Featured
-
-
Article
| Open Access Unsupervised classification of brain-wide axons reveals the presubiculum neuronal projection blueprint
Unsupervised classification of brain-wide axons reveals the presubiculum neuronal projection blueprint
The classification of different types of neurons has been a long-standing challenge in neuroscience. Here, the authors present a strategy to quantify all statistically distinct axonal patterns from a brain region based on their anatomical targeting, with this projection-driven neuron classification informing the functional architecture of the circuit.
- Diek W. Wheeler
- , Shaina Banduri
- & Giorgio A. Ascoli
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access Ultraconserved bacteriophage genome sequence identified in 1300-year-old human palaeofaeces
Ultraconserved bacteriophage genome sequence identified in 1300-year-old human palaeofaeces
Bacterial viruses (phages) are generally recognised as rapidly evolving biological entities. Here, Rozwalak et al. analyse DNA sequence datasets generated from ancient palaeofaeces and identify 298 phage genomes from the last 5300 years, including a 1300-year-old phage genome nearly identical to a present-day virus that infects human gut bacteria.
- Piotr Rozwalak
- , Jakub Barylski
- & Andrzej Zielezinski
-
Article
| Open Access Mesozoic evolution of cicadas and their origins of vocalization and root feeding
Mesozoic evolution of cicadas and their origins of vocalization and root feeding
The evolution of cicadas is unclear due to a lack of understanding of transitional features. Here, the authors assess adult and nymph mid-Cretaceous cicadas, to elucidate their morphological evolution and identify evidence of the origins of cicada sound-generation and subterranean lifestyle.
- Hui Jiang
- , Jacek Szwedo
- & Bo Wang
-
Article
| Open Access MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data
Identifying tissue structure in large-scale spatial omics datasets from multiple slices is challenging. Here, authors present MENDER, an optimisation-free spatial clustering method that can scale to million-level spatial data, enabling efficient analysis of spatial cell atlases.
- Zhiyuan Yuan
-
Article
| Open Access Data-driven grading of acute graft-versus-host disease
Data-driven grading of acute graft-versus-host disease
Acute GVHD severity grading is based on target organ assessments. Here, the authors show that data-driven grading can identify 12 distinct grades with specific aGVHD phenotypes, which are associated with clinical outcomes, and that their method outperformed conventional gradings.
- Evren Bayraktar
- , Theresa Graf
- & Amin T. Turki
-
Article
| Open Access MetaTiME integrates single-cell gene expression to characterize the meta-components of the tumor immune microenvironment
MetaTiME integrates single-cell gene expression to characterize the meta-components of the tumor immune microenvironment
Integration and comparison of multiple single cell sequencing datasets can be used to compare different studies. Here the authors propose MetaTiME which compares the gene expression of single cells from the tumour microenvironment across different tumours and uses transportable labels and metacomponents to annotate cell types and states.
- Yi Zhang
- , Guanjue Xiang
- & Clifford A. Meyer
-
Article
| Open Access Pan-cancer classification of single cells in the tumour microenvironment
Pan-cancer classification of single cells in the tumour microenvironment
The accuracy and granularity of classifying cell types in the tumour microenvironment (TME) from single-cell RNA-seq data is impacted by heterogeneity among cancer cells and similarities among functionally related immune cells. Here, the authors develop scATOMIC, a tumour and TME cell type classifier based on a hierarchical approach that can be applied to pan-cancer datasets.
- Ido Nofech-Mozes
- , David Soave
- & Sagi Abelson
-
Article
| Open Access Multilingual translation for zero-shot biomedical classification using BioTranslator
Multilingual translation for zero-shot biomedical classification using BioTranslator
Here, the authors develop the cross-modal translation method BioTranslator to translate the textual description to non-text biological data. This approach frees scientists from limiting their analysis within predefined controlled vocabularies.
- Hanwen Xu
- , Addie Woicik
- & Sheng Wang
-
Article
| Open Access Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny
Multiple sequence alignments are widely used to predict protein structure, function, and phylogeny, but are uncertain with more diverged sequences. Muscle5 generates ensembles of alternative high-accurate alignments, enabling novel confidence estimates in alignments, trees, and other inferences.
- Robert C. Edgar
-
Article
| Open Access De novo identification of microbial contaminants in low microbial biomass microbiomes with Squeegee
De novo identification of microbial contaminants in low microbial biomass microbiomes with Squeegee
Contaminant sequences in metagenomic samples can potentially impact the interpretation of findings reported in microbiome studies, especially in low biomass environments. Here the authors describe Squeegee, a computational approach designed to detect microbial contamination within low microbial biomass microbiomes and identify microbial contaminants in publicly available datasets that lack negative controls.
- Yunxi Liu
- , R. A. Leo Elworth
- & Todd J. Treangen
-
Article
| Open Access Discerning asthma endotypes through comorbidity mapping
Discerning asthma endotypes through comorbidity mapping
Asthma is a heterogeneous, complex syndrome that arises in individuals with various genetic and exposure variations. Here, the authors show that disease comorbidity patterns can serve as a surrogate for these variations, and identify asthma endotypes distinguished by comorbidity patterns, asthma risk loci, gene expression, and health-related phenotypes.
- Gengjie Jia
- , Xue Zhong
- & Julian Solway
-
Article
| Open Access Clustering by measuring local direction centrality for data with heterogeneous density and weak connectivity
Clustering by measuring local direction centrality for data with heterogeneous density and weak connectivity
Clustering is a powerful machine learning method for discovering similar patterns according to the proximity of elements in feature space. Here the authors propose a local direction centrality clustering algorithm that copes with heterogeneous density and weak connectivity issues.
- Dehua Peng
- , Zhipeng Gui
- & Huayi Wu
-
Article
| Open Access The rapid evolution of lungfish durophagy
The rapid evolution of lungfish durophagy
It is unclear how Lungfishes evolved durophagy, the consumption of hard prey, despite being the longest lineage of vertebrates with this feeding mechanism. Here, the authors describe exceptionally preserved fossils of Youngolepis from the Early Devonian, showing early adaptations to durophagy.
- Xindong Cui
- , Matt Friedman
- & Min Zhu
-
Article
| Open Access Genomic distances reveal relationships of wild and cultivated beets
Genomic distances reveal relationships of wild and cultivated beets
While a large amount of genomic resources is available, the phylogeny of wild and cultivated beets remains unclear. Here, the authors use the k-mer-based Mash method to analyze resequenced genomes of 606 accessions of the genus Beta and reveal Greece as the domestication site of sugar beet.
- Felix L. Sandell
- , Nancy Stralis-Pavese
- & Juliane C. Dohm
-
Article
| Open Access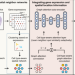 Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder
Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder
Breakthrough technologies for spatially resolved transcriptomics have enabled genome-wide profiling of gene expressions in captured locations. Here the authors integrate gene expressions and spatial locations to identify spatial domains using an adaptive graph attention auto-encoder.
- Kangning Dong
- & Shihua Zhang
-
Article
| Open Access Phylogenetically and functionally diverse microorganisms reside under the Ross Ice Shelf
Phylogenetically and functionally diverse microorganisms reside under the Ross Ice Shelf
The Ross Ice Shelf is the most extensive ice shelf of Antarctica and isolates the underlying ocean from sunlight. Here the authors use multi-omics to unravel the phylogenetic and functional diversity of microbial life in this ecosystem.
- Clara Martínez-Pérez
- , Chris Greening
- & Federico Baltar
-
Article
| Open Access Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Idiopathic pulmonary arterial hypertension is a rare and fatal disease with a heterogeneous treatment response. Here the authors show that unsupervised machine learning of whole blood transcriptomes from 359 patients with idiopathic pulmonary arterial hypertension identifies 3 subgroups (endophenotypes) that improve risk stratification and provide new molecular insights.
- Sokratis Kariotis
- , Emmanuel Jammeh
- & Richard C. Trembath
-
Article
| Open Access Extended antibody-framework-to-antigen distance observed exclusively with broad HIV-1-neutralizing antibodies recognizing glycan-dense surfaces
Extended antibody-framework-to-antigen distance observed exclusively with broad HIV-1-neutralizing antibodies recognizing glycan-dense surfaces
Here, the authors analyse the distance between the body of an antibody and a protein antigen denoted as the Antibody-Framework-to-Antigen Distance (AFAD) for about 2000 non-redundant antibody-protein antigen complexes in the Protein Data Bank. They observe that antibodies with exceptionally long AFADs were all broad HIV-1-neutralizing antibodies that targeted densely glycosylated regions on the HIV-1-envelope trimer. The connection between long AFAD and dense glycan was further validated by the cryo-EM structure of antibody 2909 recognizing a glycan hole and by glycan shielding analyses based on molecular dynamics simulations.
- Myungjin Lee
- , Anita Changela
- & Peter D. Kwong
-
Matters Arising
| Open Access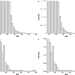 Re-evaluating the evidence for a universal genetic boundary among microbial species
Re-evaluating the evidence for a universal genetic boundary among microbial species
- Connor S. Murray
- , Yingnan Gao
- & Martin Wu
-
Matters Arising
| Open Access Reply to: “Re-evaluating the evidence for a universal genetic boundary among microbial species”
Reply to: “Re-evaluating the evidence for a universal genetic boundary among microbial species”
- Luis M. Rodriguez-R
- , Chirag Jain
- & Konstantinos T. Konstantinidis
-
Article
| Open Access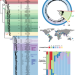 Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Whole genome sequencing is increasingly being adopted for Shigella sonnei outbreak investigation and surveillance, but there is no global classification standard. Here, the authors develop and validate a genomic framework implemented using open-source software, and demonstrate its application using surveillance data.
- Jane Hawkey
- , Kalani Paranagama
- & Kathryn E. Holt
-
Article
| Open Access Sarcoma classification by DNA methylation profiling
Sarcoma classification by DNA methylation profiling
Sarcomas are morphologically heterogeneous tumours rendering their classification challenging. Here the authors developed a classifier using DNA methylation data from several soft tissue and bone sarcoma subtypes, which has the potential to improve classification for research and clinical purposes.
- Christian Koelsche
- , Daniel Schrimpf
- & Andreas von Deimling
-
Article
| Open Access A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region
A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region
The human- and animal-adapted lineages of the Mycobacterium tuberculosis complex (MTBC) are thought to be evolved from a common progenitor in Africa. Here, the authors identify two MTBC strains isolated from patients with multidrug-resistant tuberculosis, representing an as-yet-unknown lineage further supporting an East African origin for the MTBC.
- Jean Claude Semuto Ngabonziza
- , Chloé Loiseau
- & Philip Supply
-
Article
| Open Access Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0
Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0
The increasing amount of sequenced microbial genomes and metagenomes requires platforms for efficient integrated analysis. Here, Asnicar et al. present PhyloPhlAn 3.0, a pipeline allowing large-scale microbial genome characterization and phylogenetic contextualization at multiple levels of resolution.
- Francesco Asnicar
- , Andrew Maltez Thomas
- & Nicola Segata
-
Article
| Open Access MethCORR modelling of methylomes from formalin-fixed paraffin-embedded tissue enables characterization and prognostication of colorectal cancer
MethCORR modelling of methylomes from formalin-fixed paraffin-embedded tissue enables characterization and prognostication of colorectal cancer
Molecular analysis of archival formalin-fixed clinical tissues can be difficult. Here, researchers have developed MethCORR, an approach that infers gene expression from DNA methylation data and use the approach for molecular characterization and prognostication of colorectal cancer using archival samples.
- Trine B. Mattesen
- , Mads H. Rasmussen
- & Jesper B. Bramsen
-
Article
| Open Access Re-definition of claudin-low as a breast cancer phenotype
Re-definition of claudin-low as a breast cancer phenotype
In breast cancer, the claudin-low breast cancer subtype is remarkably diverse. Here, the authors propose that claudin-low is not a classical intrinsic breast cancer subtype, but rather a complex additional phenotype that can occur across intrinsic subtypes.
- Christian Fougner
- , Helga Bergholtz
- & Therese Sørlie
-
Article
| Open Access Microbe-host interplay in atopic dermatitis and psoriasis
Microbe-host interplay in atopic dermatitis and psoriasis
Atopic dermatitis (AD) and psoriasis (PSO) are associated with dysbiosis. Here, by analyses of skin microbiome and host transcriptome of AD and PSO patients, the authors find distinct microbial and disease-related gene transcriptomic signatures that differentiate both diseases.
- Nanna Fyhrquist
- , Gareth Muirhead
- & Harri Alenius
-
Article
| Open Access Species abundance information improves sequence taxonomy classification accuracy
Species abundance information improves sequence taxonomy classification accuracy
Taxonomy classification of amplicon sequences is an important step in investigating microbial communities in microbiome analysis. Here, the authors show incorporating environment-specific taxonomic abundance information can lead to improved species-level classification accuracy across common sample types.
- Benjamin D. Kaehler
- , Nicholas A. Bokulich
- & Gavin A. Huttley
-
Article
| Open Access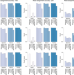 Strain-level metagenomic assignment and compositional estimation for long reads with MetaMaps
Strain-level metagenomic assignment and compositional estimation for long reads with MetaMaps
Sequencing platforms, such as Oxford Nanopore or Pacific Biosciences generate long-read data that preserve long-range genomic information but have high error rates. Here, the authors develop MetaMaps, a computational tool for strain-level metagenomic assignment and compositional estimation using long reads.
- Alexander T. Dilthey
- , Chirag Jain
- & Adam M. Phillippy
-
Article
| Open Access Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Morphological and molecular data have led to conflicting phylogenetic hypotheses for the Chelicerata. Here, the authors reconstruct the phylogeny of the Chelicerata using genomic-scale datasets, finding evidence for a monophyletic Acari and a single terrestrialisation of Arachnida.
- Jesus Lozano-Fernandez
- , Alastair R. Tanner
- & Davide Pisani
-
Article
| Open Access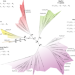 Broad phylogenetic analysis of cation/proton antiporters reveals transport determinants
Broad phylogenetic analysis of cation/proton antiporters reveals transport determinants
Cation/proton antiporters (CPAs) play a major role in maintaining living cells’ homeostasis and are divided in two main groups: CPA1 and CPA2. Here authors use a comprehensive evolutionary analysis of 6537 representative CPAs and reveal a sequence motif that determines central phenotypic characteristics.
- Gal Masrati
- , Manish Dwivedi
- & Nir Ben-Tal
-
Article
| Open Access Typing tumors using pathways selected by somatic evolution
Typing tumors using pathways selected by somatic evolution
Informative pathways driving cancer pathogenesis and subtypes can be difficult to identify in the presence of many gene interactions irrelevant to cancer. Here, the authors describe an approach for cancer gene pathway analysis based on key molecular interactions that drive cancer in relevant tissue types, and they assemble a focused map of Evolutionarily Selected Pathways (ESP) with interactions supported by both protein–protein binding and genetic epistasis during somatic tumor evolution.
- Sheng Wang
- , Jianzhu Ma
- & Trey Ideker
-
Article
| Open Access Maximal viral information recovery from sequence data using VirMAP
Maximal viral information recovery from sequence data using VirMAP
Viral taxonomic characterization from metagenomic data suffers from high background noise and signal crosstalk. Here, the authors develop VirMAP, a novel pipeline for analyses of metagenomic data that classifies viral reconstructions independent of genome coverage or read overlap.
- Nadim J Ajami
- , Matthew C. Wong
- & Joseph F. Petrosino
-
Article
| Open Access Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes
Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes
Germline mutations in homologous recombination (HR) DNA repair genes are linked to breast and ovarian cancer. Here, the authors show that mutually exclusive bi-allelic inactivation of HR genes are present in other cancer types and associated with genomic features of HR deficiency, expanding the potential use of HR-directed therapies.
- Nadeem Riaz
- , Pedro Blecua
- & Jorge S. Reis-Filho
-
Article
| Open Access Cancer-cell intrinsic gene expression signatures overcome intratumoural heterogeneity bias in colorectal cancer patient classification
Cancer-cell intrinsic gene expression signatures overcome intratumoural heterogeneity bias in colorectal cancer patient classification
Tumour expression profiling is currently used for prognostic and predictive purposes without taking into account the intra patient heterogeneity. Here the authors show that cancer cell specific signatures overcome the tumour heterogeneity effect and result in better classification of colorectal cancer patients.
- Philip D. Dunne
- , Matthew Alderdice
- & Mark Lawler
-
Article
| Open Access Fast and sensitive taxonomic classification for metagenomics with Kaiju
Fast and sensitive taxonomic classification for metagenomics with Kaiju
Here, Anders Krogh and colleagues describe Kaiju, a metagenome taxonomic classification program that uses maximum (in-)exact matches on the protein-level to account for evolutionary divergence. The authors show that Kaiju performs faster and is more sensitive compared with existing algorithms and can be used on a standard computer.
- Peter Menzel
- , Kim Lee Ng
- & Anders Krogh

 Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes
Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes