Featured
-
-
Article
| Open Access Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Suppression of unwanted CRISPR-Cas9 editing by co-administration of catalytically inactivating truncated guide RNAs
Numerous strategies exist to limit the off-target activity of CRISPR-Cas9 nucleases. Here the authors co-administer truncated gRNAs that block both Cas9 and high-fidelity Cas9 variants from cleaving at off-target sites.
- John C. Rose
- , Nicholas A. Popp
- & Douglas M. Fowler
-
Article
| Open Access Cas9-AAV6-engineered human mesenchymal stromal cells improved cutaneous wound healing in diabetic mice
Cas9-AAV6-engineered human mesenchymal stromal cells improved cutaneous wound healing in diabetic mice
Human mesenchymal stromal cells are a promising source for cell-based therapies. Here the authors use Cas9 to engineer lines that secrete PDGF-BB and VEFGA for improving wound healing.
- Waracharee Srifa
- , Nina Kosaric
- & Matthew Porteus
-
Article
| Open Access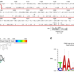 A Cas9 with PAM recognition for adenine dinucleotides
A Cas9 with PAM recognition for adenine dinucleotides
Protospacer adjacent motif (PAM) requirements limit the target range of CRISPR endonucleases. Here, the authors graft the 5\(^{\prime}\)-NAAN-3\(^{\prime}\) PAM-interacting domain of SmacCas9 onto SpyCas9 to create adenine dinucleotide targeting chimeras.
- Pranam Chatterjee
- , Jooyoung Lee
- & Noah Jakimo
-
Review Article
| Open Access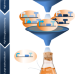 Application of combinatorial optimization strategies in synthetic biology
Application of combinatorial optimization strategies in synthetic biology
Our efforts to build complex synthetic biology circuits are impeded by limited knowledge of optimal combinations. In this review, the authors consider current combinatorial methods and look to emerging technologies.
- Gita Naseri
- & Mattheos A. G. Koffas
-
Article
| Open Access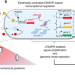 A redox-based electrogenetic CRISPR system to connect with and control biological information networks
A redox-based electrogenetic CRISPR system to connect with and control biological information networks
Redox-responsive transcriptional regulators can enable user-specified electronic control over biological functions. Here the authors demonstrate electronic control of CRISPRa and CRISPRi using redox signalling.
- Narendranath Bhokisham
- , Eric VanArsdale
- & William E. Bentley
-
Article
| Open Access Multiplex precise base editing in cynomolgus monkeys
Multiplex precise base editing in cynomolgus monkeys
Due to the polygenic nature of most diseases, simultaneous correction or introduction of single nucleotide variants is needed. Here, the authors demonstrated the feasibility of multiplex base editing for polygenes disease modeling in cynomolgus monkey embryos with high specificity.
- Wenhui Zhang
- , Tomomi Aida
- & Shihua Yang
-
Article
| Open Access Timed inhibition of CDC7 increases CRISPR-Cas9 mediated templated repair
Timed inhibition of CDC7 increases CRISPR-Cas9 mediated templated repair
Altering cellular responses to double-strand breaks in DNA could rebalance CRISPRediting outcomes. Here, the authors use a pooled CRISPR screen to identify inhibition of CDC7 as a strategy to improve HDR outcomes.
- Beeke Wienert
- , David N. Nguyen
- & Jacob E. Corn
-
Article
| Open Access Cytosine base editors with minimized unguided DNA and RNA off-target events and high on-target activity
Cytosine base editors with minimized unguided DNA and RNA off-target events and high on-target activity
Cytosine base editors have been reported to induce off-target mutations in DNA and RNA. Here the authors identify next-generation CBEs with reduced guide-independent off-target editing profiles and retain high on-target editing activity.
- Yi Yu
- , Thomas C. Leete
- & Nicole M. Gaudelli
-
Article
| Open Access Chemical modifications of adenine base editor mRNA and guide RNA expand its application scope
Chemical modifications of adenine base editor mRNA and guide RNA expand its application scope
Cas9 base editors are promising tools for correcting pathogenic single nucleotide mutations. Here the authors chemically modify mRNA encoding the editor and the gRNA to enhance editing and broaden its application.
- Tingting Jiang
- , Jordana M. Henderson
- & Wen Xue
-
Article
| Open Access Perturbing proteomes at single residue resolution using base editing
Perturbing proteomes at single residue resolution using base editing
Base editors allow for the precise modification of genes. Here the authors use Target-AID to systematically test 17,000 sites across the yeast genome.
- Philippe C. Després
- , Alexandre K. Dubé
- & Christian R. Landry
-
Article
| Open Access Pooled CRISPRi screening of the cyanobacterium Synechocystis sp PCC 6803 for enhanced industrial phenotypes
Pooled CRISPRi screening of the cyanobacterium Synechocystis sp PCC 6803 for enhanced industrial phenotypes
Developing cyanobacteria as CO2-neutral cell factories relies on the knowledge of the regulation mechanisms for growth and metabolism. Here, the authors develop an inducible CRISPRi gene repression library in Synechocystis sp. PCC 6803 and screens genes potentially affecting growth and L-lactate tolerance and production.
- Lun Yao
- , Kiyan Shabestary
- & Elton P. Hudson
-
Article
| Open Access Abundance of conserved CRISPR-Cas9 target sites within the highly polymorphic genomes of Anopheles and Aedes mosquitoes
Abundance of conserved CRISPR-Cas9 target sites within the highly polymorphic genomes of Anopheles and Aedes mosquitoes
Genetic variation in natural populations could represent gene drive resistant alleles, preventing successful application for population management. Here the authors survey 1280 genomes from three mosquito species and concludes natural variation will not be detrimental to deploying gene drive technology.
- Hanno Schmidt
- , Travis C. Collier
- & Gregory C. Lanzaro
-
Article
| Open Access Dissecting the early steps of MLL induced leukaemogenic transformation using a mouse model of AML
Dissecting the early steps of MLL induced leukaemogenic transformation using a mouse model of AML
The oncogene MLL is frequently translocated in leukemia, resulting in oncogenic fusion proteins. Here, the authors report a temporally controlled mouse model of MLL-ENL driven leukemia AND identify therapeutic targets associated with early MLL-ENL driven leukaemogenesis.
- Silvia Basilico
- , Xiaonan Wang
- & Berthold Göttgens
-
Article
| Open Access Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping
Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping
Expression of Cas9 and gRNA from viral vectors in vivo may cause off-target activity. Here the authors present NanoMEDIC, which uses nanovesicles to transiently deliver editing machinery to hard-to-transfect cells.
- Peter Gee
- , Mandy S. Y. Lung
- & Akitsu Hotta
-
Article
| Open Access Small molecule regulated sgRNAs enable control of genome editing in E. coli by Cas9
Small molecule regulated sgRNAs enable control of genome editing in E. coli by Cas9
Bacteria lack the same suite of CRISPR tools that have been developed for mammalian cells. Here, the authors link aptamers to sgRNAs to allow small molecule control of gene editing in E. coli.
- Roman S. Iwasaki
- , Bagdeser A. Ozdilek
- & Robert T. Batey
-
Article
| Open Access Prioritizing disease and trait causal variants at the TNFAIP3 locus using functional and genomic features
Prioritizing disease and trait causal variants at the TNFAIP3 locus using functional and genomic features
While genome-wide association studies have yielded thousands of trait-associated loci, identifying causal variants remains challenging. Here, the authors perform seven genomics assays in various cell types to prioritize genetic variants in the TNFAIP3 locus, and report high-priority variants within disease-associated haplotypes.
- John P. Ray
- , Carl G. de Boer
- & Nir Hacohen
-
Article
| Open Access Blackjack mutations improve the on-target activities of increased fidelity variants of SpCas9 with 5′G-extended sgRNAs
Blackjack mutations improve the on-target activities of increased fidelity variants of SpCas9 with 5′G-extended sgRNAs
Mutations to SpCas9 can improve fidelity and mitigate off-target effects. Here the authors generate ‘Blackjack’ mutations that improve fidelity while retaining effectiveness with 21G-sgRNAs.
- Péter István Kulcsár
- , András Tálas
- & Ervin Welker
-
Article
| Open Access Haploid genetic screens identify SPRING/C12ORF49 as a determinant of SREBP signaling and cholesterol metabolism
Haploid genetic screens identify SPRING/C12ORF49 as a determinant of SREBP signaling and cholesterol metabolism
The transcription factor SREBP is a well-studied and major regulator of sterol and fatty acid metabolism. Here, the authors used haploid genetic screens to identify the Golgi-resident protein SPRING as a new modulator of SREBP by regulating the level of functional SREBP cleavage-activating protein (SCAP).
- Anke Loregger
- , Matthijs Raaben
- & Noam Zelcer
-
Article
| Open Access A toxin-antidote CRISPR gene drive system for regional population modification
A toxin-antidote CRISPR gene drive system for regional population modification
CRISPR homing gene drives are highly invasive and can fail due to the rapid evolution of resistance. Here the authors present TARE drive, inspired by naturally occurring selfish genetic elements, which is less vulnerable to resistance and can potentially be confined to a target population.
- Jackson Champer
- , Esther Lee
- & Philipp W. Messer
-
Article
| Open Access A MAFG-lncRNA axis links systemic nutrient abundance to hepatic glucose metabolism
A MAFG-lncRNA axis links systemic nutrient abundance to hepatic glucose metabolism
Despite widespread transcription of LncRNA in mammalian systems, their contribution to metabolic homeostasis at the cellular and tissue level remains elusive. Here Pradas-Juni et al. describe a transcription factor–LncRNA pathway that couples hepatocyte nutrient sensing to regulation of glucose metabolism in mice.
- Marta Pradas-Juni
- , Nils R. Hansmeier
- & Jan-Wilhelm Kornfeld
-
Article
| Open Access Expanding the genome-targeting scope and the site selectivity of high-precision base editors
Expanding the genome-targeting scope and the site selectivity of high-precision base editors
Base editors can be limited by precision and the size of the target window. Here the authors test Cas9s that recognise alternative PAMs to obtain a series of high-precision editors.
- Junjie Tan
- , Fei Zhang
- & Ralph Bock
-
Article
| Open Access Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Tissues-specific gene expression requires coordinated cis-regulatory elements. Here the authors use dCas9-based enhancer targeting to remodel local epigenetic landscapes and activate or inactive transcription.
- Kailong Li
- , Yuxuan Liu
- & Jian Xu
-
Article
| Open Access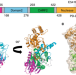 Structure and mechanism of a Type III CRISPR defence DNA nuclease activated by cyclic oligoadenylate
Structure and mechanism of a Type III CRISPR defence DNA nuclease activated by cyclic oligoadenylate
Antiviral defence type III CRISPR systems produce cyclic oligoadenylates (cOA) as second messengers that activate downstream effectors. Here the authors present the crystal structure of the type III CRISPR defence DNA nuclease Can1 in complex with cyclic tetra-adenylate (cA4) and show that Can1 nicks supercoiled DNA.
- Stephen A. McMahon
- , Wenlong Zhu
- & Tracey M. Gloster
-
Article
| Open Access A transcomplementing gene drive provides a flexible platform for laboratory investigation and potential field deployment
A transcomplementing gene drive provides a flexible platform for laboratory investigation and potential field deployment
Gene drives raise safety concerns around unintended propagation. Here the authors present a trans-complementing split-gene drive that requires inheritance of separate transgenes to assemble a fully functional drive.
- Víctor López Del Amo
- , Alena L. Bishop
- & Valentino M. Gantz
-
Article
| Open Access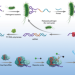 Sensitive detection of a bacterial pathogen using allosteric probe-initiated catalysis and CRISPR-Cas13a amplification reaction
Sensitive detection of a bacterial pathogen using allosteric probe-initiated catalysis and CRISPR-Cas13a amplification reaction
The detection of pathogens in food and clinical samples remains a challenge. Here, Shen et al. present a detection system, involving a combination of nucleic acid-based allosteric probes and CRISPR-Cas13a components, that can detect very low numbers of a bacterial pathogen in milk and serum samples without isolation.
- Jinjin Shen
- , Xiaoming Zhou
- & Da Xing
-
Article
| Open Access Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection
Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection
Here, Li et al. perform a genome-wide CRISPR screen to identify host dependency factors for influenza A virus infection and show that the host mRNA cap methyltransferase CMTR1 is important for viral cap snatching and that it affects expression of antiviral genes.
- Bo Li
- , Sara M. Clohisey
- & Nir Hacohen
-
Article
| Open Access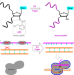 Conditional control of RNA-guided nucleic acid cleavage and gene editing
Conditional control of RNA-guided nucleic acid cleavage and gene editing
Constituitively active CRISPR systems have the risk of adverse off-target effects. Here the authors use chemical masking and activation of gRNA to control activity.
- Shao-Ru Wang
- , Ling-Yu Wu
- & Xiang Zhou
-
Article
| Open Access Multi-functional genome-wide CRISPR system for high throughput genotype–phenotype mapping
Multi-functional genome-wide CRISPR system for high throughput genotype–phenotype mapping
Genome-scale engineering is generally limited to single methods of alteration such as overexpression, repression or deletion. Here the authors present a tri-functional CRISPR system that can engineer complex synergistic interactions in a genome-wide manner.
- Jiazhang Lian
- , Carl Schultz
- & Huimin Zhao
-
Article
| Open Access A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus
A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus
Genedrives bias the inheritance of alleles in diploid organisms. Here, the authors develop a gene-drive analogous system for bacteria, selectively editing and clearing plasmids.
- J. Andrés Valderrama
- , Surashree S. Kulkarni
- & Ethan Bier
-
Article
| Open Access CRISPR-Cas3 induces broad and unidirectional genome editing in human cells
CRISPR-Cas3 induces broad and unidirectional genome editing in human cells
Class 1 CRISPR systems are not as developed for genome editing as Class 2 systems are. Here the authors show that Cas3 can be used to generate functional knockouts and knock-ins, as well as Cas3-mediated exon-skipping in DMD cells.
- Hiroyuki Morisaka
- , Kazuto Yoshimi
- & Tomoji Mashimo
-
Article
| Open Access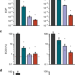 Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Bacteria use CRISPR-Cas systems to protect themselves against viral infections. Here, Watson et al. show that a type I CRISPR-Cas system can induce abortive viral infection, where infected cells do not survive but viral propagation is decreased, thus protecting the bacterial population.
- Bridget N. J. Watson
- , Reuben B. Vercoe
- & Peter C. Fineran
-
Article
| Open Access Trio deep-sequencing does not reveal unexpected off-target and on-target mutations in Cas9-edited rhesus monkeys
Trio deep-sequencing does not reveal unexpected off-target and on-target mutations in Cas9-edited rhesus monkeys
The off-target effects and on-target mutations of CRISPR editing in higher non-human primates are complex. Here the authors perform whole genome trio sequencing of rhesus monkeys and see no unexpected mutations.
- Xin Luo
- , Yaoxi He
- & Bing Su
-
Article
| Open Access Synthetic chimeric nucleases function for efficient genome editing
Synthetic chimeric nucleases function for efficient genome editing
CRISPR-Cas systems have well characterized, modular structures. Here the authors use that architecture to design a Cas12a library of 560 synthetic chimeras, with altered PAM preferences and specificities.
- R. M. Liu
- , L. L. Liang
- & R. T. Gill
-
Article
| Open Access CRISPR-Switch regulates sgRNA activity by Cre recombination for sequential editing of two loci
CRISPR-Switch regulates sgRNA activity by Cre recombination for sequential editing of two loci
Inducible genome editing systems often suffer from leakiness or reduced activity. Here the authors develop CRISPR-Switch, a Cre recombinase ON/OFF-controlled sgRNA cassette that allows consecutive editing of two loci.
- Krzysztof Chylinski
- , Maria Hubmann
- & Ulrich Elling
-
Article
| Open Access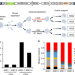 Targeting specificity of APOBEC-based cytosine base editor in human iPSCs determined by whole genome sequencing
Targeting specificity of APOBEC-based cytosine base editor in human iPSCs determined by whole genome sequencing
Cytidine base editors are powerful tools for making subtle genome alterations. Here the authors analyse edited human iPSCs with whole genome sequencing and reveal the spectrum of off-target effects.
- Erica McGrath
- , Hyunsu Shin
- & Zhaohui Ye
-
Article
| Open Access Highly efficient multiplex human T cell engineering without double-strand breaks using Cas9 base editors
Highly efficient multiplex human T cell engineering without double-strand breaks using Cas9 base editors
Multiplexed genome engineering with Cas9 can increase efficiency but also the risk of unintended alterations. Here the authors demonstrate the use of multiplexed base editors on primary T cells with reduced translocation frequency.
- Beau R. Webber
- , Cara-lin Lonetree
- & Branden S. Moriarity
-
Article
| Open Access Virus-borne mini-CRISPR arrays are involved in interviral conflicts
Virus-borne mini-CRISPR arrays are involved in interviral conflicts
Here, the authors investigate the diversity and dynamics of the CRISPRome in the hyperthermophilic archaea of the order Sulfolobales, and find the most abundant spacers to come from mini-CRISPR arrays of archaeal viruses, which might represent a strategy for superinfection exclusion and promotion of archaeal virus speciation.
- Sofia Medvedeva
- , Ying Liu
- & Mart Krupovic
-
Article
| Open Access Split selectable markers
Split selectable markers
Selectable markers are widely used in cell engineering but there is only a limited variety to choose from. Here the authors split markers using inteins, allowing up to six transgene integration events to be selected for with one marker.
- Nathaniel Jillette
- , Menghan Du
- & Albert Wu Cheng
-
Article
| Open Access Engineered amphiphilic peptides enable delivery of proteins and CRISPR-associated nucleases to airway epithelia
Engineered amphiphilic peptides enable delivery of proteins and CRISPR-associated nucleases to airway epithelia
Delivering biological cargo to airway epithelial cells is very challenging. Here, the authors use engineered amphiphilic peptides to shuttle proteins and CRISPR RNPs into airway cells in vivo.
- Sateesh Krishnamurthy
- , Christine Wohlford-Lenane
- & Paul B. McCray Jr.
-
Article
| Open Access Genome-wide microhomologies enable precise template-free editing of biologically relevant deletion mutations
Genome-wide microhomologies enable precise template-free editing of biologically relevant deletion mutations
DNA repair by microhomology-mediated end joining creates precise deletions based on flanking microhomologies. Here the authors use CRISPR-Cas9 to recreate pathogenic deletion mutations using existing microhomologies in the human genome identified by their program MHcut.
- Janin Grajcarek
- , Jean Monlong
- & Knut Woltjen
-
Article
| Open Access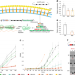 CRISPR-mediated gene silencing reveals involvement of the archaeal S-layer in cell division and virus infection
CRISPR-mediated gene silencing reveals involvement of the archaeal S-layer in cell division and virus infection
The S-layer is a proteinaceous envelope often found in bacterial and archaeal cells. Here, the authors use CRISPR-based technology to silence slaB, encoding the S-layer membrane anchor, to show that an intact S-layer is important for cell division and virus susceptibility in the archaeon Sulfolobus solfataricus.
- Isabelle Anna Zink
- , Kevin Pfeifer
- & Christa Schleper
-
Article
| Open Access Identification of atrial fibrillation associated genes and functional non-coding variants
Identification of atrial fibrillation associated genes and functional non-coding variants
The majority of disease-associated genetic variants lie in non-coding regions. Here the authors generated and compiled human transcriptomic, epigenomic and chromatin conformation datasets, to identify genes associated with atrial fibrillation and functional non-coding variants.
- Antoinette F. van Ouwerkerk
- , Fernanda M. Bosada
- & Vincent M. Christoffels
-
Article
| Open Access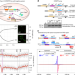 Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Detection of spacer precursors formed in vivo during primed CRISPR adaptation
Primed adaptation in the CRISPR-Cas system helps recognition of previously encountered sequence elements and promotes the formation of new memories. Here the authors characterized spacer precursors of type I-E and type I-F CRISPR-Cas system using in vivo models.
- Anna A. Shiriaeva
- , Ekaterina Savitskaya
- & Ekaterina Semenova
-
Article
| Open Access Efficient inter-species conjugative transfer of a CRISPR nuclease for targeted bacterial killing
Efficient inter-species conjugative transfer of a CRISPR nuclease for targeted bacterial killing
CRISPR nucleases can be programmed to cleave sequences in specific bacteria to induce cell death. Here, Hamilton et al. present an optimized method for conjugative delivery of CRISPR nucleases, consisting of a single plasmid that encodes both the conjugative machinery and the nuclease.
- Thomas A. Hamilton
- , Gregory M. Pellegrino
- & David R. Edgell
-
Article
| Open Access In vivo non-invasive monitoring of dystrophin correction in a new Duchenne muscular dystrophy reporter mouse
In vivo non-invasive monitoring of dystrophin correction in a new Duchenne muscular dystrophy reporter mouse
Dystrophin-deficient mice are used to test corrective strategies for Duchenne muscular dystrophy, but evaluation of dystrophin expression requires collection of tissue samples from specific muscles and time points. Here, the authors generate mice in which dystrophin expression is coupled to luciferase, and show that bioluminescence allows non-invasive monitoring of dystrophin expression following genome editing.
- Leonela Amoasii
- , Hui Li
- & Eric N. Olson
-
Article
| Open Access De novo identification of essential protein domains from CRISPR-Cas9 tiling-sgRNA knockout screens
De novo identification of essential protein domains from CRISPR-Cas9 tiling-sgRNA knockout screens
Tiling-sgRNA designs allow the in situ evaluation of protein domain functions. Here the authors present ProTiler - a computational method to predict CRISPR knockout hyper-sensitive regions, revealing previously unannotated domains.
- Wei He
- , Liang Zhang
- & Han Xu
-
Article
| Open Access High levels of AAV vector integration into CRISPR-induced DNA breaks
High levels of AAV vector integration into CRISPR-induced DNA breaks
In-depth characterization of adeno-associated virus (AAV)-mediated CRISPR delivery is still lacking. Here, the authors show high levels of integration into Cas9-induced double-strand breaks (DSBs) in therapeutically relevant genes in vivo.
- Killian S. Hanlon
- , Benjamin P. Kleinstiver
- & Bence György
-
Article
| Open Access Enhanced CRISPR-based DNA demethylation by Casilio-ME-mediated RNA-guided coupling of methylcytosine oxidation and DNA repair pathways
Enhanced CRISPR-based DNA demethylation by Casilio-ME-mediated RNA-guided coupling of methylcytosine oxidation and DNA repair pathways
DNA methylation plays an important role in regulating a wide variety of cellular processes and is implicated in a range of diseases. Here the authors present Casilio-ME to assemble protein complexes to demethylate target loci.
- Aziz Taghbalout
- , Menghan Du
- & Albert W. Cheng
-
Article
| Open Access Structure of Csx1-cOA4 complex reveals the basis of RNA decay in Type III-B CRISPR-Cas
Structure of Csx1-cOA4 complex reveals the basis of RNA decay in Type III-B CRISPR-Cas
Type III CRISPR-Cas RNases from the Csm and Csx families are activated by cyclic oligoadenylates (cOA). Here the authors present the cOA bound Sulfolobus islandicus Csx1 structure, which forms a hexamer and reveal an allosteric mechanism for the activation of Csx1 RNase.
- Rafael Molina
- , Stefano Stella
- & Guillermo Montoya

 Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards
Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards