Featured
-
-
Article
| Open Access Cre-Controlled CRISPR mutagenesis provides fast and easy conditional gene inactivation in zebrafish
Cre-Controlled CRISPR mutagenesis provides fast and easy conditional gene inactivation in zebrafish
Targeting a gene with two loxP sites is both time and labour intensive. Here the authors present Cre-Controlled CRISPR allowing conditional mutagenesis of a gene of interest and simultaneously labelling the putative mutant cells fluorescently.
- Stefan Hans
- , Daniela Zöller
- & Michael Brand
-
Article
| Open Access PrimeDesign software for rapid and simplified design of prime editing guide RNAs
PrimeDesign software for rapid and simplified design of prime editing guide RNAs
Prime editing guide RNA design is more complex than for standard CRISPR-based nucleases or base editors. Here the authors present PrimeDesign and PrimeVar for the rapid and simplified design of pegRNA and ngRNA combinations.
- Jonathan Y. Hsu
- , Julian Grünewald
- & Luca Pinello
-
Article
| Open Access Programmable human histone phosphorylation and gene activation using a CRISPR/Cas9-based chromatin kinase
Programmable human histone phosphorylation and gene activation using a CRISPR/Cas9-based chromatin kinase
Histone phosphorylation is a ubiquitous post-translational modification. Here the authors present a programmable chromatin kinase, dCas9-dMSK1, that enables controlled histone phosphorylation and specific gene activation.
- Jing Li
- , Barun Mahata
- & Isaac B. Hilton
-
Article
| Open Access In-situ generation of large numbers of genetic combinations for metabolic reprogramming via CRISPR-guided base editing
In-situ generation of large numbers of genetic combinations for metabolic reprogramming via CRISPR-guided base editing
To obtain optimal yield and productivity in bioproduction, expression of pathway genes must be appropriately coordinated. Here, the authors report repurposing of base editors for simultaneous regulation of multiple gene expression and demonstrate its application in industrially important and model microorganisms.
- Yu Wang
- , Haijiao Cheng
- & Yanhe Ma
-
Article
| Open Access Cas9-AAV6 gene correction of beta-globin in autologous HSCs improves sickle cell disease erythropoiesis in mice
Cas9-AAV6 gene correction of beta-globin in autologous HSCs improves sickle cell disease erythropoiesis in mice
CRISPR mediated gene correction of sickle cell disease (SCD) in patient-derived hematopoietic stem cells is a promising avenue for therapy. Here the authors use a humanized SCD mouse model to study gene editing in the context of autologous transplantation.
- Adam C. Wilkinson
- , Daniel P. Dever
- & Matthew H. Porteus
-
Article
| Open Access TALEN outperforms Cas9 in editing heterochromatin target sites
TALEN outperforms Cas9 in editing heterochromatin target sites
While Cas9 outperforms TALENs in euchromatin, it is less efficient in heterochromatic regions. Here the authors, using single-molecule imaging, show that Cas9 uses a less efficient search strategy compared to TALENs in these regions.
- Surbhi Jain
- , Saurabh Shukla
- & Huimin Zhao
-
Review Article
| Open Access CRISPR technologies and the search for the PAM-free nuclease
CRISPR technologies and the search for the PAM-free nuclease
One of the key limitations of CRISPR-Cas-based genome editing techniques is the PAM dependency. Here, the authors review ongoing efforts towards realizing PAM-free nucleases, address potential consequences of eliminating PAM recognition, and propose an alternative nuclease repertoire covering all possible PAM sequences.
- Daphne Collias
- & Chase L. Beisel
-
Article
| Open Access Engineered dual selection for directed evolution of SpCas9 PAM specificity
Engineered dual selection for directed evolution of SpCas9 PAM specificity
The PAM specificity of SpCas9 can be altered with positive selection during directed evolution. Here the authors use simultaneous positive and negative selection to improve activity on NAG PAMs while reducing activity on NGG PAMs.
- Gregory W. Goldberg
- , Jeffrey M. Spencer
- & Marcus B. Noyes
-
Article
| Open Access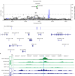 SLE non-coding genetic risk variant determines the epigenetic dysfunction of an immune cell specific enhancer that controls disease-critical microRNA expression
SLE non-coding genetic risk variant determines the epigenetic dysfunction of an immune cell specific enhancer that controls disease-critical microRNA expression
Enhancers shape gene expression patterns and are involved in disease pathogenesis. Here the authors demonstrate a strategy to screen functional regulatory elements for non-coding RNAs ― illustrated with miR-146a ― and link autoimmune disease risk genetic variants to autoimmune disease etiology.
- Guojun Hou
- , Isaac T. W. Harley
- & Nan Shen
-
Article
| Open Access Structure of human RNA polymerase III
Structure of human RNA polymerase III
The eukaryotic RNA Polymerase III transcribes tRNAs, some ribosomal and spliceosomal RNAs. Here, the authors resolve a cryo-EM structure of human RNA Polymerase III in its apo form and complemented it with crystal structures and SAXS analysis of RPC5, revealing insights into the molecular mechanisms of Pol III transcription.
- Ewan Phillip Ramsay
- , Guillermo Abascal-Palacios
- & Alessandro Vannini
-
Article
| Open Access The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
Acidothermus cellulolyticus CRISPR-Cas9 (AceCas9) is a Type II-C enzyme that cleaves DNA in a Protospacer Adjacent Motif (PAM) methylation sensitive fashion. Biochemical analysis and crystal structures of AceCas9 in complex with sgRNA and DNA bearing the correct and incorrect PAM offer insight into the structural basis for the recognition of PAM and its methylation.
- Anuska Das
- , Travis H. Hand
- & Hong Li
-
Article
| Open Access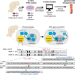 Design of efficacious somatic cell genome editing strategies for recessive and polygenic diseases
Design of efficacious somatic cell genome editing strategies for recessive and polygenic diseases
Gene correction of multiple alleles is a promising therapeutic strategy. Here the authors present bi-allelic correction of Pompe disease and provide an in silico model for predicting in vivo somatic editing outcomes.
- Jared Carlson-Stevermer
- , Amritava Das
- & Krishanu Saha
-
Article
| Open Access MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
Cas9 fused to DNA damage repair proteins can improve rates of gene knock-in but the chimeric protein is often large. Here the authors fuse Cas9 to a minimal Brex27 motif from BRCA2 consisting of thirty-six amino acids to enhance the efficacy and safety of gene editing.
- Linyuan Ma
- , Jinxue Ruan
- & Jie Xu
-
Article
| Open Access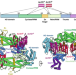 Structural basis for inhibition of an archaeal CRISPR–Cas type I-D large subunit by an anti-CRISPR protein
Structural basis for inhibition of an archaeal CRISPR–Cas type I-D large subunit by an anti-CRISPR protein
In type I-D CRISPR–Cas systems, the nuclease and helicase activities are carried out by separate subunits. The crystal structure of Sulfolobus islandicus type I-D large subunit Cas10d, containing a nuclease domain, reveals unusual architecture. The structure of Cas10d in complex with anti-CRISPR protein AcrID1 suggests that the latter sequesters Cas10d in a nonfunctional state.
- M. Cemre Manav
- , Lan B. Van
- & Ditlev E. Brodersen
-
Article
| Open Access Structural basis for assembly of non-canonical small subunits into type I-C Cascade
Structural basis for assembly of non-canonical small subunits into type I-C Cascade
Type I-C Cascade (the CRISPR-associated complex for antiviral defense) is a minimal system, comprising only three unique Cas proteins. Cryo-EM structure of the Desulfovibrio vulgaris type I-C Cascade reveals the molecular mechanisms that underlie RNA-directed complex assembly.
- Roisin E. O’Brien
- , Inês C. Santos
- & David W. Taylor
-
Article
| Open Access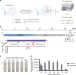 Streamlined inactivation, amplification, and Cas13-based detection of SARS-CoV-2
Streamlined inactivation, amplification, and Cas13-based detection of SARS-CoV-2
The COVID-19 pandemic has highlighted the need for user-friendly diagnostic techniques. Here, the authors present SHINE, a streamlined and optimised Cas13-based method with accompanying smartphone app for visual diagnosis.
- Jon Arizti-Sanz
- , Catherine A. Freije
- & Cameron Myhrvold
-
Article
| Open Access Discovery of multiple anti-CRISPRs highlights anti-defense gene clustering in mobile genetic elements
Discovery of multiple anti-CRISPRs highlights anti-defense gene clustering in mobile genetic elements
Mobile genetic elements have evolved anti-CRISPR (Acr) proteins to bypass the immunity provided by prokaryotic CRISPR–Cas systems. Here, the authors identify 11 type I Acrs encoded on mobile genetic elements, and show that acr loci neighborhoods can be used to discover inhibitors of other bacterial defense systems.
- Rafael Pinilla-Redondo
- , Saadlee Shehreen
- & Joseph Bondy-Denomy
-
Article
| Open Access Efficient population modification gene-drive rescue system in the malaria mosquito Anopheles stephensi
Efficient population modification gene-drive rescue system in the malaria mosquito Anopheles stephensi
Gene drives may be impeded by the generation of resistant alleles following NHEJ. Here the authors develop a recoded gene-drive rescue system for the malaria mosquito, Anopheles stephensi, that targets the drive to the kynurenine hydroxylase gene for negative selection against mutated alleles.
- Adriana Adolfi
- , Valentino M. Gantz
- & Anthony A. James
-
Article
| Open Access A catalogue of biochemically diverse CRISPR-Cas9 orthologs
A catalogue of biochemically diverse CRISPR-Cas9 orthologs
A few bacterial Cas9 nucleases have been repurposed as genome editing tools. Here, the authors use bioinformatic and biochemical analyses to characterize 79 Cas9 proteins, revealing substantial functional diversity and thus expanding the available toolbox of RNA-programmable CRISPR-associated nucleases.
- Giedrius Gasiunas
- , Joshua K. Young
- & Virginijus Siksnys
-
Article
| Open Access Structural basis for two metal-ion catalysis of DNA cleavage by Cas12i2
Structural basis for two metal-ion catalysis of DNA cleavage by Cas12i2
Cas12i, class 2 type V CRISPR-Cas system protein, uses a single RuvC domain for cleavage of both strands of target DNA. Structures of Cas12i2–crRNA–DNA complexes not only provide insight into the mechanism of DNA recognition and cleavage by Cas12i2, but also the DNA cleavage mechanism by RuvC-containing Cas proteins.
- Xue Huang
- , Wei Sun
- & Yanli Wang
-
Article
| Open Access CRISPRoff enables spatio-temporal control of CRISPR editing
CRISPRoff enables spatio-temporal control of CRISPR editing
Uncontrolled gene editing can lead to off-target effects and chromosomal translocations. Here the authors develop CRISPRoff, light degradable sgRNAs for titratable and spatially defined gene editing.
- Jared Carlson-Stevermer
- , Reed Kelso
- & Travis Maures
-
Article
| Open Access Tailoring poplar lignin without yield penalty by combining a null and haploinsufficient CINNAMOYL-CoA REDUCTASE2 allele
Tailoring poplar lignin without yield penalty by combining a null and haploinsufficient CINNAMOYL-CoA REDUCTASE2 allele
Plants with reduced amounts of lignin typically suffer from dwarfed growth, which offsets their gain in fermentable sugar yield. Here, the authors show that genome-edited poplar lines with a null and a haploinsufficient allele of CINNAMOYL-COA REDUCTASE2 (CCR2) can be obtained that have a reduced lignin level and normal growth.
- Barbara De Meester
- , Barbara Madariaga Calderón
- & Wout Boerjan
-
Article
| Open Access Viral gene drive in herpesviruses
Viral gene drive in herpesviruses
Current gene drive strategies are restricted to sexually reproducing species. Here the authors develop a gene drive in herpesviruses that allows the spread of an engineered trait through a viral population.
- Marius Walter
- & Eric Verdin
-
Article
| Open Access Engineering domain-inlaid SaCas9 adenine base editors with reduced RNA off-targets and increased on-target DNA editing
Engineering domain-inlaid SaCas9 adenine base editors with reduced RNA off-targets and increased on-target DNA editing
Off-target effects and the feasibility for AAV-mediated delivery are the major barriers impeding the clinical in vivo application of base editors. Here, the authors report the small size AAV-deliverable Cas9-ABE variant that has improved on-target editing efficiency and reduced RNA-off target footprint.
- Minh Thuan Nguyen Tran
- , Mohd Khairul Nizam Mohd Khalid
- & Alex W. Hewitt
-
Article
| Open Access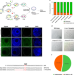 Precise allele-specific genome editing by spatiotemporal control of CRISPR-Cas9 via pronuclear transplantation
Precise allele-specific genome editing by spatiotemporal control of CRISPR-Cas9 via pronuclear transplantation
Injecting Cas9 and gRNA into an animal zygote often produces mosaicism and random biallelic targeting. Here, the authors use pronuclear transfer to reduce mosaicism and selectively target parental alleles.
- Yanhe Li
- , Yuteng Weng
- & Shaorong Gao
-
Article
| Open Access Concerted localization-resets precede YAP-dependent transcription
Concerted localization-resets precede YAP-dependent transcription
The transcriptional regulator YAP shuttles rapidly between the cytoplasm and nucleus, but whether and how dynamics such as amplitude and frequency affect target gene transcription is unclear. Here, using live imaging of endogenous YAP and target-gene transcription, the authors show that YAP-dependent signalling is encoded through rapid and concerted changes in the nucleo-cytoplasmic distribution of YAP.
- J. Matthew Franklin
- , Rajarshi P. Ghosh
- & Jan T. Liphardt
-
Article
| Open Access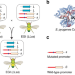 Engineering multiple species-like genetic incompatibilities in insects
Engineering multiple species-like genetic incompatibilities in insects
Natural speciation can be driven by pre-zygotic and post-zygotic barriers. Here the authors use a dominant lethal transgene coupled to a recessive resistance allele to engineer species-like barriers in Drosophila.
- Maciej Maselko
- , Nathan Feltman
- & Michael J. Smanski
-
Article
| Open Access Analysis of chromatin organization and gene expression in T cells identifies functional genes for rheumatoid arthritis
Analysis of chromatin organization and gene expression in T cells identifies functional genes for rheumatoid arthritis
Although genome-wide association studies have identified genetic variation contributing to disease risk, assigning causal genes is challenging. Here, the authors generate ATAC-seq, Hi-C, Capture Hi-C and RNA-seq data in stimulated CD4+ T cells to identify functional enhancers and demonstrate interactions of expression quantitative trait loci with target genes in rheumatoid arthritis.
- Jing Yang
- , Amanda McGovern
- & Stephen Eyre
-
Article
| Open Access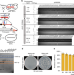 Chromosome drives via CRISPR-Cas9 in yeast
Chromosome drives via CRISPR-Cas9 in yeast
Self-propagating drives allow for non-Mendelian inheritance. Here the authors use CRISPR to build a chromosome drive, showing elimination of entire chromosomes, endoreduplication of desired chromosomes and enabling preferential transmissions of complex genetic traits on a chromosomal scale in yeast.
- Hui Xu
- , Mingzhe Han
- & Ying-Jin Yuan
-
Article
| Open Access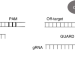 CRISPR GUARD protects off-target sites from Cas9 nuclease activity using short guide RNAs
CRISPR GUARD protects off-target sites from Cas9 nuclease activity using short guide RNAs
Off-target editing remains a concern for therapeutic applications of CRISPR-Cas9. Here the authors present CRISPR GUARD, which uses very short non-cleaving gRNAs to prevent editing at off-target sites.
- Matthew A. Coelho
- , Etienne De Braekeleer
- & Benjamin J. M. Taylor
-
Article
| Open Access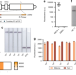 Deployable CRISPR-Cas13a diagnostic tools to detect and report Ebola and Lassa virus cases in real-time
Deployable CRISPR-Cas13a diagnostic tools to detect and report Ebola and Lassa virus cases in real-time
Outbreaks of viral hemorrhagic fevers highlight the need for sensitive, field-deployable diagnostics. Here the authors present a CRISPR-based SHERLOCK platform with field protocol and mobile app for Ebola and Lassa fever outbreaks.
- Kayla G. Barnes
- , Anna E. Lachenauer
- & Pardis C. Sabeti
-
Article
| Open Access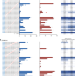 Genome-wide specificity of dCpf1 cytidine base editors
Genome-wide specificity of dCpf1 cytidine base editors
Cas12a-linked base editors can broaden the targeting scope of programmable cytidine deaminases. Here the authors assess their target specificity in an in vitro genome-wide assay.
- Daesik Kim
- , Kayeong Lim
- & Jin-Soo Kim
-
Article
| Open Access Engineering designer beta cells with a CRISPR-Cas9 conjugation platform
Engineering designer beta cells with a CRISPR-Cas9 conjugation platform
Cas9 fusions partners are often limited to natural polypeptide chains at the Cas9 termni. Here the authors present a platform for site-specific and multiple-site conjugation to both termini and internal sites of Cas9, and they apply this platform to efficiently engineer insulin-producing β cells.
- Donghyun Lim
- , Vedagopuram Sreekanth
- & Amit Choudhary
-
Article
| Open Access Engineering regulatory networks for complex phenotypes in E. coli
Engineering regulatory networks for complex phenotypes in E. coli
Regulatory networks respond to environmental and genetic perturbations by reprogramming metabolism. Here the authors screen a library of 82 regulators with 110,120 mutations to map the regulatory network of 4000 genes.
- Rongming Liu
- , Liya Liang
- & Ryan T. Gill
-
Article
| Open Access Ex vivo editing of human hematopoietic stem cells for erythroid expression of therapeutic proteins
Ex vivo editing of human hematopoietic stem cells for erythroid expression of therapeutic proteins
A platform for systemic therapeutic transgene expression independent of patient mutations needs a safe and highly transcribed locus. Here the authors ex vivo edit HPSCs using CRISPR-Cas9 to integrate transgenes under the α-globin promoter to achieve erythroid specific expression.
- Giulia Pavani
- , Marine Laurent
- & Mario Amendola
-
Article
| Open Access Machine-learning approach expands the repertoire of anti-CRISPR protein families
Machine-learning approach expands the repertoire of anti-CRISPR protein families
CRISPR-Cas is a host adaptive immunity system and viruses harbor diverse anti-CRISPR proteins (Acrs). Here, the authors develop a random forest machine-learning approach to predict Acrs, identifying 2500 candidate Acr families, which expand the current repertoire of predicted Acrs by two orders of magnitude.
- Ayal B. Gussow
- , Allyson E. Park
- & Eugene V. Koonin
-
Article
| Open Access Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment
Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment
Off-target mutations that occur at a frequency below 0.5% can be difficult to detect. Here the authors use predicted off-target amplification to increase detection sensitivity.
- Seung-Hun Kang
- , Wi-jae Lee
- & Seung Hwan Lee
-
Article
| Open Access Engineered CRISPR/Cas9 enzymes improve discrimination by slowing DNA cleavage to allow release of off-target DNA
Engineered CRISPR/Cas9 enzymes improve discrimination by slowing DNA cleavage to allow release of off-target DNA
Engineered high-fidelity Cas9s have increased discrimination against off-targets. Kinetic analyses of Cas9-HF1 and HypaCas9 engineered Cas9 variants show that their DNA cleavage is impaired by more than 100- fold, which leads to release rather than cleavage of a bound off-target substrate.
- Mu-Sen Liu
- , Shanzhong Gong
- & David W. Taylor
-
Article
| Open Access High-performance CRISPR-Cas12a genome editing for combinatorial genetic screening
High-performance CRISPR-Cas12a genome editing for combinatorial genetic screening
Reliable, multiplexed gene editing to uncover synergies between targets remains challenging. Here, the authors engineer AsCas12a and the crRNA to improve double knockout for synthetic sick/lethal interaction genetic screening.
- Rodrigo A. Gier
- , Krista A. Budinich
- & Junwei Shi
-
Article
| Open Access Engineering monocyte/macrophage−specific glucocerebrosidase expression in human hematopoietic stem cells using genome editing
Engineering monocyte/macrophage−specific glucocerebrosidase expression in human hematopoietic stem cells using genome editing
Gaucher disease is a lysosomal storage disorder caused by insufficient glucocerebrosidase expression. Here, the authors describe a CRISPR/Cas9-based gene-editing approach to re-express this enzyme in human blood stem cells and show that they can engraft in NSG mice and differentiate into functional macrophages.
- Samantha G. Scharenberg
- , Edina Poletto
- & Natalia Gomez-Ospina
-
Article
| Open Access Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing
Systemic nanoparticle delivery of CRISPR-Cas9 ribonucleoproteins for effective tissue specific genome editing
Therapeutic targets of CRISPR-Cas can often not be accessed due to lack of carriers to deliver RNPs systematically. Here, the authors engineer modified lipid nanoparticles for delivery of gene editing proteins to specific tissues.
- Tuo Wei
- , Qiang Cheng
- & Daniel J. Siegwart
-
Article
| Open Access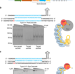 Repurposing type I–F CRISPR–Cas system as a transcriptional activation tool in human cells
Repurposing type I–F CRISPR–Cas system as a transcriptional activation tool in human cells
Class 1 type I CRISPR–Cas systems have not been as extensively developed for genome engineering as Class 2 systems. Here the authors modify the Type I–F CRISPR–Cas system for transcriptional activation of gene expression.
- Yuxi Chen
- , Jiaqi Liu
- & Zhou Songyang
-
Article
| Open Access CRISPR artificial splicing factors
CRISPR artificial splicing factors
Control over splicing could be used for both therapeutic and engineering applications. Here the authors create artificial splicing factors using RNA-targeting CRISPR systems under small molecule control.
- Menghan Du
- , Nathaniel Jillette
- & Albert Wu Cheng
-
Article
| Open Access Development of CRISPR-Cas13a-based antimicrobials capable of sequence-specific killing of target bacteria
Development of CRISPR-Cas13a-based antimicrobials capable of sequence-specific killing of target bacteria
CRISPR technology is emerging as a potential antimicrobial against antimicrobial-resistant bacteria. Here the authors develop a bacteriophage delivered Cas13a system for killing target bacteria and detecting bacterial genes.
- Kotaro Kiga
- , Xin-Ee Tan
- & Longzhu Cui
-
Article
| Open Access Synergistic gene editing in human iPS cells via cell cycle and DNA repair modulation
Synergistic gene editing in human iPS cells via cell cycle and DNA repair modulation
Precision editing with CRISPR-Cas9 often suffers from poor efficiency. Here the authors identify culture conditions and small molecules that synergize to promote homology-directed repair (HDR) in induced pluripotent stem (iPS) cells.
- Thomas L. Maurissen
- & Knut Woltjen
-
Article
| Open Access Multistable and dynamic CRISPRi-based synthetic circuits
Multistable and dynamic CRISPRi-based synthetic circuits
Synthetic circuits based on CRISPRi have not achieved multistable and dynamic behaviors. Here the authors build an oscillator, a toggle switch and an incoherent feed-forward loop using CRISPRi, and provide a mathematical model suggesting that unspecific binding in CRISPRi enables multistability.
- Javier Santos-Moreno
- , Eve Tasiudi
- & Yolanda Schaerli
-
Article
| Open Access Loss of the transcription factor MAFB limits β-cell derivation from human PSCs
Loss of the transcription factor MAFB limits β-cell derivation from human PSCs
The MAF bZIP transcription factor B (MAFB) is present in postnatal human beta cells but its role is unclear. Here, the authors show that MAFB regulates endocrine pancreatic cell fate specification.
- Ronan Russell
- , Phichitpol P. Carnese
- & Matthias Hebrok
-
Article
| Open Access Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards
Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards
Human pluripotent stem cell derived therapies can have serious safety risks. Here the authors design two drug inducible genetic safeguards to deplete undifferentiated hPSCs and hPSC-derived cell types.
- Renata M. Martin
- , Jonas L. Fowler
- & Kyle M. Loh
-
Article
| Open Access Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Numerous strategies exist to limit the off-target activity of CRISPR-Cas9 nucleases. Here the authors co-administer truncated gRNAs that block both Cas9 and high-fidelity Cas9 variants from cleaving at off-target sites.
- John C. Rose
- , Nicholas A. Popp
- & Douglas M. Fowler

 Accelerating target deconvolution for therapeutic antibody candidates using highly parallelized genome editing
Accelerating target deconvolution for therapeutic antibody candidates using highly parallelized genome editing