News & Views |
Featured
-
-
Article
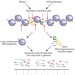
| Open Access Simultaneous single-cell three-dimensional genome and gene expression profiling uncovers dynamic enhancer connectivity underlying olfactory receptor choice
Simultaneous single-cell three-dimensional genome and gene expression profiling uncovers dynamic enhancer connectivity underlying olfactory receptor choice
LiMCA offers a tool for co-profiling 3D genome structure and gene expression at the single-cell level, enabling researchers to elucidate the olfactory receptor gene selection process.
- Honggui Wu
- , Jiankun Zhang
- & X. Sunney Xie
-
Article
| Open Access Combinatorial single-cell profiling of major chromatin types with MAbID
Combinatorial single-cell profiling of major chromatin types with MAbID
MAbID offers a multiplexing approach to uncover the genomic distributions of various epigenetic markers, enabling the study of how these markers jointly direct gene expression.
- Silke J. A. Lochs
- , Robin H. van der Weide
- & Jop Kind
-
Technology Feature |
 A tough ask: high-efficiency, large-cargo prime editing
A tough ask: high-efficiency, large-cargo prime editing
Researchers like the gene-editing method called prime editing for its precision and versatility. As they ask the method to do more, many factors shape what happens next.
- Vivien Marx
-
-
Article
| Open Access Multiplex-GAM: genome-wide identification of chromatin contacts yields insights overlooked by Hi-C
Multiplex-GAM: genome-wide identification of chromatin contacts yields insights overlooked by Hi-C
Multiplex-genome architecture mapping (multiplex-GAM) enables rapid, unbiased, ligation-free mapping of genome-wide chromatin interactions.
- Robert A. Beagrie
- , Christoph J. Thieme
- & Ana Pombo
-
Research Briefing |
 An in situ method for tagging chromatin-associated RNAs
An in situ method for tagging chromatin-associated RNAs
RNA comprises a substantial fraction of eukaryotic chromatin, but techniques to identify and map RNAs are cumbersome. We adapted existing tagmentation-based profiling techniques to enable chromatin-associated RNAs to be profiled in a simple workflow, enhancing the capability to identify regulatory RNAs.
-
Article
| Open Access Profiling RNA at chromatin targets in situ by antibody-targeted tagmentation
Profiling RNA at chromatin targets in situ by antibody-targeted tagmentation
This work presents RT&Tag, which captures RNA molecules in proximity to a protein of interest in intact nuclei.
- Nadiya Khyzha
- , Steven Henikoff
- & Kami Ahmad
-
Research Briefing |
 Sensitive, flexible and modular single-cell multi-omics profiling with ISSAAC-seq
Sensitive, flexible and modular single-cell multi-omics profiling with ISSAAC-seq
Joint profiling of multiple modalities in the same cell is challenging. We developed a method with a modular design to enable the simultaneous detection of chromatin accessibility and the transcriptome within single cells with flexible throughput.
-
Article |
 ISSAAC-seq enables sensitive and flexible multimodal profiling of chromatin accessibility and gene expression in single cells
ISSAAC-seq enables sensitive and flexible multimodal profiling of chromatin accessibility and gene expression in single cells
ISSAAC-seq offers a sensitive and modular tool for joint profiling gene expression and chromatin accessibility in single cells.
- Wei Xu
- , Weilong Yang
- & Xi Chen
-
Research Highlight |
 Sketching open and closed chromatin
Sketching open and closed chromatin
Single-cell GET-seq (scGET-seq) enables the probing of compacted as well as accessible chromatin in single cells, covering a greater portion of the genome, which provides insights into genomic and epigenetic dynamics.
- Lei Tang
-
Research Highlight |
 Arsenal of single-cell multi-omics methods expanded
Arsenal of single-cell multi-omics methods expanded
Chromatin accessibility, proteins levels and other features measured from the same cells.
- Lin Tang
-
Research Highlight |
 Walking through chromatin modifications
Walking through chromatin modifications
The Cell-TALKING technique probes DNA modifications around a histone modification in fixed cells.
- Lei Tang
-
Article |
 Simultaneous profiling of chromatin accessibility and methylation on human cell lines with nanopore sequencing
Simultaneous profiling of chromatin accessibility and methylation on human cell lines with nanopore sequencing
Using nanopore sequencing as readout, nanoNOMe–seq enables chromosome-level allele-specific profiles of CpG methylation and chromatin accessibility on human cells, in which the chromatin accessibility is profiled through exogenous GpC methylation introduced by a GpC methyltransferase.
- Isac Lee
- , Roham Razaghi
- & Winston Timp
-
Article |
 Long-range single-molecule mapping of chromatin accessibility in eukaryotes
Long-range single-molecule mapping of chromatin accessibility in eukaryotes
SMAC-seq combines long-read sequencing with open chromatin methylation by DNA methyltransferases to enable mapping of nucleosome position and chromatin accessibility.
- Zohar Shipony
- , Georgi K. Marinov
- & William J. Greenleaf
-
Research Highlight |
HACs with non-repetitive centromeres
Engineering human centromeres will allow versatile applications.
- Nicole Rusk
-
Brief Communication |
 Joint profiling of DNA methylation and chromatin architecture in single cells
Joint profiling of DNA methylation and chromatin architecture in single cells
Methyl-HiC combines the elucidation of chromatin architecture with the reading of DNA methylomes in pools and single cells. Regions that are distant on the linear-genome but close in three-dimensional space show coordinated DNA methylation.
- Guoqiang Li
- , Yaping Liu
- & Bing Ren
-
-
Research Highlight |
Variability in genome structure
Comparisons of genome organization in many individual cells with high-throughput FISH show extensive variation.
- Nicole Rusk
-
-
-
Correspondence |
 ATAC Primer Tool for targeted analysis of accessible chromatin
ATAC Primer Tool for targeted analysis of accessible chromatin
- Kathryn E Yost
- , Ava C Carter
- & Howard Y Chang
-
Methods in Brief |
Chromosome accessibility and methylation status in one
-
Brief Communication |
 An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues
An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues
The Omni-ATAC protocol improves the signal-to-background ratio in chromatin accessibility profiles and is suitable for a range of cell lines and primary cell types, as well as frozen tissue.
- M Ryan Corces
- , Alexandro E Trevino
- & Howard Y Chang
-
Research Highlights |
 Native chromosome conformation
Native chromosome conformation
Isolation of nuclei in an isotonic buffer retains chromosome loops and allows the probing of intrinsic loop conformation.
- Nicole Rusk
-
Review Article |
 How best to identify chromosomal interactions: a comparison of approaches
How best to identify chromosomal interactions: a comparison of approaches
In this Review, the authors compare commonly used chromosome conformation capture techniques, describing their respective strengths and weaknesses, and provide advice for the end user on which approach and analysis method to use.
- James O J Davies
- , A Marieke Oudelaar
- & Jim R Hughes
-
Brief Communication |
 Massively multiplex single-cell Hi-C
Massively multiplex single-cell Hi-C
Single-cell combinatorial indexed Hi-C (sciHi-C) is a streamlined protocol for generating thousands of high-quality single-cell chromosome conformation data sets that resemble bulk Hi-C data in aggregate.
- Vijay Ramani
- , Xinxian Deng
- & Jay Shendure
-
Brief Communication |
 HiChIP: efficient and sensitive analysis of protein-directed genome architecture
HiChIP: efficient and sensitive analysis of protein-directed genome architecture
HiChIP combines chromosome conformation capture with immunoprecipitation- and tagmentation-based library preparation to uncover the 3D chromatin architecture focused around a protein of interest.
- Maxwell R Mumbach
- , Adam J Rubin
- & Howard Y Chang
-
Methods in Brief |
Are super-enhancers really super?
-
Research Highlights |
A single cell's open chromatin
Increasing the sensitivity of DNase-seq allows chromatin accessibility to be profiled from very low numbers of cells.
- Nicole Rusk
-
Methods in Brief |
Chromatin accessibility in single cells
-
Research Highlights |
Tinkering with chromatin
Protein trans-splicing is used to study chromatin signaling in the cell nucleus.
- Allison Doerr
-
Methods in Brief |
Low-input ChIP for immunogenomics
-
News & Views |
 Chromatin biochemistry enters the next generation of code 'seq-ing'
Chromatin biochemistry enters the next generation of code 'seq-ing'
Barcoded semisynthetic nucleosomes combined with massively parallel sequencing provide an innovative new platform for analyzing the histone-recognition and histone-modifying activities of chromatin-associated proteins.
- Erin K Shanle
- , Scott B Rothbart
- & Brian D Strahl
-
This Month |
 Tom Muir
Tom Muir
From the netherworld between biology and chemistry comes a new method that leverages both fields and delivers acceleration to chromatin biochemistry.
- Vivien Marx
-
Article |
 Accelerated chromatin biochemistry using DNA-barcoded nucleosome libraries
Accelerated chromatin biochemistry using DNA-barcoded nucleosome libraries
A synthetic library of nucleosomes, each with a DNA-barcode characteristic for a defined post-translational modification (PTM), is used to probe PTM-based recruitment and modulation of histone mark readers and writers.
- Uyen T T Nguyen
- , Lenka Bittova
- & Tom W Muir
-
Methods in Brief |
Bias in ChIP-seq data
-
News & Views |
 The genome shows its sensitive side
The genome shows its sensitive side
New methods for measuring the sensitivity of chromatin to DNase digestion and Tn5 transposition help us map and interpret the genome's regulatory sequences.
- Anil Raj
- & Graham McVicker
-
Article |
 High-resolution mapping of transcription factor binding sites on native chromatin
High-resolution mapping of transcription factor binding sites on native chromatin
A method to map transcription factor binding across the genome, at high resolution and without cross-linking.
- Sivakanthan Kasinathan
- , Guillermo A Orsi
- & Steven Henikoff
-
Article |
 Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification
Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification
Detailed analysis of DNase-seq protocols reveals the importance of choosing the right enzyme concentration and fragment length and cautions that many transcription factor footprints may represent cutting bias.
- Housheng Hansen He
- , Clifford A Meyer
- & Myles Brown
-
Article |
 Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position
Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position
ATAC-seq queries the location of open chromatin, the binding of DNA-associated proteins and chromatin compaction at nucleotide resolution.
- Jason D Buenrostro
- , Paul G Giresi
- & William J Greenleaf
-
Methods in Brief |
Profiling chromatin at specific loci
-
Research Highlights |
 Writing the histone code
Writing the histone code
Recruiting chromatin modifiers to a specific locus allows the regulation of heterochromatin formation in living cells.
- Nicole Rusk
-
Brief Communication |
 Unsupervised pattern discovery in human chromatin structure through genomic segmentation
Unsupervised pattern discovery in human chromatin structure through genomic segmentation
Segway, a method using dynamic Bayesian network techniques, segments a genome and produces functional labels defined by histone modifications, transcription-factor binding, locations of open chromatin and other genome-wide functional data.
- Michael M Hoffman
- , Orion J Buske
- & William Stafford Noble
-
Correspondence |
 ChromHMM: automating chromatin-state discovery and characterization
ChromHMM: automating chromatin-state discovery and characterization
- Jason Ernst
- & Manolis Kellis

 LiMCA: Hi-C gets an RNA twist
LiMCA: Hi-C gets an RNA twist Diving deeper in the 3D genome
Diving deeper in the 3D genome