« Prev Next »
The evolutionary history of viruses represents a fascinating, albeit murky, topic for virologists and cell biologists. Because of the great diversity among viruses, biologists have struggled with how to classify these entities and how to relate them to the conventional tree of life. They may represent genetic elements that gained the ability to move between cells. They may represent previously free-living organisms that became parasites. They may be the precursors of life as we know it.
The Basics of Viruses
We know that viruses are quite diverse. Unlike all other biological entities, some viruses, like poliovirus, have RNA genomes and some, like herpesvirus, have DNA genomes. Further, some viruses (like influenza virus) have single-stranded genomes, while others (like smallpox) have double-stranded genomes. Their structures and replication strategies are equally diverse. Viruses, do, however, share a few features: First, they generally are quite small, with a diameter of less than 200 nanometers (nm). Second, they can replicate only within a host cell. Third, no known virus contains ribosomes, a necessary component of a cell's protein-making translational machinery.
Are Viruses Alive?
Do viruses conform to these criteria? Yes and no. We probably all realize that viruses reproduce in some way. We can become infected with a small number of virus particles — by inhaling particles expelled when another person coughs, for instance — and then become sick several days later as the viruses replicate within our bodies. Likewise we probably all realize that viruses evolve over time. We need to get a flu vaccine every year primarily because the influenza virus changes, or evolves, from one year to the next (Nelson & Holmes 2007).
Viruses do not, however, carry out metabolic processes. Most notably, viruses differ from living organisms in that they cannot generate ATP. Viruses also do not possess the necessary machinery for translation, as mentioned above. They do not possess ribosomes and cannot independently form proteins from molecules of messenger RNA. Because of these limitations, viruses can replicate only within a living host cell. Therefore, viruses are obligate intracellular parasites. According to a stringent definition of life, they are nonliving. Not everyone, though, necessarily agrees with this conclusion. Perhaps viruses represent a different type of organism on the tree of life — the capsid-encoding organisms, or CEOs (Figure 1; Raoult & Forterre 2008).
Where Did Viruses Come From?
There is much debate among virologists about this question. Three main hypotheses have been articulated: 1. The progressive, or escape, hypothesis states that viruses arose from genetic elements that gained the ability to move between cells; 2. the regressive, or reduction, hypothesis asserts that viruses are remnants of cellular organisms; and 3. the virus-first hypothesis states that viruses predate or coevolved with their current cellular hosts.
The Progressive Hypothesis
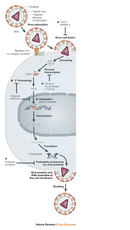
Retroviruses have a single-stranded RNA genome. When the virus enters a host cell, a viral enzyme, reverse transcriptase, converts that single-stranded RNA into double-stranded DNA. This viral DNA then migrates to the nucleus of the host cell. Another viral enzyme, integrase, inserts the newly formed viral DNA into the host cell's genome. Viral genes can then be transcribed and translated. The host cell's RNA polymerase can produce new copies of the virus's single-stranded RNA genome. Progeny viruses assemble and exit the cell to begin the process again (Figure 2).
This process very closely mirrors the movement of an
important, though somewhat unusual, component of most eukaryotic genomes: retrotransposons.
These mobile genetic elements make up an astonishing 42% of the human genome
(Lander et al. 2001) and can move
within the genome via an RNA intermediate. Like retroviruses, certain classes
of retrotransposons, the viral-like retrotransposons, encode a reverse
transcriptase and, often, an integrase. With these enzymes, these elements can
be transcribed into RNA, reverse-transcribed into DNA, and then integrated into
a new location within the genome (Figure 3). We can speculate that the
acquisition of a few structural proteins could allow the element to exit a cell
and enter a new cell, thereby becoming an infectious agent. Indeed, the genetic
structures of retroviruses and viral-like retrotransposons show remarkable
similarities.
The Regressive Hypothesis
In contrast to the progressive process just described, viruses may have originated via a regressive, or reductive, process. Microbiologists generally agree that certain bacteria that are obligate intracellular parasites, like Chlamydia and Rickettsia species, evolved from free-living ancestors. Indeed, genomic studies indicate that the mitochondria of eukaryotic cells and Rickettsia prowazekii may share a common, free-living ancestor (Andersson et al. 1998). It follows, then, that existing viruses may have evolved from more complex, possibly free-living organisms that lost genetic information over time, as they adopted a parasitic approach to replication.
Viruses of one particular group, the nucleocytoplasmic large DNA viruses (NCLDVs), best illustrate this hypothesis. These viruses, which include smallpox virus and the recently discovered giant of all viruses, Mimivirus, are much bigger than most viruses (La Scola et al. 2003). A typical brick-shaped poxvirus, for instance, may be 200 nm wide and 300 nm long. About twice that size, Mimivirus exhibits a total diameter of roughly 750 nm (Xiao et al. 2005). Conversely, spherically shaped influenza virus particles may be only 80 nm in diameter, and poliovirus particles have a diameter of only 30 nm, roughly 10,000 times smaller than a grain of salt. The NCLDVs also possess large genomes. Again, poxvirus genomes often approach 200,000 base pairs, and Mimivirus has a genome of 1.2 million base pairs; while poliovirus has a genome of only 7,500 nucleotides total. In addition to their large size, the NCLDVs exhibit greater complexity than other viruses have and depend less on their host for replication than do other viruses. Poxvirus particles, for instance, include a large number of viral enzymes and related factors that allow the virus to produce functional messenger RNA within the host cell cytoplasm.
Because of the size and complexity of NCLDVs, some virologists have hypothesized that these viruses may be descendants of more complex ancestors. According to proponents of this hypothesis, autonomous organisms initially developed a symbiotic relationship. Over time, the relationship turned parasitic, as one organism became more and more dependent on the other. As the once free-living parasite became more dependent on the host, it lost previously essential genes. Eventually it was unable to replicate independently, becoming an obligate intracellular parasite, a virus. Analysis of the giant Mimivirus may support this hypothesis. This virus contains a relatively large repertoire of putative genes associated with translation — genes that may be remnants of a previously complete translation system. Interestingly, Mimivirus does not differ appreciably from parasitic bacteria, such as Rickettsia prowazekii (Raoult et al. 2004).
The Virus-First Hypothesis
Most biologists now agree that the very first replicating molecules consisted of RNA, not DNA. We also know that some RNA molecules, ribozymes, exhibit enzymatic properties; they can catalyze chemical reactions. Perhaps, simple replicating RNA molecules, existing before the first cell formed, developed the ability to infect the first cells. Could today's single-stranded RNA viruses be descendants of these precellular RNA molecules?
Others have argued that precursors of today's NCLDVs led to the emergence of eukaryotic cells. Villarreal and DeFilippis (2000) and Bell (2001) described models explaining this proposal. Perhaps, both groups postulate, the current nucleus in eukaryotic cells arose from an endosymbiotic-like event in which a complex, enveloped DNA virus became a permanent resident of an emerging eukaryotic cell.
No Single Hypothesis May Be Correct
Where viruses came from is not a simple question to answer. One can argue quite convincingly that certain viruses, such as the retroviruses, arose through a progressive process. Mobile genetic elements gained the ability to travel between cells, becoming infectious agents. One can also argue that large DNA viruses arose through a regressive process whereby once-independent entities lost key genes over time and adopted a parasitic replication strategy. Finally, the idea that viruses gave rise to life as we know it presents very intriguing possibilities. Perhaps today's viruses arose multiple times, via multiple mechanisms. Perhaps all viruses arose via a mechanism yet to be uncovered. Today's basic research in fields like microbiology, genomics, and structural biology may provide us with answers to this basic question.
Summary
Contemplating the origins of life fascinates both scientists and the general public. Understanding the evolutionary history of viruses may shed some light on this interesting topic. To date, no clear explanation for the origin(s) of viruses exists. Viruses may have arisen from mobile genetic elements that gained the ability to move between cells. They may be descendants of previously free-living organisms that adapted a parasitic replication strategy. Perhaps viruses existed before, and led to the evolution of, cellular life. Continuing studies may provide us with clearer answers. Or future studies may reveal that the answer is even murkier than it now appears.
References and Recommended Reading
Andersson, S. G. E. et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396, 133–143 (1998) doi:10.1038/24094.
Bell, P. J. L. Viral eukaryogenesis: Was the ancestor of the nucleus a complex DNA virus? Journal of Molecular Evolution 53, 251–256 (2001) doi:10.1007/s002390010215.
Koonin, E. V. & Martin, W. On the origin of genomes and cells within inorganic compartments. Trends in Genetics 21, 647–654 (2005).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001) doi:10.1038/35057062.
La Scola, B. et al. A giant virus in Amoebae. Science 299, 2033 (2003) doi:10.1126/science.1081867.
Nelson, M. I. & Holmes, E. C. The evolution of epidemic influenza. Nature Reviews Genetics 8, 196–205 (2007) doi:10-1038/nrg2053.
Prangishvili, D., Forterre, P. & Garrett, R. A. Viruses of the Archaea: A unifying view. Nature Reviews Microbiology 4, 837–848 (2006) doi:10.1038/nrmicro1527.
Raoult, D. & Forterre, P. Redefining viruses: Lessons from mimivirus. Nature Reviews Microbiology 6, 315–319 (2008) doi:10.1038/nrmicro1858.
Raoult, D. et al. The 1.2-megabase genome sequence of Mimivirus. Science 306, 1344–1350 (2004) doi:10.1126/science.1101485.
Villarreal, L. P. & DeFilippis, V. R. A hypothesis for DNA viruses as the origin of eukaryotic replication proteins. Journal of Virology 74, 7079–7084 (2000).
Xiao, C. et al. Cryo-electron microscopy of the giant Mimivirus. Journal of Molecular Biology 353, 493–496 (2005) doi:10.1016/j.jmb.2005.08.060.



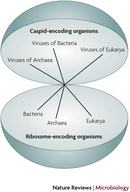
 Figure 1
Figure 1