« Prev Next »
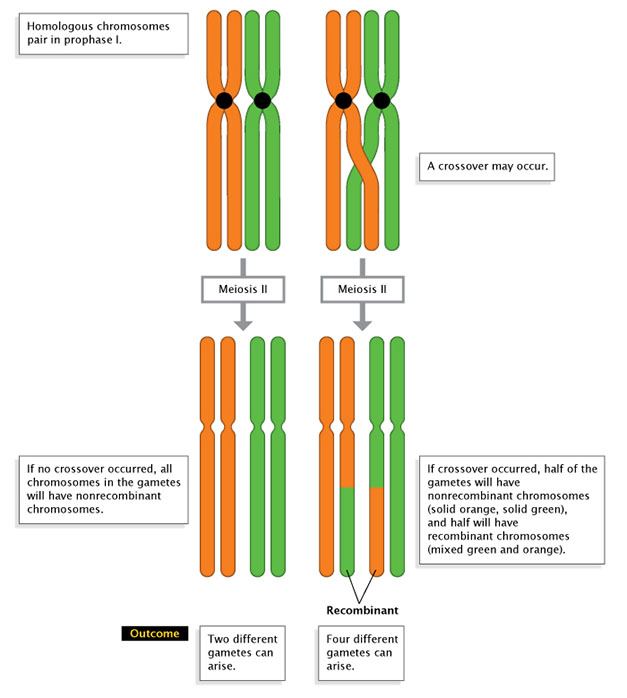
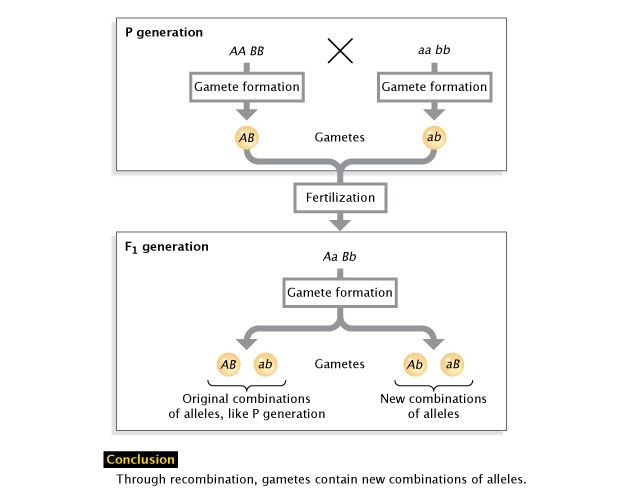
In 1911, while studying the chromosome theory of heredity, biologist Thomas Hunt Morgan had a major breakthrough. Morgan occasionally noticed that "linked" traits would separate. Meanwhile, other traits on the same chromosome showed little detectable linkage. Morgan considered the evidence and proposed that a process of crossing over, or recombination, might explain his results. Specifically, he proposed that the two paired chromosomes could "cross over" to exchange information. Today, we know that recombination does indeed occur during prophase of meiosis (Figure 1), and it creates different combinations of alleles in the gametes that result (i.e., the F1 generation; Figure 2).
When proposing the idea of crossing over, Morgan also hypothesized that the frequency of recombination was related to the distance between the genes on a chromosome, and that the interchange of genetic information broke the linkage between genes. Morgan imagined that genes on chromosomes were similar to pearls on a string (Weiner, 1999); in other words, they were physical objects. The closer two genes were to one another on a chromosome, the greater their chance of being inherited together. In contrast, genes located farther away from one another on the same chromosome were more likely to be separated during recombination. Therefore, Morgan correctly proposed that the strength of linkage between two genes depends upon the distance between the genes on the chromosome. This proposition became the basis for construction of the earliest maps of the human genome.
Sturtevant Uses Crossing-Over Data to Construct the First Genetic Map
Soon after Morgan presented his hypothesis, Alfred Henry Sturtevant, a 19-year-old Columbia University undergraduate who was working with Morgan, realized that if the frequency of crossing over was related to distance, one could use this information to map out the genes on a chromosome. After all, the farther apart two genes were on a chromosome, the more likely it was that these genes would separate during recombination. Therefore, as Sturtevant explained it, the "proportion of crossovers could be used as an index of the distance between any two factors" (Sturtevant, 1913). Collecting a stack of laboratory data, Sturtevant went home and spent most of the night drawing the first chromosomal linkage map for the genes located on the X chromosome of fruit flies (Weiner, 1999).
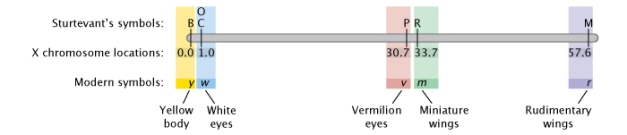
When creating his map, Sturtevant started by placing six X-linked genes in order. B was a gene for black body color. C was a gene that allowed color to appear in the eyes. Flies with the P gene had vermilion eyes instead of the ordinary red, and flies with two copies of the recessive O gene had eyes that appeared a shade known as eosin. The R and M factors both affected the wings. Sturtevant placed C and O at the same point because they were completely linked and were always inherited together — in other words, he never saw any evidence for recombination between C and O. Sturtevant then placed the remainder of the genes in the order shown in Figure 3 (Sturtevant, 1913). Crossover events were tracked by examining the F2 progeny in crosses for "new" phenotypes.

For example, to find the distance between P (vermilion eyes) and M (long wings), Sturtevant performed crosses between flies that had long wings and vermilion eyes and flies that had small wings and red eyes. These crosses resulted in F1 flies that either had long wings and red eyes or long wings and vermilion eyes. Sturtevant then crossed these two types of F1 flies and analyzed the offspring for evidence of recombination. Unexpected phenotypes observed in the male F2 progeny from this cross were then examined. (Because very little recombination occurs in the male germ line of Drosophila, only the female F1 chromosomes are considered for predicting phenotypes [Figure 4].) Sturtevant noted four classes of male flies in this F2 generation, as shown in Table 1.
The two additional classes of flies that appeared in this generation (long wings with red eyes and rudimentary wings with vermilion eyes) could only be explained by recombination occurring in the female germ line.
| Phenotype | Number of Flies | Nature of Related Gametes |
| Long wings, red eyes | 105 | Recombinant |
| Rudimentary wings, red eyes | 33 | Nonrecombinant |
| Long wings, vermilion eyes | 316 | Nonrecombinant |
| Rudimentary wings, vermilion eyes | 4 | Recombinant |
| Table 1: Class of male files in the F2 generation | ||
Sturtevant then worked out the order and the linear distances between these linked genes, thus forming a linkage map. In doing so, he computed the distance in an arbitrary unit he called the "map unit," which represented a recombination frequency of 0.01, or 1%. Later, the map unit was renamed the centimorgan (cM), in honor of Thomas Hunt Morgan, and it is still used today as the unit of measurement of distances along chromosomes.
In addition to describing the order of the genes on the X chromosome of fruit flies, Sturtevant's 1913 paper elucidated a number of other interesting points, including the following:
- The relationship between crossing over and genetic map distance
- The effects of multiple crossover events
- The fact that a first crossover can inhibit a second crossover (a phenomenon called interference, which is described later in this article)
Mapping Genes Using Recombination Frequency
To better understand how Sturtevant arrived as his results, let's take a closer look at the process he followed. In Figure 5, the gray-eosin and yellow-red flies are the parental lines, and all the alleles for these traits are linked on the X chromosome. Therefore, any gray-red or yellow-eosin male offspring are recombinants. As you can see, two recombinants result from the cross. We count only the male progeny because the males have one X chromosome and dominance will not obscure any phenotypes (Robbins, 2000). Of course, crossing over can occur only in the female fruit flies, which have two X chromosomes. Thus, in this cross, the female F1 gametes provide the parental and recombinant gametes that we observe in the F2 progeny.
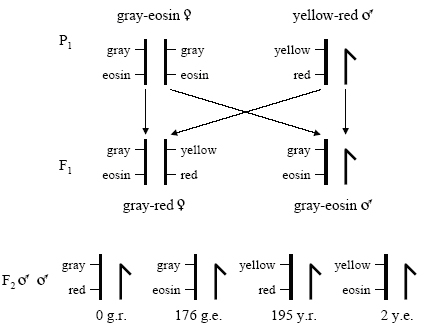
Therefore, the two genes are 0.5 map units apart.
Deviations from Expected Results Revealed Genetic Interference
In short, Sturtevant realized that double recombination events could occur if genes were far apart. Moreover, not only did Sturtevant's data suggest that double-crossing over occurred, but it also suggested that an initial crossover event could inhibit subsequent events by way of a phenomenon Sturtevant referred to as interference.
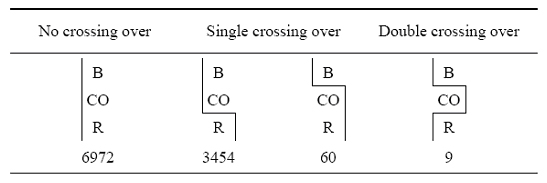
To understand how Sturtevant arrived at this conclusion, take a look at the data shown in Figure 6 (Sturtevant, 1913). As you can see, Sturtevant examined recombination events between B (body color), CO (two eye color genes that were closely linked), and R (rudimentary wings), and compared the frequencies of crossover events. When B and CO did not separate, Sturtevant noticed that the "gametic ratio," or presence, of CO/R recombinants was approximately 1:2 (3,454:6,972). However, when a crossover between B/CO (N = 60) occurred, there was a much lower likelihood (approximately 1:6.5) of a crossover between CO/R (N = 9). This finding is indicative of interference.
Interference phenomena are still being studied today, and research has shown that interference can act over extremely large distances of the genome. For example, Kenneth J. Hillers and Anne M. Villeneuve recently demonstrated that in Caenorhabditis elegans, interference can actually occur over half the genome of the organism. They demonstrated this by fusing multiple chromosomes together and observing that crossovers still occurred a single time (Hillers & Villeneuve, 2003).
Complete and Incomplete Linkage
When Sturtevant drew his chromosomal map, he placed the C and O genes at the same location because they were always inherited together (Figure 3; Sturtevant, 1913). Genes that are so close together on a chromosome that they are always inherited as a single unit show a relationship referred to as complete linkage. In fact, two genes that are completely linked can only be differentiated as separate genes when a mutation occurs in one of them. There is no other way to identify genes with complete linkage from single genes that show multiple phenotypes.
On the other hand, the phenomenon known as incomplete linkage occurs when two genes show linkage with a recombination level greater than 0% and less than 50%. In incomplete linkage, all expected types of gametes are formed, but the recombinant gametes occur less often than the parental gametes.
In addition, if two genes are on the same chromosome and are far enough apart that they undergo recombination at least 50% of the time, the genes are independently assorting and do not show linkage. Genes independently assort at a distance of 50 cM or more apart. This means that no statistical test would allow researchers to measure linkage.
Finally, linked genes that do not independently assort show statistical linkage. Statistical linkage is detected as deviation from independent assortment that favors the parental gametes. Syntenic genes are genes that are physically located on the same chromosome, whether or not the genes themselves exhibit linkage (Passarge et al., 1999). Therefore, all linked genes are syntenic, but not all syntenic genes show genetic linkage.
References and Recommended Reading
Blixt, S. Why didn't Gregor Mendel find linkage? Nature 256, 206 (1975) doi:10.1038/256206a0 (link to article)
Bridges, C. B. Salivary chromosome maps with a key to the banding of the chromosomes of Drosophila melanogaster. Journal of Heredity 26, 60–64 (1935)
———. A revised map of the salivary gland X chromosome. Journal of Heredity 29, 1113 (1938)
Hillers, K., & Villeneuve, A. Chromosome-wide control of meiotic crossing over in C. elegans. Current Biology 13, 1641–1647 (2003) doi:10.1016/j.cub.2003.08.026
Morgan, T. H. Random segregation versus coupling in Mendelian inheritance. Science 34, 384 (1911)
Passarge, E., et al. Incorrect use of the term "synteny." Nature Genetics 23, 387 (1999) (link to article)
Pierce, B. Genetics: A Conceptual Approach. (New York, W. H. Freeman & Co., 2005)
Punnett, R. C. Linkage in the sweet pea (Lathyrus odoratus). Journal of Genetics 13, 101–123 (1923)
———. Linkage groups and chromosome number in Lathyrus. Proceedings of the Royal Society of London: Series B, Containing Papers of a Biological Character 102 236–238. (1927)
Robbins, R. J. Introduction to sex-limited inheritance in Drosophila. Electronic Scholarly Publishing Foundations of Classical Genetics Project. http://www.esp.org/foundations/genetics/classical/thm-10a.pdf (2000) (accessed May 19, 2008)
Sturtevant, A. H. The linear arrangement of six sex-linked factors in Drosophila, as shown by their mode of association. Journal of Experimental Zoology 14, 43–59 (1913)
Weiner, J. Time, Love, Memory: A Great Biologist and His Quest for the Origins of Behavior (New York, Random House, 1999)
































