Featured
-
-
Article
| Open Access Precise atom-to-atom mapping for organic reactions via human-in-the-loop machine learning
Precise atom-to-atom mapping for organic reactions via human-in-the-loop machine learning
Precise atom mapping is crucial for data-driven reaction prediction, but currently lacks the required accuracy. Here, authors introduce a human-in-the-loop machine learning scheme for that purpose, and achieve high accuracy on a wide spectrum of reaction datasets.
- Shuan Chen
- , Sunggi An
- & Yousung Jung
-
Article
| Open Access Difficulty in chirality recognition for Transformer architectures learning chemical structures from string representations
Difficulty in chirality recognition for Transformer architectures learning chemical structures from string representations
There has been limited research on how NLP models comprehend diverse chemical structures despite its popularity. Here, the authors examine the learning process of Transformer for chemical structures and show inherent issues for chirality recognition.
- Yasuhiro Yoshikai
- , Tadahaya Mizuno
- & Hiroyuki Kusuhara
-
Article
| Open Access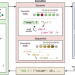 An integrated self-optimizing programmable chemical synthesis and reaction engine
An integrated self-optimizing programmable chemical synthesis and reaction engine
A limitation of robotic platforms in chemistry is the lack of feedback loops to adjust the conditions in-operando. Here the authors present a dynamically programmable robotic system that uses sensors for real-time adaptation, achieving yield improvements in syntheses and discovering new molecules.
- Artem I. Leonov
- , Alexander J. S. Hammer
- & Leroy Cronin
-
Article
| Open Access SQM2.20: Semiempirical quantum-mechanical scoring function yields DFT-quality protein–ligand binding affinity predictions in minutes
SQM2.20: Semiempirical quantum-mechanical scoring function yields DFT-quality protein–ligand binding affinity predictions in minutes
The paper presents the universal QM-based scoring function that accurately and rapidly predicts protein-ligand binding affinities, outperforming current computational tools. This is demonstrated on the PL-REX experimental benchmark dataset.
- Adam Pecina
- , Jindřich Fanfrlík
- & Jan Řezáč
-
Article
| Open Access Predictive Minisci late stage functionalization with transfer learning
Predictive Minisci late stage functionalization with transfer learning
Regioselectivity prediction for many reactions remains a challenging target for a priori prediction. Here, the authors develop a machine learning model that predicts the outcomes of Minisci reactions.
- Emma King-Smith
- , Felix A. Faber
- & Alpha A. Lee
-
Article
| Open Access On-tissue dataset-dependent MALDI-TIMS-MS2 bioimaging
On-tissue dataset-dependent MALDI-TIMS-MS2 bioimaging
There is a need for dataset-dependent MS2 acquisition in trapped ion mobility spectrometry imaging. Here the authors report spatial ion mobility-scheduled exhaustive fragmentation (SIMSEF) which enables on-tissue metabolite and lipid annotation in mass spectrometry bioimaging studies, and use this to visualise the chemical space in rat brains.
- Steffen Heuckeroth
- , Arne Behrens
- & Robin Schmid
-
Article
| Open Access Developing a class of dual atom materials for multifunctional catalytic reactions
Developing a class of dual atom materials for multifunctional catalytic reactions
This work developed a class of dual atom materials that can act as efficient and stable catalysts for multifunctional catalytic reactions in an uninterrupted water splitting system.
- Xingkun Wang
- , Liangliang Xu
- & Minghua Huang
-
Article
| Open Access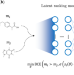 Extracting medicinal chemistry intuition via preference machine learning
Extracting medicinal chemistry intuition via preference machine learning
Over their careers, medicinal chemists develop a gut feeling for what is a promising molecule. Here, the authors use machine learning models to learn this intuition and show that it can be successfully applied in several drug discovery scenarios.
- Oh-Hyeon Choung
- , Riccardo Vianello
- & José Jiménez-Luna
-
Comment
| Open AccessLimitations of representation learning in small molecule property prediction
Machine learning is a powerful tool for the study and design of molecules. Here the authors comment a recent publication in Nature Communications which highlights the challenges of different molecular representations for data-driven property predictions.
- Ana Laura Dias
- , Latimah Bustillo
- & Tiago Rodrigues
-
Article
| Open Access A systematic study of key elements underlying molecular property prediction
A systematic study of key elements underlying molecular property prediction
AI has become a crucial tool for drug discovery, but how to properly represent molecules for data-driven property prediction is still an open question. Here the authors evaluate 62,820 models to highlight existing challenges, the impact of activity cliffs, and the crucial role of dataset size.
- Jianyuan Deng
- , Zhibo Yang
- & Fusheng Wang
-
Article
| Open Access Retrosynthesis prediction with an interpretable deep-learning framework based on molecular assembly tasks
Retrosynthesis prediction with an interpretable deep-learning framework based on molecular assembly tasks
Automating retrosynthesis prediction in organic chemistry is a major application of ML. Here the authors present RetroExplainer, which offers a high-performance, transparent and interpretable deep-learning framework providing valuable insights for drug development.
- Yu Wang
- , Chao Pang
- & Leyi Wei
-
Article
| Open Access First fully-automated AI/ML virtual screening cascade implemented at a drug discovery centre in Africa
First fully-automated AI/ML virtual screening cascade implemented at a drug discovery centre in Africa
Streamlined data-driven drug discovery remains challenging, especially in resource-limited settings. Here, the authors present ZairaChem, an AI/ML tool that streamlines QSAR/QSPR modelling, implemented for the first time at the H3D Centre in South Africa.
- Gemma Turon
- , Jason Hlozek
- & Miquel Duran-Frigola
-
Article
| Open Access DECIMER.ai: an open platform for automated optical chemical structure identification, segmentation and recognition in scientific publications
DECIMER.ai: an open platform for automated optical chemical structure identification, segmentation and recognition in scientific publications
Chemical structures are typically published as nonmachine-readable images in scientific literature. Here, the authors present DECIMER.ai, an open platform for translating chemical structures in publications into machine-readable representations.
- Kohulan Rajan
- , Henning Otto Brinkhaus
- & Christoph Steinbeck
-
Article
| Open Access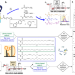 Ozone-enabled fatty acid discovery reveals unexpected diversity in the human lipidome
Ozone-enabled fatty acid discovery reveals unexpected diversity in the human lipidome
Fatty acids are fundamental biomolecular building blocks that are characterized by extraordinary structural diversity and present a formidable analytical challenge. Here the authors introduce a discovery workflow for de novo identification that adds more than 100 fatty acids to the human lipidome.
- Jan Philipp Menzel
- , Reuben S. E. Young
- & Stephen J. Blanksby
-
Article
| Open Access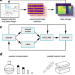 Rapid planning and analysis of high-throughput experiment arrays for reaction discovery
Rapid planning and analysis of high-throughput experiment arrays for reaction discovery
High-throughput experimentation is an increasingly important tool in reaction discovery, while there remains a need for software solutions to navigate data-rich experiments. Here the authors report phactor™, a software that facilitates the performance and analysis of high-throughput experimentation in a chemical laboratory.
- Babak Mahjour
- , Rui Zhang
- & Tim Cernak
-
Article
| Open Access Ultra-fast and accurate electron ionization mass spectrum matching for compound identification with million-scale in-silico library
Ultra-fast and accurate electron ionization mass spectrum matching for compound identification with million-scale in-silico library
Accuracy loss and slow speed affect the identification of compounds through matching of mass spectra using a large-scale spectral library. Here the authors use Word2vec spectral embedding and hierarchical navigable small-world graph to improve accuracy and speed of spectral matching on their own million-scale in-silico library.
- Qiong Yang
- , Hongchao Ji
- & Zhimin Zhang
-
Article
| Open Access Reaction performance prediction with an extrapolative and interpretable graph model based on chemical knowledge
Reaction performance prediction with an extrapolative and interpretable graph model based on chemical knowledge
Predictive modelling remains a key challenge for designing synthetic transformations. Here, the authors develop a knowledge-based graph model to predict reaction yield and stereoselectivity, offering an extrapolative and interpretable approach for evaluating reaction performance.
- Shu-Wen Li
- , Li-Cheng Xu
- & Xin Hong
-
Article
| Open Access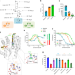 From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia
From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia
Deoxycytidine kinase is the rate-limiting enzyme of the salvage pathway and it has recently emerged as a target for antiproliferative therapies for cancers where it is essential. Here, the authors develop a potent inhibitor applying an iterative multidisciplinary approach, which relies on computational design coupled with experimental evaluations.
- Magali Saez-Ayala
- , Laurent Hoffer
- & Xavier Morelli
-
Article
| Open Access Retrosynthesis prediction using an end-to-end graph generative architecture for molecular graph editing
Retrosynthesis prediction using an end-to-end graph generative architecture for molecular graph editing
Retrosynthesis prediction is a fundamental problem in organic synthesis. Here, inspired by simplified arrow-pushing reaction mechanisms, the authors develop a graph-to-edits framework, Graph2Edits, based on graph neural network for retrosynthesis prediction.
- Weihe Zhong
- , Ziduo Yang
- & Calvin Yu-Chian Chen
-
Article
| Open Access Architector for high-throughput cross-periodic table 3D complex building
Architector for high-throughput cross-periodic table 3D complex building
Rare-earth and actinide complexes are critical for a wealth of clean-energy applications but Three dimensional (3D) structural generation and prediction for these organometallic systems remains challenging. Here, the authors propose a high-throughput in-silico synthesis code for s-, p-, d-, and f-block mononuclear organometallic complexes.
- Michael G. Taylor
- , Daniel J. Burrill
- & Ping Yang
-
Article
| Open Access Chemistry-intuitive explanation of graph neural networks for molecular property prediction with substructure masking
Chemistry-intuitive explanation of graph neural networks for molecular property prediction with substructure masking
Attempts to explain molecular property predictions of neural networks are not always compatible with chemical intuition based on chemical substructures. Here the authors propose the substructure mask explanation method to tackle this challenge.
- Zhenxing Wu
- , Jike Wang
- & Tingjun Hou
-
Article
| Open Access Single-step retrosynthesis prediction by leveraging commonly preserved substructures
Single-step retrosynthesis prediction by leveraging commonly preserved substructures
Retrosynthesis is a critical task for organic chemistry with numerous industrial applications. Here, the authors build a machine learning model to learn the concept of substructures from a large reaction dataset to achieve chemist-like intuitions.
- Lei Fang
- , Junren Li
- & Jian-Guang Lou
-
Article
| Open Access Predicting compound activity from phenotypic profiles and chemical structures
Predicting compound activity from phenotypic profiles and chemical structures
Experimental assays are used to determine if compounds cause a desired activity in cells. Here the authors demonstrate that computational methods can predict compound bioactivity given their chemical structure, imaging and gene expression data from historic screening libraries.
- Nikita Moshkov
- , Tim Becker
- & Juan C. Caicedo
-
Article
| Open Access Digital circuits and neural networks based on acid-base chemistry implemented by robotic fluid handling
Digital circuits and neural networks based on acid-base chemistry implemented by robotic fluid handling
The complementarity of acids and bases is a fundamental chemical concept. Here, the authors use simple acid-base chemistry to encode binary information and perform information processing including digital circuits and neural networks using robotic fluid handling.
- Ahmed A. Agiza
- , Kady Oakley
- & Sherief Reda
-
Article
| Open Access Leveraging molecular structure and bioactivity with chemical language models for de novo drug design
Leveraging molecular structure and bioactivity with chemical language models for de novo drug design
Generative Deep Learning holds promise for mining the unexplored “chemical universe” for new drugs. Here, the authors demonstrate the de novo design of phosphoinositide 3-kinase gamma (PI3Kγ) inhibitors for the PI3K/Akt pathway in human tumor cells.
- Michael Moret
- , Irene Pachon Angona
- & Gisbert Schneider
-
Article
| Open Access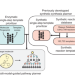 Merging enzymatic and synthetic chemistry with computational synthesis planning
Merging enzymatic and synthetic chemistry with computational synthesis planning
The identification of synthetic routes combining enzymatic and non-enzymatic reactions has been challenging and requiring expert knowledge. Here, the authors describe a computational retrosynthetic approach relying on neural network models for planning synthetic routes using both strategies.
- Itai Levin
- , Mengjie Liu
- & Connor W. Coley
-
Article
| Open Access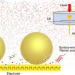 Unconventional interfacial water structure of highly concentrated aqueous electrolytes at negative electrode polarizations
Unconventional interfacial water structure of highly concentrated aqueous electrolytes at negative electrode polarizations
Water-in-salt electrolytes can be useful for future electrochemical energy storage systems. Here, the authors investigate the potential-dependent double-layer structures at the interface between a gold electrode and a highly concentrated aqueous electrolyte solution via in situ Raman measurements.
- Chao-Yu Li
- , Ming Chen
- & Tianquan Lian
-
Article
| Open Access Impedance-based forecasting of lithium-ion battery performance amid uneven usage
Impedance-based forecasting of lithium-ion battery performance amid uneven usage
Accurate forecasts of lithium-ion battery performance will ease concerns about the reliability of electric vehicles. Here, the authors leverage electrochemical impedance spectroscopy and machine learning to show that future capacity can be predicted amid uneven use, with no historical data requirement.
- Penelope K. Jones
- , Ulrich Stimming
- & Alpha A. Lee
-
Article
| Open Access Language models can learn complex molecular distributions
Language models can learn complex molecular distributions
Generative models for the novo molecular design attract enormous interest for exploring the chemical space. Here the authors investigate the application of chemical language models to challenging modeling tasks demonstrating their capability of learning complex molecular distributions.
- Daniel Flam-Shepherd
- , Kevin Zhu
- & Alán Aspuru-Guzik
-
Article
| Open Access The pocketome of G-protein-coupled receptors reveals previously untargeted allosteric sites
The pocketome of G-protein-coupled receptors reveals previously untargeted allosteric sites
G-protein-coupled receptors bind endogenous ligands at sites that are frequently highly conserved. Here, authors computationally describe alternative allosteric pockets, several of which have not been targeted by synthetic ligands before.
- Janik B. Hedderich
- , Margherita Persechino
- & Peter Kolb
-
Article
| Open Access SIMILE enables alignment of tandem mass spectra with statistical significance
SIMILE enables alignment of tandem mass spectra with statistical significance
Interrelating metabolites by their fragmentation spectra is central to metabolomics. Here the authors align fragmentation spectra with both statistical significance and allowance for multiple chemical differences using Significant Interrelation of MS/MS Ions via Laplacian Embedding (SIMILE).
- Daniel G. C. Treen
- , Mingxun Wang
- & Benjamin P. Bowen
-
Article
| Open Access Implicitly perturbed Hamiltonian as a class of versatile and general-purpose molecular representations for machine learning
Implicitly perturbed Hamiltonian as a class of versatile and general-purpose molecular representations for machine learning
Molecular representations are fundamental tools for machine-learning models. The current work introduces a new set of molecular representations demonstrated to enable accurate predictions of molecular conformational energy and solvation free energy.
- Amin Alibakhshi
- & Bernd Hartke
-
Article
| Open Access Retrosynthetic reaction pathway prediction through neural machine translation of atomic environments
Retrosynthetic reaction pathway prediction through neural machine translation of atomic environments
Reaction route planning remains a major challenge in organic synthesis. The authors present a retrosynthetic prediction model using the fragment-based representation of molecules and the Transformer architecture in neural machine translation.
- Umit V. Ucak
- , Islambek Ashyrmamatov
- & Juyong Lee
-
Comment
| Open Access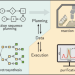 Autonomous platforms for data-driven organic synthesis
Autonomous platforms for data-driven organic synthesis
Achieving autonomous multi-step synthesis of novel molecular structures in chemical discovery processes is a goal shared by many researchers. In this Comment, we discuss key considerations of what an ideal platform may look like and the apparent state of the art. While most hardware challenges can be overcome with clever engineering, other challenges will require advances in both algorithms and data curation.
- Wenhao Gao
- , Priyanka Raghavan
- & Connor W. Coley
-
Article
| Open Access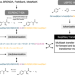 Biocatalysed synthesis planning using data-driven learning
Biocatalysed synthesis planning using data-driven learning
As of now, only rule-based systems support retrosynthetic planning using biocatalysis, while initial data-driven approaches are limited to forward predictions. Here, the authors extend the data-driven forward reaction as well as retrosynthetic pathway prediction models based on the Molecular Transformer architecture to biocatalysis.
- Daniel Probst
- , Matteo Manica
- & Teodoro Laino
-
Article
| Open Access Reinforcing the supply chain of umifenovir and other antiviral drugs with retrosynthetic software
Reinforcing the supply chain of umifenovir and other antiviral drugs with retrosynthetic software
COVID-19 has exposed the fragility of supply chains, particularly for goods that are essential or may suddenly become essential, such as repurposed pharmaceuticals. Here the authors develop a methodology to provide routes to pharmaceutical targets that allow low-supply starting materials or intermediates to be avoided, with representative pathways validated experimentally.
- Yingfu Lin
- , Zirong Zhang
- & Tim Cernak
-
Article
| Open Access SpookyNet: Learning force fields with electronic degrees of freedom and nonlocal effects
SpookyNet: Learning force fields with electronic degrees of freedom and nonlocal effects
Current machine-learned force fields typically ignore electronic degrees of freedom. SpookyNet is a deep neural network that explicitly treats electronic degrees of freedom, closing an important remaining gap for models in quantum chemistry.
- Oliver T. Unke
- , Stefan Chmiela
- & Klaus-Robert Müller
-
Article
| Open Access Coupling complementary strategy to flexible graph neural network for quick discovery of coformer in diverse co-crystal materials
Coupling complementary strategy to flexible graph neural network for quick discovery of coformer in diverse co-crystal materials
Experimental determination of new cocrystals remains challenging due to the need of a systematic screening with a large range of coformers. Here the authors develop a flexible deep learning framework based on graph neural network demonstrated to quickly predict the formation of co-crystals.
- Yuanyuan Jiang
- , Zongwei Yang
- & Xuemei Pu
-
Article
| Open Access Teaching a neural network to attach and detach electrons from molecules
Teaching a neural network to attach and detach electrons from molecules
Quantum mechanical calculations of molecular ionized states are computationally quite expensive. This work reports a successful extension of a previous deep-neural networks approach towards transferable neural-network models for predicting multiple properties of open shell anions and cations.
- Roman Zubatyuk
- , Justin S. Smith
- & Olexandr Isayev
-
Article
| Open Access Machine learning based energy-free structure predictions of molecules, transition states, and solids
Machine learning based energy-free structure predictions of molecules, transition states, and solids
Accurate computational prediction of atomistic structure with traditional methods is challenging. The authors report a kernel-based machine learning model capable of reconstructing 3D atomic coordinates from predicted interatomic distances across a variety of system classes.
- Dominik Lemm
- , Guido Falk von Rudorff
- & O. Anatole von Lilienfeld
-
Article
| Open Access Bioactivity descriptors for uncharacterized chemical compounds
Bioactivity descriptors for uncharacterized chemical compounds
Small molecules bioactivity descriptors are enriched representations of compounds, reaching beyond chemical structures and capturing their known biological properties. Here the authors present a collection of deep neural networks able to infer bioactivity signatures for any compound of interest, even when little or no experimental information is available for them.
- Martino Bertoni
- , Miquel Duran-Frigola
- & Patrick Aloy
-
Article
| Open Access A robotic prebiotic chemist probes long term reactions of complexifying mixtures
A robotic prebiotic chemist probes long term reactions of complexifying mixtures
The transition of prebiotic chemistry to present-day chemistry lasted a very long period of time, but the current laboratory investigations of this process are mostly limited to a couple of days. Here, the authors develop a fully automated robotic prebiotic chemist designed for long-term chemical experiments exploring unconstrained multicomponent reactions, which can run autonomously and uses simple chemical inputs.
- Silke Asche
- , Geoffrey J. T. Cooper
- & Leroy Cronin
-
Article
| Open Access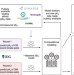 Crowdsourced mapping of unexplored target space of kinase inhibitors
Crowdsourced mapping of unexplored target space of kinase inhibitors
The IDG-DREAM Challenge carried out crowdsourced benchmarking of predictive algorithms for kinase inhibitor activities on unpublished data. This study provides a resource to compare emerging algorithms and prioritize new kinase activities to accelerate drug discovery and repurposing efforts.
- Anna Cichońska
- , Balaguru Ravikumar
- & Tero Aittokallio
-
Article
| Open Access Masked graph modeling for molecule generation
Masked graph modeling for molecule generation
Generating new sensible molecular structures is a key problem in computer aided drug discovery. Here the authors propose a graph-based molecular generative model that outperforms previously proposed graph-based generative models of molecules and performs comparably to several SMILES-based models.
- Omar Mahmood
- , Elman Mansimov
- & Kyunghyun Cho
-
Article
| Open Access Identifying molecules as biosignatures with assembly theory and mass spectrometry
Identifying molecules as biosignatures with assembly theory and mass spectrometry
The search for life in the universe is difficult due to issues with defining signatures of living systems. Here, the authors present an approach based on the molecular assembly number and tandem mass spectrometry that allows identification of molecules produced by biological systems, and use it to identify biosignatures from a range of samples, including ones from outer space.
- Stuart M. Marshall
- , Cole Mathis
- & Leroy Cronin
-
Article
| Open Access Inferring experimental procedures from text-based representations of chemical reactions
Inferring experimental procedures from text-based representations of chemical reactions
In organic chemistry, synthetic routes for new molecules are often specified in terms of reacting molecules only. The current work reports an artificial intelligence model to predict the full sequence of experimental operations for an arbitrary chemical equation.
- Alain C. Vaucher
- , Philippe Schwaller
- & Teodoro Laino
-
Article
| Open Access Bias free multiobjective active learning for materials design and discovery
Bias free multiobjective active learning for materials design and discovery
Identifying optimal materials in multiobjective optimization problems represents a challenge for new materials design approaches. Here the authors develop an active-learning algorithm to optimize the Pareto-optimal solutions successfully applied to the in silico polymer design for a dispersant-based application.
- Kevin Maik Jablonka
- , Giriprasad Melpatti Jothiappan
- & Brian Yoo
-
Article
| Open Access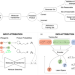 Quantitative interpretation explains machine learning models for chemical reaction prediction and uncovers bias
Quantitative interpretation explains machine learning models for chemical reaction prediction and uncovers bias
Machine learning algorithms offer new possibilities for automating reaction procedures. The present paper investigates automated reaction’s prediction with Molecular Transformer, the state-of-the-art model for reaction prediction, proposing a new debiased dataset for a realistic assessment of the model’s performance.
- Dávid Péter Kovács
- , William McCorkindale
- & Alpha A. Lee
-
Article
| Open Access Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences
Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences
Large-scale sequencing efforts have uncovered a large number of secondary metabolic pathways, but the chemicals they synthesise remain unknown. Here the authors present PRISM 4, which predicts the chemical structures encoded by microbial genome sequences, including all classes of bacterial antibiotics in clinical use.
- Michael A. Skinnider
- , Chad W. Johnston
- & Nathan A. Magarvey

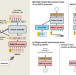 Bidirectional generation of structure and properties through a single molecular foundation model
Bidirectional generation of structure and properties through a single molecular foundation model