Article
|
Open Access
Featured
-
-
Article
| Open Access Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM
Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM
Gene blaNDM, conferring resistance to carbapenem antibiotics, is globally distributed across Gram-negative bacteria on multiple plasmids. Here, Acman et al. study the dynamics underlying blaNDM dissemination across over 6000 bacterial genomes, and identify mobile genetic elements and specific mobilisation events likely involved in the gene’s global spread.
- Mislav Acman
- , Ruobing Wang
- & Francois Balloux
-
Article
| Open Access A role for ColV plasmids in the evolution of pathogenic Escherichia coli ST58
A role for ColV plasmids in the evolution of pathogenic Escherichia coli ST58
Escherichia coli ST58 has recently emerged as a globally disseminated extra-intestinal pathogen. Here, Reid et al. present a pan-genomic analysis of a global collection of ST58 isolates from animal and human sources, showing that ColV plasmid acquisition likely contributed to the divergence of a major sub-lineage that has a broad host range but is more commonly found in poultry and swine.
- Cameron J. Reid
- , Max L. Cummins
- & Steven P. Djordjevic
-
Article
| Open Access Population structure analysis and laboratory monitoring of Shigella by core-genome multilocus sequence typing
Population structure analysis and laboratory monitoring of Shigella by core-genome multilocus sequence typing
Lab-based surveillance of Shigella has traditionally been based on serotyping but increasing availability of whole genome sequencing could enable higher resolution typing. Here, the authors apply a core genome multilocus sequence typing scheme to Shigella sequence data and describe its population structure.
- Iman Yassine
- , Sophie Lefèvre
- & François-Xavier Weill
-
Article
| Open Access Population analysis of Legionella pneumophila reveals a basis for resistance to complement-mediated killing
Population analysis of Legionella pneumophila reveals a basis for resistance to complement-mediated killing
The bacterium Legionella pneumophila can cause severe respiratory infection, but is typically a symbiont of free-living amoeba. Here, the authors analyse the genomes of 902 clinical and environmental isolates, and identify a bacterial gene that is strongly associated with human infection and confers resistance to complement-mediated killing.
- Bryan A. Wee
- , Joana Alves
- & J. Ross Fitzgerald
-
Article
| Open Access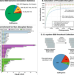 Generation of a Gluconobacter oxydans knockout collection for improved extraction of rare earth elements
Generation of a Gluconobacter oxydans knockout collection for improved extraction of rare earth elements
Bioleaching of rare earth elements using microorganisms offers an environmentally friendly alternative to thermochemical extraction. Here, Schmitz et al. generate a whole-genome knockout collection of mutants for one such microorganism, Gluconobacter oxydans, and identify genes affecting the production of acidic biolixiviant and thus bioleaching efficacy.
- Alexa M. Schmitz
- , Brooke Pian
- & Buz Barstow
-
Article
| Open Access Type VI secretion system mutations reduced competitive fitness of classical Vibrio cholerae biotype
Type VI secretion system mutations reduced competitive fitness of classical Vibrio cholerae biotype
The bacterium Vibrio cholerae has caused seven recorded cholera pandemics. The factors responsible for the decline of 6th pandemic classical biotype strains are not well understood. Here, Kostiuk et al. propose that classical strains underwent sequential mutations in type-six secretion system genes that disadvantaged them when confronted with 7th pandemic El Tor biotype strains.
- Benjamin Kostiuk
- , Francis J. Santoriello
- & Stefan Pukatzki
-
Article
| Open Access Population structure, biogeography and transmissibility of Mycobacterium tuberculosis
Population structure, biogeography and transmissibility of Mycobacterium tuberculosis
Mycobacterium tuberculosis is a clonal pathogen that has co-evolved with humans for millennia. Here, Freschi et al. reevaluate the population structure of M. tuberculosis, providing an in-depth analysis of the ancient Indo-Oceanic Lineage 1 and the modern Central Asian Lineage 3, and expanding our understanding of Lineages 2 and 4.
- Luca Freschi
- , Roger Vargas Jr.
- & Maha Reda Farhat
-
Article
| Open Access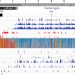 Dynamics of the compartmentalized Streptomyces chromosome during metabolic differentiation
Dynamics of the compartmentalized Streptomyces chromosome during metabolic differentiation
Streptomyces bacteria have a linear chromosome, with core genes located in the central region and gene clusters for specialized metabolite biosynthesis found in the ‘arms’. Here, Lioy et al. show that such chromosome structure correlates with genetic compartmentalization, and the onset of metabolic differentiation is accompanied by a rearrangement of chromosome architecture.
- Virginia S. Lioy
- , Jean-Noël Lorenzi
- & Stéphanie Bury-Moné
-
Article
| Open Access Global phylogenomic analyses of Mycobacterium abscessus provide context for non cystic fibrosis infections and the evolution of antibiotic resistance
Global phylogenomic analyses of Mycobacterium abscessus provide context for non cystic fibrosis infections and the evolution of antibiotic resistance
Mycobacterium abscessus is an emerging infection that usually affects patients with structural lung diseases such as cystic fibrosis (CF). Here, the authors use phylogenetic analyses to demonstrate close relationships between isolates from CF and non-CF patients and identify antibiotic resistance markers.
- Ryan A. Bronson
- , Chhavi Gupta
- & Keira A. Cohen
-
Article
| Open Access Conversion of dietary inositol into propionate and acetate by commensal Anaerostipes associates with host health
Conversion of dietary inositol into propionate and acetate by commensal Anaerostipes associates with host health
Here, the authors report an anaerobic metabolic pathway from the dominant gut butyrogen Anaerostipes, showing several strains of this genus to be capable of producing propionate from dietary myo-inositol that associates with reduced fasting-glucose levels in mice.
- Thi Phuong Nam Bui
- , Louise Mannerås-Holm
- & Willem M. deVos
-
Article
| Open Access A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
Klebsiella pneumoniae is a pathogen of increasing public health concern and antimicrobial resistance is becoming more prevalent. Here, the authors describe a K. pneumoniae genotyping tool, Kleborate, that can be used to identify lineages and detect antimicrobial resistance and virulence loci.
- Margaret M. C. Lam
- , Ryan R. Wick
- & Kathryn E. Holt
-
Article
| Open Access Genomic analyses of Mycobacterium tuberculosis from human lung resections reveal a high frequency of polyclonal infections
Genomic analyses of Mycobacterium tuberculosis from human lung resections reveal a high frequency of polyclonal infections
Polyclonal infections occur when at least two unrelated strains of the same pathogen are detected in an individual. Here, Moreno-Molina et al. analyse sputum and surgical resections from tuberculosis patients, showing that the magnitude of polyclonal infections can be underestimated when only testing sputum samples.
- Miguel Moreno-Molina
- , Natalia Shubladze
- & Iñaki Comas
-
Article
| Open Access Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii
Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii
Mycobacterium kansasii can cause serious pulmonary disease. Here, the authors present a population genomics analysis of 358 environmental and clinical isolates from around the world, supporting the idea that municipal water is a main source of infection, and shedding light into the pathogen’s diversity and adaptation to the human host.
- Tao Luo
- , Peng Xu
- & Qian Gao
-
Article
| Open Access Reductive evolution and unique predatory mode in the CPR bacterium Vampirococcus lugosii
Reductive evolution and unique predatory mode in the CPR bacterium Vampirococcus lugosii
The Candidate Phyla Radiation (CPR) constitutes a large group of bacterial lineages with small cell sizes and limited biosynthetic capabilities. Here, Moreira et al. study the biology and genome of Vampirococcus lugosii, an epibiotic parasite of other bacteria, supporting parasitism as a common lifestyle of CPR bacteria.
- David Moreira
- , Yvan Zivanovic
- & Purificación López-García
-
Article
| Open Access Forecasting the dissemination of antibiotic resistance genes across bacterial genomes
Forecasting the dissemination of antibiotic resistance genes across bacterial genomes
Antibiotic resistance spreads among bacteria through horizontal transfer of antibiotic resistance genes (ARGs). Here, Ellabaan et al. use a statistical approach to identify putative mobilisation elements and other features associated with ARG transfer among bacterial clades to predict the potential future dissemination of known ARGs.
- Mostafa M. H. Ellabaan
- , Christian Munck
- & Morten O. A. Sommer
-
Article
| Open Access High-throughput fitness screening and transcriptomics identify a role for a type IV secretion system in the pathogenesis of Crohn’s disease-associated Escherichia coli
High-throughput fitness screening and transcriptomics identify a role for a type IV secretion system in the pathogenesis of Crohn’s disease-associated Escherichia coli
Adherent-invasive E. coli (AIEC) are frequently isolated from Crohn’s disease (CD) patients. Here, Elhenawy et al. conduct a genome-wide screen to identify AIEC genes required for in vivo intestinal colonization, and show that a type IV secretion system contributes to AIEC persistence in the gut and is enriched in CD patients’ isolates.
- Wael Elhenawy
- , Sarah Hordienko
- & Brian K. Coombes
-
Article
| Open Access Apparent nosocomial adaptation of Enterococcus faecalis predates the modern hospital era
Apparent nosocomial adaptation of Enterococcus faecalis predates the modern hospital era
Enterococcus faecalis is a commensal microorganism of animals, insects and humans, but also a nosocomial pathogen. Here, the authors analyse genomic sequences from E. faecalis isolates from animals and humans, and find that the last common ancestors of multiple hospital-associated lineages date to the pre-antibiotic era.
- Anna K. Pöntinen
- , Janetta Top
- & Jukka Corander
-
Article
| Open Access Spatiotemporal persistence of multiple, diverse clades and toxins of Corynebacterium diphtheriae
Spatiotemporal persistence of multiple, diverse clades and toxins of Corynebacterium diphtheriae
Cases of diphtheria have increased in recent years. Here, the authors analyse the genomes of 502 Corynebacterium diphtheriae isolates across 16 countries and territories over 122 years, describing an increase in antimicrobial resistance genes and identifying toxin variants.
- Robert C. Will
- , Thandavarayan Ramamurthy
- & Ankur Mutreja
-
Article
| Open Access Hologenome analysis reveals dual symbiosis in the deep-sea hydrothermal vent snail Gigantopelta aegis
Hologenome analysis reveals dual symbiosis in the deep-sea hydrothermal vent snail Gigantopelta aegis
Symbiotic partners are rarely studied in equal depth. By assembling new genomes, Lan et al. report a novel dual symbiosis in the snail Gigantopelta aegis with two evolutionarily distant gammaproteobacterial endosymbionts: one which oxidises sulfur, the other, methane in a metabolically mutualistic relationship.
- Yi Lan
- , Jin Sun
- & Pei-Yuan Qian
-
Article
| Open Access Insertion-sequence-mediated mutations both promote and constrain evolvability during a long-term experiment with bacteria
Insertion-sequence-mediated mutations both promote and constrain evolvability during a long-term experiment with bacteria
Insertion sequences (IS) are common mobile genetic elements in bacteria, but their effects on bacterial evolution are not well understood. Here, Consuegra and colleagues investigate the dynamics and fitness consequences of IS elements in E. coli over 50,000 generations.
- Jessika Consuegra
- , Joël Gaffé
- & Dominique Schneider
-
Article
| Open Access A collection of bacterial isolates from the pig intestine reveals functional and taxonomic diversity
A collection of bacterial isolates from the pig intestine reveals functional and taxonomic diversity
The authors present a public collection of 117 bacterial isolates from the pig gut, including the description of 38 novel taxa. Interesting functions discovered in these organisms include a new fucosyltransferease and sactipeptide-like molecules encoded by biosynthetic gene clusters.
- David Wylensek
- , Thomas C. A. Hitch
- & Thomas Clavel
-
Article
| Open Access Construction of a complete set of Neisseria meningitidis mutants and its use for the phenotypic profiling of this human pathogen
Construction of a complete set of Neisseria meningitidis mutants and its use for the phenotypic profiling of this human pathogen
The bacterium Neisseria meningitidis causes life-threatening meningitis and sepsis. Here, Muir et al. construct a complete collection of defined mutants in protein-coding genes of this organism, which they use to identify its essential genome and to shed light on the functions of multiple genes.
- Alastair Muir
- , Ishwori Gurung
- & Vladimir Pelicic
-
Article
| Open Access Increased power from conditional bacterial genome-wide association identifies macrolide resistance mutations in Neisseria gonorrhoeae
Increased power from conditional bacterial genome-wide association identifies macrolide resistance mutations in Neisseria gonorrhoeae
The mechanisms underlying resistance of Neisseria gonorrhoeae to the antibiotic azithromycin are incompletely understood. Here, Ma et al. conduct a conditional genome-wide association study to identify new resistance mutations and experimentally confirm that a mutation in ribosomal protein L4 confers increased resistance.
- Kevin C. Ma
- , Tatum D. Mortimer
- & Yonatan H. Grad
-
Article
| Open Access Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates
Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates
Some bacterial lineages appear to be pre-disposed to evolving antibiotic resistance. Here, the authors show that differential expression of an efflux pump causes widespread variation in evolvability across Staphylococcus aureus isolates, and chemical inhibition of the pump prevents resistance evolution.
- Andrei Papkou
- , Jessica Hedge
- & R. Craig MacLean
-
Article
| Open Access A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron
A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron
Bacteroides thetaiotaomicron is a human gut microbe and an emergent model organism. Here, Ryan et al. generate single-nucleotide resolution RNA-seq data for this bacterium and map transcription start sites and noncoding RNAs, one of which modulates expression of metabolic enzymes.
- Daniel Ryan
- , Laura Jenniches
- & Alexander J. Westermann
-
Article
| Open Access A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region
A sister lineage of the Mycobacterium tuberculosis complex discovered in the African Great Lakes region
The human- and animal-adapted lineages of the Mycobacterium tuberculosis complex (MTBC) are thought to be evolved from a common progenitor in Africa. Here, the authors identify two MTBC strains isolated from patients with multidrug-resistant tuberculosis, representing an as-yet-unknown lineage further supporting an East African origin for the MTBC.
- Jean Claude Semuto Ngabonziza
- , Chloé Loiseau
- & Philip Supply
-
Article
| Open Access Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines
Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines
Whole-genome sequencing (WGS) can support drug resistance surveillance, but is rare in low- and middle-income countries. Here, the authors integrate WGS within the national antimicrobial resistance surveillance program in the Philippines and conduct a retrospective sequencing survey to characterize bacterial populations and dissect resistance phenotypes.
- Silvia Argimón
- , Melissa A. L. Masim
- & Celia C. Carlos
-
Article
| Open Access Large-scale network analysis captures biological features of bacterial plasmids
Large-scale network analysis captures biological features of bacterial plasmids
Plasmids can mediate the exchange of genetic material between bacterial cells. Here, Acman et al. use network analyses to study the population structure and dynamics of over 10,000 plasmids, assigning them into cliques that correlate with gene content, host range, and existing classifications based on replicon and mobility types.
- Mislav Acman
- , Lucy van Dorp
- & Francois Balloux
-
Article
| Open Access CdbA is a DNA-binding protein and c-di-GMP receptor important for nucleoid organization and segregation in Myxococcus xanthus
CdbA is a DNA-binding protein and c-di-GMP receptor important for nucleoid organization and segregation in Myxococcus xanthus
The second messenger c-di-GMP modulates multiple responses to environmental and cellular signals in bacteria. Here, Skotnicka et al. identify a protein that binds c-di-GMP and contributes to chromosome organization and segregation in Myxococcus xanthus, with DNA-binding activity regulated by c-di-GMP.
- Dorota Skotnicka
- , Wieland Steinchen
- & Lotte Søgaard-Andersen
-
Article
| Open Access A megaplasmid family driving dissemination of multidrug resistance in Pseudomonas
A megaplasmid family driving dissemination of multidrug resistance in Pseudomonas
The emergence of multidrug resistance (MDR) in bacteria represents a global threat to human health. Here, Cazares et al. identify a family of MDR megaplasmids carrying large arrays of antibiotic resistance genes in Pseudomonas strains from various sources, including P. aeruginosa clinical isolates.
- Adrian Cazares
- , Matthew P. Moore
- & Craig Winstanley
-
Article
| Open Access A comparative genomics methodology reveals a widespread family of membrane-disrupting T6SS effectors
A comparative genomics methodology reveals a widespread family of membrane-disrupting T6SS effectors
Gram-negative bacteria deliver effectors via the type VI secretion system (T6SS) to outcompete their rivals. Here, Fridman et al. present an approach to identify T6SS effectors encoded in bacterial genomes without prior knowledge of their domain content or genetic neighbourhood, and identify a new family of membrane-disrupting effectors.
- Chaya M. Fridman
- , Kinga Keppel
- & Dor Salomon
-
Article
| Open Access Integrating multiple genomic technologies to investigate an outbreak of carbapenemase-producing Enterobacter hormaechei
Integrating multiple genomic technologies to investigate an outbreak of carbapenemase-producing Enterobacter hormaechei
Antibiotic-resistant bacteria are an urgent threat to human health. Here, Roberts et al. characterise and monitor an ongoing hospital outbreak of carbapenemase-producing Enterobacter hormaechei by integrating several technologies for whole-genome sequencing and shotgun metagenomics.
- Leah W. Roberts
- , Patrick N. A. Harris
- & Scott A. Beatson
-
Article
| Open Access The C. difficile toxin B membrane translocation machinery is an evolutionarily conserved protein delivery apparatus
The C. difficile toxin B membrane translocation machinery is an evolutionarily conserved protein delivery apparatus
Large Clostridial toxins infiltrate host cells using a translocation domain (LCT-T). Here, using a genomics-driven approach and functional assays, the authors uncover the presence of distant LCT-T homologs in bacteria outside clostridia and provide evidence for a toxic effector function in the gammaproteobacterium Serratia marcescens.
- Kathleen E. Orrell
- , Michael J. Mansfield
- & Roman A. Melnyk
-
Article
| Open Access Phagocytosis-like cell engulfment by a planctomycete bacterium
Phagocytosis-like cell engulfment by a planctomycete bacterium
Phagocytosis is a typically eukaryotic feature that could be behind the origin of eukaryotic cells. Here, the authors describe a bacterium that can engulf other bacteria and small eukaryotic cells through a phagocytosis-like mechanism.
- Takashi Shiratori
- , Shigekatsu Suzuki
- & Ken-ichiro Ishida
-
Article
| Open Access A hybrid sub-lineage of Listeria monocytogenes comprising hypervirulent isolates
A hybrid sub-lineage of Listeria monocytogenes comprising hypervirulent isolates
Listeria monocytogenes isolates are highly heterogeneous and exhibit different levels of virulence. Here, the authors identify hypervirulent isolates that represent a hybrid sub-lineage of the major lineage II harbouring virulence factors from Listeria ivanovii and wall teichoic acids found in major lineage I.
- Yuelan Yin
- , Hao Yao
- & Xin’an Jiao
-
Article
| Open Access Comparative fitness analysis of D-cycloserine resistant mutants reveals both fitness-neutral and high-fitness cost genotypes
Comparative fitness analysis of D-cycloserine resistant mutants reveals both fitness-neutral and high-fitness cost genotypes
D-cycloserine (DCS) has been used for decades to treat Mycobacterium tuberculosis (Mtb) but resistance is rarely observed in clinical isolates. Here, the authors report ultra-low rate of emergence of resistance mutations as the underlying mechanism of DCS resistance evasion in Mtb.
- Dimitrios Evangelopoulos
- , Gareth A. Prosser
- & Luiz Pedro S. de Carvalho
-
Article
| Open Access The distinction of CPR bacteria from other bacteria based on protein family content
The distinction of CPR bacteria from other bacteria based on protein family content
Recent studies have identified a large, phylogenetically distinct clade of bacteria, the candidate phyla radiation (CPR). Here, Méheust and colleagues analyze almost 3600 genomes to characterize the protein family content of CPR versus other bacteria and archaea.
- Raphaël Méheust
- , David Burstein
- & Jillian F. Banfield
-
Article
| Open Access Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
5-carboxymethoxyuridine (cmo5U) is one of the RNA modifications found in bacterial tRNA anticodons. Here the authors show that the first step of cmo5U biosynthesis from uridine is mediated by either one of two parallel factors, TrhP or TrhO, and that cmo5U modification is required for efficient translation.
- Yusuke Sakai
- , Satoshi Kimura
- & Tsutomu Suzuki
-
Article
| Open Access Bacteroidetes use thousands of enzyme combinations to break down glycans
Bacteroidetes use thousands of enzyme combinations to break down glycans
Bacteroidetes genomes contain polysaccharide utilization loci (PULs), each of which encodes enzymes for the breakdown of one particular glycan. By analyzing the enzyme composition of 13,537 PULs, the authors suggest that the natural glycan diversity is orders of magnitude lower than previously proposed.
- Pascal Lapébie
- , Vincent Lombard
- & Bernard Henrissat
-
Article
| Open Access The phylogeography and incidence of multi-drug resistant typhoid fever in sub-Saharan Africa
The phylogeography and incidence of multi-drug resistant typhoid fever in sub-Saharan Africa
Typhoid fever is caused by the bacterium Salmonella Typhi. Here, Park et al. analyse the genomes of 249 S. Typhi isolates from 11 sub-Saharan African countries, identifying genes and plasmids associated with antibiotic resistance and showing that multi-drug resistance is highly pervasive in sub-Saharan Africa.
- Se Eun Park
- , Duy Thanh Pham
- & Stephen Baker
-
Article
| Open Access Microevolution of Neisseria lactamica during nasopharyngeal colonisation induced by controlled human infection
Microevolution of Neisseria lactamica during nasopharyngeal colonisation induced by controlled human infection
Carriage of Neisseria lactamica, a harmless coloniser of the human respiratory tract, is inversely correlated with Neisseria meningitidis infection. Here, Pandey et al. provide insights into micro-evolutionary processes in N. lactamica during controlled infection of healthy volunteers.
- Anish Pandey
- , David W. Cleary
- & Robert C. Read
-
Article
| Open Access Synthetic microbe communities provide internal reference standards for metagenome sequencing and analysis
Synthetic microbe communities provide internal reference standards for metagenome sequencing and analysis
Complex microbial communities pose a challenge to metagenomic analysis. Here the authors develop ‘sequins’, internal DNA standards that represent a synthetic community of artificial genomes.
- Simon A. Hardwick
- , Wendy Y. Chen
- & Tim R. Mercer
-
Article
| Open Access Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination
Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination
Since the 1980s, hypervirulent clonal-group CG23 serotype K1 Klebsiella pneumoniae has been recognised as a prominent cause of community-acquired liver abscess and other severe infections. Here, the authors investigate the genomic evolutionary history of CG23 and suggest a new reference strain for CG23.
- Margaret M. C. Lam
- , Kelly L. Wyres
- & Kathryn E. Holt
-
Article
| Open Access Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen
Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen
Microbes in the cow rumen are crucial for the breakdown of plant material. Here, Stewart et al. assemble over 900 bacterial and archaeal genomes from the cow rumen microbiome, revealing new species and genes encoding enzymes with potential roles in carbohydrate metabolism.
- Robert D. Stewart
- , Marc D. Auffret
- & Mick Watson
-
Article
| Open Access Phylogenomics and antimicrobial resistance of the leprosy bacillus Mycobacterium leprae
Phylogenomics and antimicrobial resistance of the leprosy bacillus Mycobacterium leprae
Leprosy is caused by the yet-uncultured pathogen Mycobacterium leprae. Here, Benjak et al. obtain M. leprae genome sequences from DNA extracted from patients' skin biopsies and, by analysing 154 genomes from 25 countries, provide insight into the pathogen’s evolution and antimicrobial resistance.
- Andrej Benjak
- , Charlotte Avanzi
- & Stewart T. Cole
-
Article
| Open Access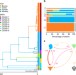 Distinct Campylobacter fetus lineages adapted as livestock pathogens and human pathobionts in the intestinal microbiota
Distinct Campylobacter fetus lineages adapted as livestock pathogens and human pathobionts in the intestinal microbiota
Human infections with Campylobacter fetus are often assumed to be derived from livestock. Here, Iraola et al. provide evidence that healthy humans may act as carriers and dispersers, and C. fetus may have originated in humans as an intestinal pathobiont and then adapted as a livestock pathogen.
- Gregorio Iraola
- , Samuel C. Forster
- & Trevor D. Lawley
-
Article
| Open Access The chromosomal organization of horizontal gene transfer in bacteria
The chromosomal organization of horizontal gene transfer in bacteria
Horizontal gene transfer (HGT) is an important mechanism for genome evolution and adaptation in bacteria. Here, Oliveira and colleagues find HGT hotspots comprising ~ 1% of the chromosomal regions in 80 bacterial species.
- Pedro H. Oliveira
- , Marie Touchon
- & Eduardo P. C. Rocha
-
Article
| Open Access Community-like genome in single cells of the sulfur bacterium Achromatium oxaliferum
Community-like genome in single cells of the sulfur bacterium Achromatium oxaliferum
The cells of Achromatium bacteria are remarkably large and contain multiple chromosome copies. Here, Ionescu et al. show that chromosome copies within individual cells display high diversity, similar to that of bacterial communities, and contain tens of transposable elements.
- Danny Ionescu
- , Mina Bizic-Ionescu
- & Hans-Peter Grossart
-
Article
| Open Access Nanoscopy of bacterial cells immobilized by holographic optical tweezers
Nanoscopy of bacterial cells immobilized by holographic optical tweezers
Nanoscopy of non-adherent cells is currently not possible, due to their movement in solution. Here the authors immobilize and manipulate fixedE. coli by multiple optical traps; their holographic optical tweezers enable dSTORM imaging of orthogonal planes via 3D realignment of the sample.
- Robin Diekmann
- , Deanna L. Wolfson
- & Thomas Huser

 The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages
The core and accessory Hfq interactomes across Pseudomonas aeruginosa lineages