Article
|
Open Access
Featured
-
-
Article
| Open Access Whole-cortex in situ sequencing reveals input-dependent area identity
Whole-cortex in situ sequencing reveals input-dependent area identity
BARseq interrogates the expression of 104 cell-type marker genes in 10.3 million cells over nine mouse forebrain hemispheres to reveal the role of peripheral inputs on cortical area development.
- Xiaoyin Chen
- , Stephan Fischer
- & Anthony M. Zador
-
Article
| Open Access Cellular development and evolution of the mammalian cerebellum
Cellular development and evolution of the mammalian cerebellum
Single-nucleus RNA-sequencing data from the cerebellum of human, mouse and opossum is used to analyse the developmental dynamics of cell types and states in mammalian cerebellum and provide evolutionary insights.
- Mari Sepp
- , Kevin Leiss
- & Henrik Kaessmann
-
Article |
 Molecular landscapes of human hippocampal immature neurons across lifespan
Molecular landscapes of human hippocampal immature neurons across lifespan
Single-nucleus RNA-sequencing analysis supports the presence of immature dentate granule cells throughout the human lifespan and shows that these cells are reduced in number and dysregulated in Alzheimer's disease.
- Yi Zhou
- , Yijing Su
- & Hongjun Song
-
Article |
 Tbx2 is a master regulator of inner versus outer hair cell differentiation
Tbx2 is a master regulator of inner versus outer hair cell differentiation
Tbx2 is a master regulator of cochlear inner hair cells.
- Jaime García-Añoveros
- , John C. Clancy
- & Anne Duggan
-
Article |
 A complete temporal transcription factor series in the fly visual system
A complete temporal transcription factor series in the fly visual system
A complex regulatory network of temporally expressed transcription factors in Drosophila optic lobe stem cells regulates the generation of all neuronal diversity.
- Nikolaos Konstantinides
- , Isabel Holguera
- & Claude Desplan
-
Article
| Open Access The development and evolution of inhibitory neurons in primate cerebrum
The development and evolution of inhibitory neurons in primate cerebrum
Evolutionary modelling shows that an initial set of inhibitory neurons serving olfactory bulbs may have been repurposed to diversify the taxonomy of interneurons found in the expanded striata and cortices in primates.
- Matthew T. Schmitz
- , Kadellyn Sandoval
- & Alex A. Pollen
-
Article
| Open Access Single-cell delineation of lineage and genetic identity in the mouse brain
Single-cell delineation of lineage and genetic identity in the mouse brain
Single-cell RNA sequencing with parallel tagging of progenitor cells is used to track clonal relationships and transcriptomic signatures during development of the mouse forebrain.
- Rachel C. Bandler
- , Ilaria Vitali
- & Christian Mayer
-
Article |
 Temporal controls over inter-areal cortical projection neuron fate diversity
Temporal controls over inter-areal cortical projection neuron fate diversity
Combined analysis of the connectome and transcriptome in the mouse cortex indicates that dynamic differences in expression levels of largely generic sets of genes regulate differential targeting within neuronal subtypes.
- Esther Klingler
- , Ugo Tomasello
- & Denis Jabaudon
-
Article |
 Neuronal diversity and convergence in a visual system developmental atlas
Neuronal diversity and convergence in a visual system developmental atlas
The neuronal diversity of the Drosophila optic lobe is described throughout pupal development by single-cell sequencing, leading to the discovery of transient extrinsic neurons and a dorsoventral asymmetry of the visual circuits.
- Mehmet Neset Özel
- , Félix Simon
- & Claude Desplan
-
Article |
 Unique homeobox codes delineate all the neuron classes of C. elegans
Unique homeobox codes delineate all the neuron classes of C. elegans
Each one of the complete set of 118 neuron classes in Caenorhabditis elegans can be specified individually by the expression of unique combinations of homeodomain proteins.
- Molly B. Reilly
- , Cyril Cros
- & Oliver Hobert
-
Article |
 Molecular design of hypothalamus development
Molecular design of hypothalamus development
Single-cell RNA sequencing reveals molecular determinants of the developmental programs that orchestrate the intermingling of neuronal subtypes in the hypothalamus.
- Roman A. Romanov
- , Evgenii O. Tretiakov
- & Tibor Harkany
-
Article |
 Cell stress in cortical organoids impairs molecular subtype specification
Cell stress in cortical organoids impairs molecular subtype specification
Single-cell RNA sequencing clarifies the development and specification of neurons in the human cortex and shows that cell stress impairs this process in cortical organoids.
- Aparna Bhaduri
- , Madeline G. Andrews
- & Arnold R. Kriegstein
-
Letter |
 Individual brain organoids reproducibly form cell diversity of the human cerebral cortex
Individual brain organoids reproducibly form cell diversity of the human cerebral cortex
Single-cell RNA-sequencing analysis demonstrates that individual human brain organoids generate the cellular diversity of the cerebral cortex with organoid-to-organoid variability that is comparable to that of individual endogenous brains.
- Silvia Velasco
- , Amanda J. Kedaigle
- & Paola Arlotta
-
Letter |
 Subcellular transcriptomes and proteomes of developing axon projections in the cerebral cortex
Subcellular transcriptomes and proteomes of developing axon projections in the cerebral cortex
A subcellular sorting approach enables quantitative analysis of subtypes of growth cones in the brain, and reveals subcellular relationships between local mRNA and local proteomes in developing projection neurons.
- Alexandros Poulopoulos
- , Alexander J. Murphy
- & Jeffrey D. Macklis
-
Article |
 LHX2- and LDB1-mediated trans interactions regulate olfactory receptor choice
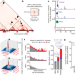
LHX2- and LDB1-mediated trans interactions regulate olfactory receptor choice
Specific interchromosomal contacts in olfactory sensory neurons form a super-enhancer that controls the expression of a single olfactory receptor in each neuron.
- Kevin Monahan
- , Adan Horta
- & Stavros Lomvardas
-
Letter |
 A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex
A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex
WebAnalysis of gene expression at single-cell resolution in the developing prefrontal cortex of the human embryo reveals a diversity of cell types, elucidates cell lineages and identifies signalling pathways that regulate development.
- Suijuan Zhong
- , Shu Zhang
- & Xiaoqun Wang
-
Article |
 Integration of temporal and spatial patterning generates neural diversity
Integration of temporal and spatial patterning generates neural diversity
Combinatorial inputs from temporal and spatial axes act together to promote medullary neural diversity in the optic lobes of Drosophila.
- Ted Erclik
- , Xin Li
- & Claude Desplan
-
Letter |
 Early myeloid lineage choice is not initiated by random PU.1 to GATA1 protein ratios
Early myeloid lineage choice is not initiated by random PU.1 to GATA1 protein ratios
Live imaging and single-cell analyses are used to show that decision-making by differentiating haematopoietic stem cells between the megakaryocytic–erythroid and granulocytic–monocytic lineages is not initiated by stochastic switching between the lineage-specific transcription factors PU.1 and GATA1, which challenges the previous model of early myeloid lineage choice.
- Philipp S. Hoppe
- , Michael Schwarzfischer
- & Timm Schroeder
-
Letter |
 Molecular logic behind the three-way stochastic choices that expand butterfly colour vision
Molecular logic behind the three-way stochastic choices that expand butterfly colour vision
Butterflies diversify their retinal mosaics by producing three stochastic types of ommatidia instead of the two types found in Drosophila; this study shows that butterfly retinas use two R7-like photoreceptors per ommatidium that each make an independent stochastic decision to express the transcription factor Spineless, which controls photoreceptor and ommatidial fate.
- Michael Perry
- , Michiyo Kinoshita
- & Claude Desplan
-
Outlook |
 Cell physiology: The changing colour of fat
Cell physiology: The changing colour of fat
The different functions of white, brown and beige fat might yield new targets in the fight against obesity and metabolic disease.
- Brian Owens

 Chromatin accessibility during human first-trimester neurodevelopment
Chromatin accessibility during human first-trimester neurodevelopment