Article
|
Open Access
Featured
-
-
Article
| Open Access Enhancer-promoter hubs organize transcriptional networks promoting oncogenesis and drug resistance
Enhancer-promoter hubs organize transcriptional networks promoting oncogenesis and drug resistance
The role of enhancer-promoter hubs in the regulation of gene expression in cancer remains to be explored. Here, the authors identify enhancer-promoter hubs in breast cancer, lymphoma, and leukemia and suggest their potential role in promoting oncogenesis and drug resistance.
- Brent S. Perlman
- , Noah Burget
- & Robert B. Faryabi
-
Article
| Open Access Multiscale mapping of transcriptomic signatures for cardiotoxic drugs
Multiscale mapping of transcriptomic signatures for cardiotoxic drugs
Using a new computational pipeline for identification of drug-selective transcriptomic responses and FAERS data, the authors identified potential pathways and genomic variants indicative of cancer drug cardiotoxicity in iPSC-derived cardiomyocytes.
- Jens Hansen
- , Yuguang Xiong
- & Ravi Iyengar
-
Article
| Open Access Systematic identification of post-transcriptional regulatory modules
Systematic identification of post-transcriptional regulatory modules
RNA binding proteins (RBPs) regulate various RNA processes, yet their interactions remain poorly understood. Here, authors generate a comprehensive map of RBP interactions using multimodal data, uncovering context-specific functions and revealing complex post-transcriptional regulatory networks.
- Matvei Khoroshkin
- , Andrey Buyan
- & Hani Goodarzi
-
Article
| Open Access scConfluence: single-cell diagonal integration with regularized Inverse Optimal Transport on weakly connected features
scConfluence: single-cell diagonal integration with regularized Inverse Optimal Transport on weakly connected features
The abundance of unpaired multimodal single-cell data drives the need for improved integration methods. Here, authors introduce scConfluence, a method combining uncoupled autoencoders with regularized Inverse Optimal Transport to tackle robustly diverse integration scenarios.
- Jules Samaran
- , Gabriel Peyré
- & Laura Cantini
-
Article
| Open Access Herbarium collections remain essential in the age of community science
Herbarium collections remain essential in the age of community science
Here, the authors compare the diversity of vascular plants found in community science observations and digitized herbarium specimens, finding that with only one-third the records, herbaria still capture more data by several metrics.
- Isaac Eckert
- , Anne Bruneau
- & Laura J. Pollock
-
Article
| Open Access Leveraging multiple data types for improved compound-kinase bioactivity prediction
Leveraging multiple data types for improved compound-kinase bioactivity prediction
This study presents a method that enhances compound bioactivity modeling by integrating diverse data types, enabling cost-effective training data generation. In experimental testing, the best model achieves a 40% hit rate and a negative predictive value of 78%.
- Ryan Theisen
- , Tianduanyi Wang
- & Anna Cichońska
-
Article
| Open Access STIE: Single-cell level deconvolution, convolution, and clustering in in situ capturing-based spatial transcriptomics
STIE: Single-cell level deconvolution, convolution, and clustering in in situ capturing-based spatial transcriptomics
In situ capturing-based spatial transcriptomics cannot precisely capture randomly located single cells, regardless of its spot resolution. Here, authors integrate spot-level gene expression with histology images, computationally achieving single-cell level spatial transcriptomics.
- Shijia Zhu
- , Naoto Kubota
- & Yujin Hoshida
-
Article
| Open Access XENTURION is a population-level multidimensional resource of xenografts and tumoroids from metastatic colorectal cancer patients
XENTURION is a population-level multidimensional resource of xenografts and tumoroids from metastatic colorectal cancer patients
Improvement of preclinical models is critical for ensuring effective treatment discovery for colorectal cancer. Here, the authors develop a platform of 128 PDX models from metastatic colorectal cancer with matched tumouroid cultures, and use these to demonstrate molecular concordance between PDX-tumouroid pairs, cetuximab sensitivity heterogeneity, and adaptive upregulation of druggable targets under cetuximab pressure.
- Simonetta M. Leto
- , Elena Grassi
- & Livio Trusolino
-
Article
| Open Access Secondary integrated analysis of multi-tissue transcriptomic responses to a combined lifestyle intervention in older adults from the GOTO nonrandomized trial
Secondary integrated analysis of multi-tissue transcriptomic responses to a combined lifestyle intervention in older adults from the GOTO nonrandomized trial
Molecular effects of lifestyle interventions are typically studied within one tissue, neglecting potential shared responses across tissues. Here, the authors show that subcutaneous adipose tissue RNA levels best capture health benefits of the intervention and identified joint effects among blood, subcutaneous adipose tissue and muscle tissue RNA levels.
- F. A. Bogaards
- , T. Gehrmann
- & P. E. Slagboom
-
Article
| Open Access HMGA1 orchestrates chromatin compartmentalization and sequesters genes into 3D networks coordinating senescence heterogeneity
HMGA1 orchestrates chromatin compartmentalization and sequesters genes into 3D networks coordinating senescence heterogeneity
HMGA1 helps regulate the topology of the chromatin network, supporting compartmentalization. Here, Olan et al. show that, in oncogene-induced senescence, genes are included or excluded from HMGA1 cores, suggesting HMGA1 has a fine-tuning role.
- Ioana Olan
- , Masami Ando-Kuri
- & Masashi Narita
-
Article
| Open Access Prediction of plant complex traits via integration of multi-omics data
Prediction of plant complex traits via integration of multi-omics data
Translating genotype to phenotype is a grand challenge in biology. Here, the authors investigate the utility of genome, transcriptome, and methylome data and their combinations in predicting six plant complex traits and uncovering key genes and genetic interactions in Arabidopsis.
- Peipei Wang
- , Melissa D. Lehti-Shiu
- & Shin-Han Shiu
-
Article
| Open Access A unified model for interpretable latent embedding of multi-sample, multi-condition single-cell data
A unified model for interpretable latent embedding of multi-sample, multi-condition single-cell data
Single-cell analysis of multi-condition cohorts requires modelling the interaction between sample variables and cell states. Here, authors develop GEDI to enable integration, cluster-free differential expression analysis and regulon analysis for both gene expression and alternative splicing modalities.
- Ariel Madrigal
- , Tianyuan Lu
- & Hamed S. Najafabadi
-
Article
| Open Access Evaluating batch correction methods for image-based cell profiling
Evaluating batch correction methods for image-based cell profiling
Batch effects can limit the usefulness of image-based profiling data. Here, authors benchmark ten popular batch correction techniques on a large Cell Painting dataset, evaluating multiple metrics. They identify Harmony and Seurat RPCA as top methods across diverse complex scenarios.
- John Arevalo
- , Ellen Su
- & Shantanu Singh
-
Article
| Open Access Whole brain alignment of spatial transcriptomics between humans and mice with BrainAlign
Whole brain alignment of spatial transcriptomics between humans and mice with BrainAlign
Comparative transcriptomics of whole brains across species is vital in neuroscience. Here, authors develop a deep learning method, BrainAlign, to align spatial transcriptomics across human and mouse brains. BrainAlign identifies conserved brain regions and uncovers similar patterns for marker genes.
- Biao Zhang
- , Shuqin Zhang
- & Shihua Zhang
-
Article
| Open Access Multiomic profiling of medulloblastoma reveals subtype-specific targetable alterations at the proteome and N-glycan level
Multiomic profiling of medulloblastoma reveals subtype-specific targetable alterations at the proteome and N-glycan level
Medulloblastomas (MBs) are highly heterogeneous paediatric brain tumours that remain challenging to treat. Here, the authors integrate proteomics, epigenomics, transcriptomics and post-translational modification analyses to find molecular subgroups and potential therapeutic targets in MB tumours.
- Shweta Godbole
- , Hannah Voß
- & Julia E. Neumann
-
Article
| Open Access SANTO: a coarse-to-fine alignment and stitching method for spatial omics
SANTO: a coarse-to-fine alignment and stitching method for spatial omics
Spatial omics technologies require alignment and stitching of slices for a 3D molecular profile. Here, the authors present SANTO, a coarse-to-fine method that rapidly determines spatial positions and overlap regions, then refines them, enabling integration across platforms and modalities.
- Haoyang Li
- , Yingxin Lin
- & Xin Gao
-
Article
| Open Access Directional integration and pathway enrichment analysis for multi-omics data
Directional integration and pathway enrichment analysis for multi-omics data
The data fusion method presented here integrates multi-omics datasets for gene prioritisation, biomarker discovery, and pathway enrichment analysis by finding genes and proteins with significant and directionally consistent changes across the data modalities.
- Mykhaylo Slobodyanyuk
- , Alexander T. Bahcheli
- & Jüri Reimand
-
Article
| Open Access Multi-modal generative modeling for joint analysis of single-cell T cell receptor and gene expression data
Multi-modal generative modeling for joint analysis of single-cell T cell receptor and gene expression data
Although single-cell RNA sequencing analysis now allows simultaneous examination of transcriptome and T cell receptor repertoire sequences, integrating these two modalities remains a challenge. Here, the authors develop mvTCR, a generative deep learning model that integrates transcriptome and T cell receptor data into a joint representation capturing cell functions and phenotypes.
- Felix Drost
- , Yang An
- & Benjamin Schubert
-
Article
| Open Access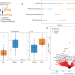 Distinct pulmonary and systemic effects of dexamethasone in severe COVID-19
Distinct pulmonary and systemic effects of dexamethasone in severe COVID-19
Dexamethasone has been used in the treatment of critically ill COVID-19 patients. Here the authors apply transcriptomics to investigate the effects of dexamethasone treatment in COVID-19 patients, and show both systemic and compartment-specific effects.
- Lucile P. A. Neyton
- , Ravi K. Patel
- & Gabriela K. Fragiadakis
-
Article
| Open Access Dissecting tumor microenvironment from spatially resolved transcriptomics data by heterogeneous graph learning
Dissecting tumor microenvironment from spatially resolved transcriptomics data by heterogeneous graph learning
Dissecting the relations between cells, genes, and histological regions in the tumor microenvironment (TME) remains challenging. Here, the authors develop stKeep, a heterogeneous graph learning method that integrates multimodal data and gene-gene interactions to identify cell states and composition in the TME from spatial transcriptomics.
- Chunman Zuo
- , Junjie Xia
- & Luonan Chen
-
Article
| Open Access DOT: a flexible multi-objective optimization framework for transferring features across single-cell and spatial omics
DOT: a flexible multi-objective optimization framework for transferring features across single-cell and spatial omics
Single-cell and spatial omics come with a trade-off between resolution and gene coverage. Here, authors bridge this gap via DOT, a multi-objective optimisation model for localising cell features in high/low-resolution spatial data considering cell composition, heterogeneity, and technical effects.
- Arezou Rahimi
- , Luis A. Vale-Silva
- & Julio Saez-Rodriguez
-
Article
| Open Access A variational expectation-maximization framework for balanced multi-scale learning of protein and drug interactions
A variational expectation-maximization framework for balanced multi-scale learning of protein and drug interactions
Multi-scale learning still struggles with imbalanced information and greedy characteristics. Here the authors present MUSE, an Expectation-Maximization-based multi-scale framework, improving predictions across molecular interactions and atomic interfaces.
- Jiahua Rao
- , Jiancong Xie
- & Yuedong Yang
-
Article
| Open Access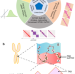 A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains
A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains
TAD hierarchy demonstrates cell-to-cell variability, leading to the development of numerous callers. Here, authors present a comprehensive benchmark of TAD hierarchy callers and introduce the ‘air conditioner’ model to illustrate TAD hierarchy’s role in transcription.
- Jingxuan Xu
- , Xiang Xu
- & Hebing Chen
-
Article
| Open Access 1q amplification and PHF19 expressing high-risk cells are associated with relapsed/refractory multiple myeloma
1q amplification and PHF19 expressing high-risk cells are associated with relapsed/refractory multiple myeloma
Translocations and copy number variations that affect multiple myeloma (MM) have not been investigated at the single cell level. Here, single cell multi-omics reveal the relationship between epigenetic regulation and cytogenetic events that lead to the increase of cell proliferation in MM.
- Travis S. Johnson
- , Parvathi Sudha
- & Brian A. Walker
-
Article
| Open Access The impact of exercise on gene regulation in association with complex trait genetics
The impact of exercise on gene regulation in association with complex trait genetics
It is known that exercise influences many human traits, but not which tissues and genes are most important. This study connects transcriptome data collected across 15 tissues during exercise training in rats as part of the Molecular Transducers of Physical Activity Consortium with human data to identify traits with similar tissue specific gene expression signatures to exercise.
- Nikolai G. Vetr
- , Nicole R. Gay
- & Stephen B. Montgomery
-
Article
| Open Access Integrated proteogenomic and metabolomic characterization of papillary thyroid cancer with different recurrence risks
Integrated proteogenomic and metabolomic characterization of papillary thyroid cancer with different recurrence risks
Papillary thyroid cancers (PTC) generally have good prognosis, but their recurrence rate remains high. Here, the authors use proteogenomics and metabolomics to identify molecular features in PTC tumours and determine PTC subtypes that are associated with prognosis and potential targeted therapies.
- Ning Qu
- , Di Chen
- & Rongliang Shi
-
Article
| Open Access Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Here, the authors employ multi-omics on a cohort comprising three generations of family members, showing that fecal microbiota populations, functions, and metabolome of infants vary greatly from their maternal lineage, exhibiting a less diverse microbiota and differences in various metabolite classes including short- and branched-chain fatty acids.
- Tomás Clive Barker-Tejeda
- , Elisa Zubeldia-Varela
- & Alma Villaseñor
-
Article
| Open Access scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
Technical limitations of simultaneously multi-omics profiling lead to highly noisy multi-modal data and substantial costs. Here, authors proposed a versatile framework and data augmentation schemes, capable of single-cell cross-modality translation and multiple extensive applications.
- Yichuan Cao
- , Xiamiao Zhao
- & Shengquan Chen
-
Article
| Open Access Multi-omic integration of microbiome data for identifying disease-associated modules
Multi-omic integration of microbiome data for identifying disease-associated modules
Here, Muller et al. introduce MintTea, a method for analyzing multi-omic microbiome data and identifying disease-associated modules comprising mixed sets of features that collectively shift in disease, offering insights into microbiome-disease interactions.
- Efrat Muller
- , Itamar Shiryan
- & Elhanan Borenstein
-
Article
| Open Access Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)
Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)
Use of cloud-based personal health records are increasing globally. Here, authors introduce the Global Patient co-Owned Cloud (GPOC) concept. The systematic review and meta-analysis examine factors like data security, efficiency, privacy, and cost. It aims to establish a scientific basis for a GPOC, which may disseminate global artificial intelligence for healthcare.
- Niklas Lidströmer
- , Joe Davids
- & Eric Herlenius
-
Article
| Open Access Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
By developing computational algorithms, the authors annotated translated open reading frames in five eukaryotes and found many stable peptides are encoded by putative ‘noncoding’ regions of genomes.
- Haiwang Yang
- , Qianru Li
- & Zhe Ji
-
Article
| Open Access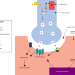 Rare disease research workflow using multilayer networks elucidates the molecular determinants of severity in Congenital Myasthenic Syndromes
Rare disease research workflow using multilayer networks elucidates the molecular determinants of severity in Congenital Myasthenic Syndromes
Congenital myasthenic syndromes are rare inherited neuromuscular disorders. Here, the authors attempt to explain diverse disease severity seen in 20 patients with shared CHRNE gene mutations with a multilayer network analysis that identifies individual-level impairments at the neuromuscular junction.
- Iker Núñez-Carpintero
- , Maria Rigau
- & Alfonso Valencia
-
Article
| Open Access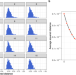 Imputation of plasma lipid species to facilitate integration of lipidomic datasets
Imputation of plasma lipid species to facilitate integration of lipidomic datasets
Advancements in plasma lipidomic profiling increase specificity of measurements but pose challenges in aligning datasets created at different times or platforms. Here the authors present a predictive framework for harmonising such datasets with different levels of granularity in their lipid measurements.
- Aleksandar Dakic
- , Jingqin Wu
- & Peter J. Meikle
-
Article
| Open Access scDisInFact: disentangled learning for integration and prediction of multi-batch multi-condition single-cell RNA-sequencing data
scDisInFact: disentangled learning for integration and prediction of multi-batch multi-condition single-cell RNA-sequencing data
Here the authors propose a deep learning model that integrates multi-condition, multi-batch single-cell RNA-sequencing datasets. The model disentangles biological variation (condition effect) from technical confounders (batch effect) and overcomes some limitations of existing approaches.
- Ziqi Zhang
- , Xinye Zhao
- & Xiuwei Zhang
-
Article
| Open Access Human whole-exome genotype data for Alzheimer’s disease
Human whole-exome genotype data for Alzheimer’s disease
The heterogeneity of whole-exome sequencing (WES) data generation methods presents a challenge to joint analysis. Here, the authors present a bioinformatics strategy to generate high-quality data from processing diversely generated WES samples, as applied in the Alzheimer’s Disease Sequencing Project.
- Yuk Yee Leung
- , Adam C. Naj
- & Li-San Wang
-
Article
| Open Access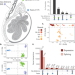 Molecular quantitative trait loci in reproductive tissues impact male fertility in cattle
Molecular quantitative trait loci in reproductive tissues impact male fertility in cattle
Investigating the genetics of male fertility requires comprehensive genotype and phenotype data. Here, the authors characterize the transcriptional complexity of bovine male reproductive tissues to identify loci associated with male fertility.
- Xena Marie Mapel
- , Naveen Kumar Kadri
- & Hubert Pausch
-
Article
| Open Access Longitudinal single cell atlas identifies complex temporal relationship between type I interferon response and COVID-19 severity
Longitudinal single cell atlas identifies complex temporal relationship between type I interferon response and COVID-19 severity
Single cell transcriptomics can reveal at high resolution the body’s response to infection. Here the authors have applied this technology to a longitudinal SARS-CoV-2 infected cohort and identified gene expression changes that may predict disease severity and reveal the underlying molecular mechanisms.
- Quy Xiao Xuan Lin
- , Deepa Rajagopalan
- & Shyam Prabhakar
-
Article
| Open Access PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Understanding biological mechanisms requires a thorough exploration of spatiotemporal transcriptional patterns in complex tissues. Here, authors present PROST to quantify spatial gene expression patterns and detect spatial domains using spatial transcriptomics data of varying resolutions.
- Yuchen Liang
- , Guowei Shi
- & Zhonghui Tang
-
Article
| Open Access BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
Subcellular in situ spatial transcriptomics offers the promise to address biological problems that were previously inaccessible but requires accurate cell segmentation to uncover insights. Here, authors present BIDCell, a biologically informed, deep learning-based cell segmentation framework.
- Xiaohang Fu
- , Yingxin Lin
- & Jean Y. H. Yang
-
Article
| Open Access MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data
Identifying tissue structure in large-scale spatial omics datasets from multiple slices is challenging. Here, authors present MENDER, an optimisation-free spatial clustering method that can scale to million-level spatial data, enabling efficient analysis of spatial cell atlases.
- Zhiyuan Yuan
-
Article
| Open Access Integrative genotyping of cancer and immune phenotypes by long-read sequencing
Integrative genotyping of cancer and immune phenotypes by long-read sequencing
Single-cell transcriptomics excel in cell subset classification and can be augmented by suitable genotype information. Here the authors devise a long-read sequencing workflow, termed nanoranger, for detection of molecular barcodes from single-cell cDNA and apply this to clonal tracking of acute myeloid leukemia and identification of complex immune phenotypes.
- Livius Penter
- , Mehdi Borji
- & Catherine J. Wu
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access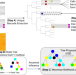 LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
Inferring lineage trees while incorporating gene expressions and lineage barcodes is a challenging task. Here, authors present LinRace, which infers improved cell lineage trees and ancestral cell states using the proposed asymmetric division model.
- Xinhai Pan
- , Hechen Li
- & Xiuwei Zhang
-
Article
| Open Access Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Co-fractionation mass spectrometry (CF-MS) is a powerful technique for mapping protein interactions under physiological conditions. Here, the authors uniformly re-process 411 CF-MS experiments and carry out meta-analyses of protein abundance, protein-protein interactions, and phosphorylation sites in the resulting resource.
- Michael A. Skinnider
- , Mopelola O. Akinlaja
- & Leonard J. Foster
-
Article
| Open Access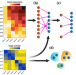 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
Integration of single-cell datasets is essential to gain a comprehensive understanding of complex biological systems. Here, the authors develop scDREAMER, a deep generative framework for performing unsupervised and supervised atlas-level integration, demonstrating improved bio-conservation and batch-correction.
- Ajita Shree
- , Musale Krushna Pavan
- & Hamim Zafar
-
Article
| Open Access Paired single-cell multi-omics data integration with Mowgli
Paired single-cell multi-omics data integration with Mowgli
Mowgli is a novel paired single-cell multi-omics integration method leveraging matrix factorization and Optimal Transport. In-depth benchmarking demonstrates promising cell clustering results and improved biological interpretability.
- Geert-Jan Huizing
- , Ina Maria Deutschmann
- & Laura Cantini
-
Article
| Open Access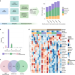 Distinct transcriptomic profiles in children prior to the appearance of type 1 diabetes-linked islet autoantibodies and following enterovirus infection
Distinct transcriptomic profiles in children prior to the appearance of type 1 diabetes-linked islet autoantibodies and following enterovirus infection
Although type-1 diabetes has a clear genetic component, not all children who are at risk eventually develop autoimmunity, suggesting the existence of environmental triggers. In this longitudinal transcriptomic study, the authors find that children who later develop autoimmunity have a distinct profile before the appearance of autoantibodies and may have impaired responses to enterovirus infection.
- Jake Lin
- , Elaheh Moradi
- & Matti Nykter

 Identifying genetic variants associated with chromatin looping and genome function
Identifying genetic variants associated with chromatin looping and genome function