospective cohorts
Hypomethylation of smoking-related genes is associated with future lung cancer in four prospective cohorts
Smoking tobacco is known to alter DNA methylation. Here, the authors show that hypomethylation of smoke-related genes is associated with future increase in lung cancer risk.
- Francesca Fasanelli
- , Laura Baglietto
- & Paolo Vineis
-
Article
| Open Access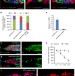 Histone H1-mediated epigenetic regulation controls germline stem cell self-renewal by modulating H4K16 acetylation
Histone H1-mediated epigenetic regulation controls germline stem cell self-renewal by modulating H4K16 acetylation
Epigenetics plays critical roles in controlling stem cell self-renewal and differentiation. Here, Sun et al. show that H1 is intrinsically required in the regulation of germline stem cells in the Drosophilaovary by antagonizing MOF, a histone acetyltransferase specific for H4K16.
- Jin Sun
- , Hui-Min Wei
- & Jian-Quan Ni
-
Article |
 Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3
Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3
Epigenetic modulation of hosts by pathogenic bacteria is underexplored. Here, Yaseen et al. show that protein Rv1988 from Mycobacterium tuberculosisenhances microbial survival by methylating histone H3 in the host cell nucleus and thus altering host gene expression.
- Imtiyaz Yaseen
- , Prabhjot Kaur
- & Sanjeev Khosla
-
Article
| Open Access Ikaros mediates gene silencing in T cells through Polycomb repressive complex 2
Ikaros mediates gene silencing in T cells through Polycomb repressive complex 2
Haematopoietic stem and progenitor cell-specific genes are epigenetically silenced during T cell differentiation. Here the authors show that Ikaros represses over 500 loci in developing T cells in cooperation with PRC2 and independently of its well established partner NuRD.
- Attila Oravecz
- , Apostol Apostolov
- & Philippe Kastner
-
Article
| Open Access Microscale insights into pneumococcal antibiotic mutant selection windows
Microscale insights into pneumococcal antibiotic mutant selection windows
The emergence of antibiotic resistance in bacteria is driven by inhibitory but non-lethal antibiotic concentrations. Here, Sorg and Veening study the effects of different antibiotics on the pneumococcus, with a focus on inhibition dynamics, metabolic activity and processes at the single-cell level.
- Robin A. Sorg
- & Jan-Willem Veening
-
Article
| Open Access Enhancer repertoires are reshaped independently of early priming and heterochromatin dynamics during B cell differentiation
Enhancer repertoires are reshaped independently of early priming and heterochromatin dynamics during B cell differentiation
Enhancers in differentiated haematopoietic cells are generally believed to be primed prior to lineage commitment. Here, the authors show that early priming and Polycomb group mediated silencing have minor roles in shaping the enhancer repertoire in differentiated B cells and that most active enhancers are generated de novo.
- Mohamed-Amin Choukrallah
- , Shuang Song
- & Patrick Matthias
-
Article
| Open Access Ribozyme-enhanced single-stranded Ago2-processed interfering RNA triggers efficient gene silencing with fewer off-target effects
Ribozyme-enhanced single-stranded Ago2-processed interfering RNA triggers efficient gene silencing with fewer off-target effects
Short hairpin RNAs are widely used to produce small interfering RNAs (siRNAs) for gene silencing. Here, the authors show that an alternative siRNA precursor in the presence of a self-cleaving ribozyme has enhanced silencing activity and reduced off-target effects, providing a potential RNAi tool.
- Renfu Shang
- , Fengjuan Zhang
- & Ligang Wu
-
Article
| Open Access Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Das et al. present a novel Bayesian approach called expression Quantitative Trait enhancer Loci (eQTeL), which effectively integrates genetic and epigenetic information to identify combination of regulatory genomic variants underlying expression variance. Using various functional data, the authors show the variants identified by eQTeL are likely to be causal.
- Avinash Das
- , Michael Morley
- & Sridhar Hannenhalli
-
Article
| Open Access An LSC epigenetic signature is largely mutation independent and implicates the HOXA cluster in AML pathogenesis
An LSC epigenetic signature is largely mutation independent and implicates the HOXA cluster in AML pathogenesis
Leukaemic stem cells are prevalent in acute myeloid leukemia. Here, Jung and colleagues derive a signature of 71 methylated genes that characterise these stem cells and find multiple HOXAgenes within the signature.
- Namyoung Jung
- , Bo Dai
- & Andrew P. Feinberg
-
Article
| Open Access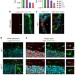 Differential genomic imprinting regulates paracrine and autocrine roles of IGF2 in mouse adult neurogenesis
Differential genomic imprinting regulates paracrine and autocrine roles of IGF2 in mouse adult neurogenesis
Selective biallelic expression of certain genes through genomic imprinting are known to play a role in controlling neurogenesis in the adult mammalian brain. Here the authors investigate the role of imprinting in the dosage control of Igf2 and its relevance for the function of IGF2 as a neurogenic regulator in the mouse brain.
- S. R. Ferrón
- , E. J. Radford
- & A. C. Ferguson-Smith
-
Article
| Open Access Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
In contrast to genetic information, epigenetic state varies greatly under different conditions. Here, the authors develop ChromDiff and apply it to ENCODE and Roadmap Epigenomics datasets to find chromatin state differences associated with sex, tissue, and developmental age.
- Angela Yen
- & Manolis Kellis
-
Article
| Open Access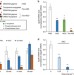 DNA/RNA heteroduplex oligonucleotide for highly efficient gene silencing
DNA/RNA heteroduplex oligonucleotide for highly efficient gene silencing
Antisense oligonucleotides (ASOs) can repress the expression of specific genes. Here, the authors show that a DNA/RNA heteroduplex oligonucleotide (HDO) with a structure different from ASOs is more potent in suppressing target gene expression, and causes a less adverse effect in mouse liver.
- Kazutaka Nishina
- , Wenying Piao
- & Takanori Yokota
-
Article
| Open Access An epigenetic regulator emerges as microtubule minus-end binding and stabilizing factor in mitosis
An epigenetic regulator emerges as microtubule minus-end binding and stabilizing factor in mitosis
The heptameric KAT8-associated nonspecific lethal complex consists of highly conserved chromatin modifier proteins. Here, the authors show a role for the members of the complex in regulating microtubule assembly during mitosis.
- Sylvain Meunier
- , Maria Shvedunova
- & Asifa Akhtar
-
Article
| Open Access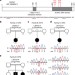 Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome
Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome
Immunodeficiency-centromeric instability-facial anomalies syndrome is a life threatening autosomal recessive disorder caused by mutations in DNMT3B and ZBTB24. Here Thijssen et al. identify mutations in CDCA7 and HELLSin previously unexplained cases.
- Peter E. Thijssen
- , Yuya Ito
- & Hiroyuki Sasaki
-
Article
| Open Access A biphasic epigenetic switch controls immunoevasion, virulence and niche adaptation in non-typeable Haemophilus influenzae
A biphasic epigenetic switch controls immunoevasion, virulence and niche adaptation in non-typeable Haemophilus influenzae
Non-typeable Haemophilus influenzae, which causes ear and lung infections, has a DNA methyltransferase encoded by alternative alleles that are subject to random ON/OFF switching. Here, Atack et al.show that this epigenetic switch controls the expression of key proteins involved in virulence.
- John M. Atack
- , Yogitha N. Srikhanta
- & Michael P. Jennings
-
Article
| Open Access Evolution of dosage compensation under sexual selection differs between X and Z chromosomes
Evolution of dosage compensation under sexual selection differs between X and Z chromosomes
Complete sex chromosome dosage compensation is largely limited to male heterogametic species, with the majority of female heterogametic species displaying incomplete dosage compensation. Here, the authors show that sexual conflict over gene expression combined with sexual selection in males can explain this pattern.
- Charles Mullon
- , Alison E. Wright
- & Judith E. Mank

 Early programming of the oocyte epigenome temporally controls late prophase I transcription and chromatin remodelling
Early programming of the oocyte epigenome temporally controls late prophase I transcription and chromatin remodelling