Featured
-
-
Letter |
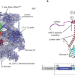 Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM
Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM
A cryo-electron microscopy structure of the mammalian mitochondrial translation initiation complex reveals unique features that are necessary to initiate mitochondrial translation and highlights a possible membrane-targeting peptide.
- Eva Kummer
- , Marc Leibundgut
- & Nenad Ban
-
Letter |
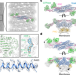 Cryo-EM of the dynamin polymer assembled on lipid membrane
Cryo-EM of the dynamin polymer assembled on lipid membrane
A cryo-electron microscopy structure of human dynamin-1 demonstrates conformational changes and sheds light on the fission of membranes during endocytosis.
- Leopold Kong
- , Kem A. Sochacki
- & Jenny E. Hinshaw
-
Article |
 Cryo-EM structure of a fungal mitochondrial calcium uniporter
Cryo-EM structure of a fungal mitochondrial calcium uniporter
A cryo-electron microscopy structure of fungal mitochondrial calcium uniporter shows that the channel is tetrameric and sheds light on channel assembly and function.
- Nam X. Nguyen
- , Jean-Paul Armache
- & Youxing Jiang
-
Article |
 Cryo-EM structures of fungal and metazoan mitochondrial calcium uniporters
Cryo-EM structures of fungal and metazoan mitochondrial calcium uniporters
Structures of the mitochondrial calcium uniporter from fungal and metazoan organisms reveal a tetrameric architecture and shed light on the function of the channel.
- Rozbeh Baradaran
- , Chongyuan Wang
- & Stephen Barstow Long
-
Letter |
 Structures of human Patched and its complex with native palmitoylated sonic hedgehog
Structures of human Patched and its complex with native palmitoylated sonic hedgehog
High-resolution structures of the human plasma membrane protein patched 1 alone and in complex with the native form of the ligand sonic hedgehog are determined.
- Xiaofeng Qi
- , Philip Schmiege
- & Xiaochun Li
-
Letter |
 Prespliceosome structure provides insights into spliceosome assembly and regulation
Prespliceosome structure provides insights into spliceosome assembly and regulation
The cryo-electron microscopy structure of the Saccharomyces cerevisiae prespliceosome provides insights into splice-site selection and early spliceosome assembly events.
- Clemens Plaschka
- , Pei-Chun Lin
- & Kiyoshi Nagai
-
Article |
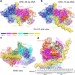 Structure of the origin recognition complex bound to DNA replication origin
Structure of the origin recognition complex bound to DNA replication origin
The cryo-EM structure of the yeast origin recognition complex (ORC) bound to a 72-base-pair origin DNA sequence provides insights into the basis of the origin selection mechanism.
- Ningning Li
- , Wai Hei Lam
- & Bik-Kwoon Tye
-
Letter |
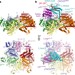 Mechanism for remodelling of the cell cycle checkpoint protein MAD2 by the ATPase TRIP13
Mechanism for remodelling of the cell cycle checkpoint protein MAD2 by the ATPase TRIP13
Structural analysis demonstrates how TRIP13 and p31comet disassemble the mitotic checkpoint complex.
- Claudio Alfieri
- , Leifu Chang
- & David Barford
-
Letter |
 Cryo-EM structure of an essential Plasmodium vivax invasion complex
Cryo-EM structure of an essential Plasmodium vivax invasion complex
Structural studies show that conserved residues in Plasmodium vivax reticulocyte-binding protein 2b determine interactions with transferrin receptor 1 that are essential for host invasion, suggesting avenues for designing vaccines that work across P. vivax strains.
- Jakub Gruszczyk
- , Rick K. Huang
- & Wai-Hong Tham
-
Article |
 Structure of a human synaptic GABAA receptor
Structure of a human synaptic GABAA receptor
The cryo-electron microscopy structure of the type A GABA receptor bound to GABA and the benzodiazepine site antagonist flumazenil reveals structural mechanisms that underlie intersubunit interactions and ligand selectivity of the receptor.
- Shaotong Zhu
- , Colleen M. Noviello
- & Ryan E. Hibbs
-
Letter |
 Cryo-EM structure of the serotonin 5-HT1B receptor coupled to heterotrimeric Go
Cryo-EM structure of the serotonin 5-HT1B receptor coupled to heterotrimeric Go
The high-resolution structure of the serotonin 5-HT1B receptor in complex with the agonist donitriptan and a Go heterotrimer highlights features that may underlie the specificity of receptor–G-protein coupling and kinetics of signalling.
- Javier García-Nafría
- , Rony Nehmé
- & Christopher G. Tate
-
Article |
 Structure of the adenosine-bound human adenosine A1 receptor–Gi complex
Structure of the adenosine-bound human adenosine A1 receptor–Gi complex
The cryo-electron microscopy structure of the human adenosine A1 receptor in complex with adenosine and heterotrimeric Gi2 protein provides molecular insights into receptor and G-protein selectivity.
- Christopher J. Draper-Joyce
- , Maryam Khoshouei
- & Arthur Christopoulos
-
Article |
 Cryo-EM structure of human rhodopsin bound to an inhibitory G protein
Cryo-EM structure of human rhodopsin bound to an inhibitory G protein
The cryo-electron microscopy structure of human rhodopsin bound to the inhibitory Gi protein-coupled receptor provides insights into ligand–receptor–G-protein interactions.
- Yanyong Kang
- , Oleg Kuybeda
- & H. Eric Xu
-
Article |
 Structure of the µ-opioid receptor–Gi protein complex
Structure of the µ-opioid receptor–Gi protein complex
A cryo-electron structure of the µ-opioid receptor in complex with the peptide agonist DAMGO and the inhibitory G protein Gi reveals structural determinants of its G protein-binding specificity.
- Antoine Koehl
- , Hongli Hu
- & Brian K. Kobilka
-
Letter |
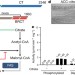 Structural basis for regulation of human acetyl-CoA carboxylase
Structural basis for regulation of human acetyl-CoA carboxylase
Cryo-electron microscopy studies of distinct, catalytically active and inactive filaments of human acetyl-CoA carboxylase 1 reveal the structural basis of its regulation.
- Moritz Hunkeler
- , Anna Hagmann
- & Timm Maier
-
Article |
 Structural basis of mitochondrial receptor binding and constriction by DRP1
Structural basis of mitochondrial receptor binding and constriction by DRP1
Cryo-electron microscopy is used to resolve the structure of human dynamin-related protein 1 co-assembled with its receptor mitochondrial dynamics protein of 49 kDa, along with an analysis of structure- and disease-based mutations.
- Raghav Kalia
- , Ray Yu-Ruei Wang
- & Adam Frost
-
Article |
 Visualizing late states of human 40S ribosomal subunit maturation
Visualizing late states of human 40S ribosomal subunit maturation
Cryo-EM structures of late intermediates in the assembly of human 40S ribosomal subunits help to define the principles by which immature rRNA conformations and ribosomal biogenesis factors shape the 40S maturation process.
- Michael Ameismeier
- , Jingdong Cheng
- & Roland Beckmann
-
Article |
 Structure of a volume-regulated anion channel of the LRRC8 family
Structure of a volume-regulated anion channel of the LRRC8 family
The structure of a homomeric channel of subunit A of leucine-rich repeat-containing protein 8 (LRRC8) determined by cryo-electron microscopy and X-ray crystallography reveals the basis for anion selectivity.
- Dawid Deneka
- , Marta Sawicka
- & Raimund Dutzler
-
Letter |
 Structural principles of distinct assemblies of the human α4β2 nicotinic receptor
Structural principles of distinct assemblies of the human α4β2 nicotinic receptor
Cryo-electron microscopy structures of two stoichiometries of heteromeric acetylcholine receptors in complex with nicotine reveal principles of subunit assembly and the structural basis of the distinctive biophysical and pharmacological properties of the different stoichiometries.
- Richard M. Walsh Jr
- , Soung-Hun Roh
- & Ryan E. Hibbs
-
Letter |
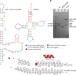 Architecture of an HIV-1 reverse transcriptase initiation complex
Architecture of an HIV-1 reverse transcriptase initiation complex
A cryo-EM structure of an initiation complex of HIV-1 reverse transcriptase sheds light on the initiation of reverse transcription of viral RNA.
- Kevin P. Larsen
- , Yamuna Kalyani Mathiharan
- & Elisabetta Viani Puglisi
-
Letter |
 Structure of the alternative complex III in a supercomplex with cytochrome oxidase
Structure of the alternative complex III in a supercomplex with cytochrome oxidase
The structure of alternative complex III, a key enzyme in the bacterial electron transport chain, is reported both alone and as a supercomplex with an aa3-type cytochrome c oxidase.
- Chang Sun
- , Samir Benlekbir
- & Robert B. Gennis
-
Article |
 Cryo-EM structure of the gasdermin A3 membrane pore
Cryo-EM structure of the gasdermin A3 membrane pore
High-resolution cryo-electron microscopy structures of the membrane-pore-forming domain of the mouse gasdermin GSDMA3 show that it forms pores with 26-, 27- or 28-fold symmetry and indicate that it may also form a parallel, soluble, pre-pore ring structure.
- Jianbin Ruan
- , Shiyu Xia
- & Hao Wu
-
Article |
 Cryo-EM structure of substrate-bound human telomerase holoenzyme
Cryo-EM structure of substrate-bound human telomerase holoenzyme
A cryo-electron microscopy structure of the substrate-bound human telomerase holoenzyme, which lengthens the protective caps on chromosomes.
- Thi Hoang Duong Nguyen
- , Jane Tam
- & Kathleen Collins
-
Letter |
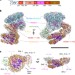 Structure and regulation of the human INO80–nucleosome complex
Structure and regulation of the human INO80–nucleosome complex
Cryo-electron microscopy structure of the human INO80 chromatin remodeller in complex with a bound nucleosome reveals that its motor domains are located at the DNA wrap around the histone core.
- Rafael Ayala
- , Oliver Willhoft
- & Xiaodong Zhang
-
Letter |
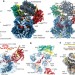 Structural basis for ATP-dependent chromatin remodelling by the INO80 complex
Structural basis for ATP-dependent chromatin remodelling by the INO80 complex
Cryo-electron microscopy structures of the evolutionarily conserved core of a fungal INO80 complex bound to the nucleosomal substrate reveal the mechanism underlying nucleosome sliding and histone editing used by this ATP-dependent chromatin remodeller.
- Sebastian Eustermann
- , Kevin Schall
- & Karl-Peter Hopfner
-
Letter |
 Molecular nucleation mechanisms and control strategies for crystal polymorph selection
Molecular nucleation mechanisms and control strategies for crystal polymorph selection
Cryo-transmission electron microscopy reveals the initiation of glucose isomerase crystals and their growth into different crystalline or gel polymorphs, and shows that mutating specific amino acids controls which polymorph grows.
- Alexander E. S. Van Driessche
- , Nani Van Gerven
- & Mike Sleutel
-
Article |
 Cryo-EM structure of the
Blastochloris viridis LH1–RC complex at 2.9 Å
Cryo-EM structure of the
Blastochloris viridis LH1–RC complex at 2.9 Å
A cryo-electron microscopy structure of the light-harvesting–reaction centre (LH1–RC) complex of the photosynthetic bacterium Blastochloris viridis suggests factors that underlie the large redshift in the absorption spectrum of bacteriochlorophyll in the complex and that promote quinone–quinol translocation across the LH1 ring.
- Pu Qian
- , C. Alistair Siebert
- & C. Neil Hunter
-
Article |
 Architecture of the human GATOR1 and GATOR1–Rag GTPases complexes
Architecture of the human GATOR1 and GATOR1–Rag GTPases complexes
Cryo-electron microscopy and biochemical analyses of the GATOR1 protein complex reveal that two binding modes underpin its ability to regulate Rag GTPases as a GTPase-activating protein for RAGA.
- Kuang Shen
- , Rick K. Huang
- & David M. Sabatini
-
Letter |
 Structural insights into the voltage and phospholipid activation of the mammalian TPC1 channel
Structural insights into the voltage and phospholipid activation of the mammalian TPC1 channel
Structures of the voltage-gated and phosphatidylinositol 3,5-bisphosphate-activated mouse two-pore channel TPC1 in apo and ligand-bound states provide insights into the selectivity and gating mechanisms of mammalian two-pore channels.
- Ji She
- , Jiangtao Guo
- & Xiao-chen Bai
-
Letter |
 Modular assembly of the nucleolar pre-60S ribosomal subunit
Modular assembly of the nucleolar pre-60S ribosomal subunit
Cryo-electron microscopy structures of multiple states along the assembly pathway of the nucleolar pre-60S ribosomal subunit suggest that folding of the pre-ribosomal RNA and sequential binding of transient assembly factors are regulated by steric hindrance among these factors.
- Zahra Assur Sanghai
- , Linamarie Miller
- & Sebastian Klinge
-
Letter |
 Structure of the insulin receptor–insulin complex by single-particle cryo-EM analysis
Structure of the insulin receptor–insulin complex by single-particle cryo-EM analysis
Cryo-electron microscopy structures of insulin in a complex with the insulin receptor define the S2 binding site on the receptor and suggest a mechanism for downstream propagation of insulin signalling.
- Giovanna Scapin
- , Venkata P. Dandey
- & Bridget Carragher
-
Article |
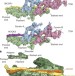 Cryo-EM shows how dynactin recruits two dyneins for faster movement
Cryo-EM shows how dynactin recruits two dyneins for faster movement
Cryo-electron microscopy and single-molecule studies reveal that the adaptors BICDR1 and HOOK3 recruit two dynein molecules to dynactin and thereby increase the force and speed of the dynein–dynactin microtubule motor.
- Linas Urnavicius
- , Clinton K. Lau
- & Andrew P. Carter
-
Article |
 Structure and mechanogating mechanism of the Piezo1 channel
Structure and mechanogating mechanism of the Piezo1 channel
The electron cryo-microscopy structure of full-length mouse Piezo1 reveals unique topological features such as the repetitive transmembrane helical units that constitute the highly curved transmembrane region, and identifies regions and single residues that are crucial for the mechanical activation of the channel.
- Qiancheng Zhao
- , Heng Zhou
- & Bailong Xiao
-
Article |
 The atomic structure of a eukaryotic oligosaccharyltransferase complex
The atomic structure of a eukaryotic oligosaccharyltransferase complex
The high-resolution structure of the entire eukaryotic oligosaccharyltransferase complex is determined, revealing the role of membrane lipids in mediating inter-subunit interactions, and the mechanism by which the complex enables protein N-glycosylation.
- Lin Bai
- , Tong Wang
- & Huilin Li
-
Article |
 Molecular mechanism of promoter opening by RNA polymerase III
Molecular mechanism of promoter opening by RNA polymerase III
Cryo-EM structures of Pol III preinitiation complexes are presented, comprising Pol III and the transcription factor TFIIIB bound to a natural promoter in different functional states.
- Matthias K. Vorländer
- , Heena Khatter
- & Christoph W. Müller
-
Article |
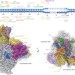 Structural basis of RNA polymerase III transcription initiation
Structural basis of RNA polymerase III transcription initiation
Detailed structures of yeast RNA polymerase III and its initiation complex shed light on how the transcription of essential non-coding RNAs begins and allow comparisons with other RNA polymerases.
- Guillermo Abascal-Palacios
- , Ewan Phillip Ramsay
- & Alessandro Vannini
-
Letter |
 Structure and mutagenesis reveal essential capsid protein interactions for KSHV replication
Structure and mutagenesis reveal essential capsid protein interactions for KSHV replication
Cryo-electron microscopy reveals the structure of the Kaposi’s sarcoma-associated herpesvirus capsid, and experiments with polypeptides that mimic the smallest capsid protein demonstrate the potential for structure-derived insights to help to develop antiviral agents.
- Xinghong Dai
- , Danyang Gong
- & Z. Hong Zhou
-
Article |
 Structure of the mechanically activated ion channel Piezo1
Structure of the mechanically activated ion channel Piezo1
The cryo-electron microscopy structure of full-length mouse Piezo1 reveals six Piezo repeats, and 26 transmembrane helices per protomer, and shows that a kinked helical beam and anchor domain link the Piezo repeats to the pore and control gating allosterically.
- Kei Saotome
- , Swetha E. Murthy
- & Andrew B. Ward
-
Letter |
 Opening of the human epithelial calcium channel TRPV6
Opening of the human epithelial calcium channel TRPV6
The cryo-electron microscopy structure of the calcium channel TRPV6 in its open and closed states demonstrates a novel gating mechanism involving an alanine hinge.
- Luke L. McGoldrick
- , Appu K. Singh
- & Alexander I. Sobolevsky
-
Letter |
 Cryo-EM structures of the TMEM16A calcium-activated chloride channel
Cryo-EM structures of the TMEM16A calcium-activated chloride channel
Electron cryo-microscopy density maps of mouse TMEM16A reconstituted in nanodiscs or solubilized in detergent reveal two functional states of calcium-activated chloride channels.
- Shangyu Dang
- , Shengjie Feng
- & Lily Yeh Jan
-
Article |
 Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40
Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40
The cryo-electron microscopy and crystal structures of several mTORC1 complexes, and accompanying biochemical analyses, shed light on how mTORC1 is regulated and how cancer mutations lead to its hyperactivation.
- Haijuan Yang
- , Xiaolu Jiang
- & Nikola P. Pavletich
-
Letter |
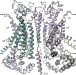 Activation mechanism of the calcium-activated chloride channel TMEM16A revealed by cryo-EM
Activation mechanism of the calcium-activated chloride channel TMEM16A revealed by cryo-EM
Cryo-electron microscopy mapping of the calcium-activated chloride channel TMEM16A combined with functional experiments reveals that calcium ions interact directly with the pore to activate the channel.
- Cristina Paulino
- , Valeria Kalienkova
- & Raimund Dutzler
-
Article |
 Electron cryo-microscopy structure of a human TRPM4 channel
Electron cryo-microscopy structure of a human TRPM4 channel
The structure of the Ca2+-activated, non-selective ion channel TRPM4 bound to the agonist Ca2+ and a modulator decavanadate, solved using electron cryo-microscopy.
- Paige A. Winkler
- , Yihe Huang
- & Wei Lü
-
Article |
 Structures of the calcium-activated, non-selective cation channel TRPM4
Structures of the calcium-activated, non-selective cation channel TRPM4
Electron cryo-microscopy structures of mouse TRPM4, a calcium-activated, non-selective cation channel, in the apo and ATP-bound states.
- Jiangtao Guo
- , Ji She
- & Youxing Jiang
-
Letter |
 Structural basis for the initiation of eukaryotic transcription-coupled DNA repair
Structural basis for the initiation of eukaryotic transcription-coupled DNA repair
Cryo-electron microscopy analysis of yeast Rad26 bound to RNA polymerase II provides insight into the initiation of the transcription-coupled DNA repair mechanism in eukaryotes.
- Jun Xu
- , Indrajit Lahiri
- & Dong Wang
-
Letter |
 Structure of the human MHC-I peptide-loading complex
Structure of the human MHC-I peptide-loading complex
Electron cryo-microscopy structures of the human peptide-loading complex shed light on its operation and on the onset of adaptive immune responses.
- Andreas Blees
- , Dovile Januliene
- & Robert Tampé
-
Article |
 Visualization of chemical modifications in the human 80S ribosome structure
Visualization of chemical modifications in the human 80S ribosome structure
A high-resolution structure of the human ribosome determined by cryo-electron microscopy visualizes numerous RNA modifications that are concentrated at functional sites with an extended shell, and suggests the possibility of designing more specific ribosome-targeting drugs.
- S. Kundhavai Natchiar
- , Alexander G. Myasnikov
- & Bruno P. Klaholz
-
Article |
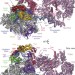 Structures of transcription pre-initiation complex with TFIIH and Mediator
Structures of transcription pre-initiation complex with TFIIH and Mediator
Cryo-electron microscopy structures of the yeast pre-initiation complex (PIC) and its complex with core Mediator provide insights into the opening of promoter DNA and the initiation of transcription.
- S. Schilbach
- , M. Hantsche
- & P. Cramer
-
Article |
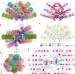 Structure of phycobilisome from the red alga Griffithsia pacifica
Structure of phycobilisome from the red alga Griffithsia pacifica
Single-particle cryo-electron microscopy is used to resolve the structure of the phycobilisome, a 16.8-megadalton light-harvesting megacomplex, from the red alga Griffithsia pacifica at a resolution of 3.5 Å.
- Jun Zhang
- , Jianfei Ma
- & Sen-Fang Sui

 Cryo-EM structure of the insect olfactory receptor Orco
Cryo-EM structure of the insect olfactory receptor Orco