« Prev Next »
Dominance relationships between alleles for a given trait can impact phenotypic ratios, but interactions between different genes can also impact phenotype. Such traits that result from the interaction among multiple genes and their environment are called complex traits. So, given a specific trait, how can we tell whether it is complex? One way to recognize a complex trait is through inconsistent inheritance patterns in successive generations. For example, a dominant trait might skip an entire generation yet be expressed in the subsequent generation. How is this possible? The answer to this question lies in the concepts of penetrance and expressivity.
Penetrance
In fact, large population studies are necessary for measuring penetrance, and studies of penetrance help us predict how likely it is that a trait will be evident in those who carry the underlying alleles. In general, when we know that the genotype is present but the phenotype is not observable, the trait shows incomplete penetrance. Basically, anything that shows less than 100% penetrance is an example of incomplete penetrance. Therefore, although the penetrance of a trait is a statistically calculated value based on the appearance of a phenotype among known genotypes, incomplete penetrance is simply a qualitative description about a group of known genotypes.
A specific example of incomplete penetrance is the human bone disease osteogenesis imperfecta (OI). The majority of people with this disease have a dominant mutation in one of the two genes that produce type 1 collagen, COL1A1 or COL1A2. Collagen is a tissue that strengthens bones and muscles and multiple body tissues. People with OI have weak bones, bluish color in the whites of their eyes, and a variety of afflictions that cause weakness in their joints and teeth. However, this disease doesn't affect everyone who has COLIA1 and COLIA2 mutations in the same way. In fact, some people can carry the mutation but have no symptoms. Thus, families can unknowingly transmit the mutation from one generation to the next through someone who carries the mutation but does not express the OI phenotype.
Incomplete penetrance examples such as OI demonstrate that even monogenic diseases do not have predictable expression patterns in a population. Is there a way to explain this unpredictability? Let's think about it. If two people have the same dominant mutation in COL1A1, why might only one of them actually display OI symptoms? Could it have to do with other genes that rescue the bad effect of a mutated collagen gene? Could it be that those who have OI simply express more mutated collagen than the person who is unaffected? To consider the possible explanations for incomplete penetrance, we have to remember how many steps there are between gene transcription and protein expression.
Note that the expression of other genes, such as transcriptional or translational regulators, can influence the final effect of a gene product. Anything that interferes with the pathway from transcription to protein activation is known as an epigenetic factor. Indeed, there are multiple points at which another gene product can intervene in the stages prior to the production of a protein. Interference at these stages might stop production entirely, create an altered form of the protein that might never be active, or do any number of other things that renders the gene silent. So, the final stage of an active protein reflects many different processes that lead to the amino acid sequence and ultimate protein shape, all of which can be interfered with by other genes. Furthermore, some genes can up- or downregulate rates of transcription, which changes the total amount of protein produced. Thus, genes that affect the final form and expression amount of another gene can be influential in the formation of the phenotype derived from the regulated gene (Figure 1).
So, if so many different possible modification points for a gene product exist, how can we narrow down the question of what causes incomplete penetrance? Interestingly, some scientists have actually tried to do this by observing how the genetic mutations that cause OI affect mice. These investigators inserted a mutated form of COL1A1 into mice and bred them so that they all contained this mutation. The mice were affected in similar ways to those with human OI: Many had severe bone weakness and multiple bone fractures, even at birth. In fact, when the researchers examined the mouse bones closely, they found that 70% of mice with the mutated COL1A1 gene showed evidence of OI (bone fractures); however, the remaining 30% appeared completely normal. In these mice with no OI phenotype, there was the same amount of COL1A1 expression as in those mice that did show the phenotype. Furthermore, the investigators used a purebred strain of mice that had little variability in their genomes to begin with. This means that the genetic context in which COL1A1 was expressed did not vary among the mice studied. Yet, despite the fact that all the mice had extremely similar genomes and all of them expressed the same amount of COL1A1, 30% of them did not show any OI phenotype. These results continue to be perplexing.
Therefore, even the powerful experimental techniques currently available cannot explain penetrance. The two most popular explanations for incomplete penetrance, genetic background and variable expression levels, did not explain the lack of phenotype in 30% of the mice (Pereira et al., 1994).
Expressivity
Another example of expressivity at work is the occurrence of extra toes, or polydactyly, in cats. The presence of extra toes on a cat's paw is a phenotype that emerges in groups of cats who have interbred for generations. In fact, there are several well-known groups of these cats, such as those on Key West Island (known as "Hemingway's cats"), as well as those in breeding clusters in the eastern U.S. and shores of the British Isles (Figure 2). The first to report on this phenomenon was C. H. Danforth, who studied the inheritance of polydactyly among 55 generations of cats. He observed that the polydactyly phenotype showed "good penetrance, but variable expression" because the gene always causes extra toes on the paw, but the number of extra toes varies widely from cat to cat (Danforth, 1947). Through his breeding studies, Danforth found that although a dominant allele underlies the cause of polydactyly, the degree of polydactyly depends on the condition of adjacent layered tissues in the developing limb; that is, the expression of genes in tissues surrounding tissue that will become the toe determines the degree of polydactyly (Willier, 1974).
Future Directions
The relationship between genotype and phenotype is not simple. Sometimes, dominant alleles can be silenced by other genes that minimize the appearance of the phenotype. In other cases, gene expression can be changed in subtle ways, yet it can have a large impact on phenotype. The exact causes of penetrance and expressivity are still not well understood. The more we learn about the molecular mechanisms governing genetic regulation, the more we can formulate and test hypotheses about how this variability arises.
References and Recommended Reading
Danforth, C. H. Heredity of polydactyly in the cat. Journal of Heredity 38, 107–112 (1947)
Jin, C., et al. Novel FBN1 mutations associated with predominant ectopia lentis and marfanoid habitus in Chinese patients. Molecular Vision 13, 1280–1284 (2007)
Li, D., et al. The roles of two novel FBN1 gene mutations in the genotype-phenotype correlations of Marfan syndrome and ectopia lentis patients with marfanoid habitus. Genetic Testing 12, 325–330 (2008) doi:10.1089/gte.2008.0002
Pereira, R., et al. Phenotypic variability and incomplete penetrance of spontaneous fractures in an inbred strain of transgenic mice expressing a mutated collagen gene (COL1A1). Clinical Investigation 93, 1765 (1994)
Willier, B. H. Chapter 1: Charles Haskell Danforth. In Biographical Memoirs (Washington, DC, National Academies Press, 1974)



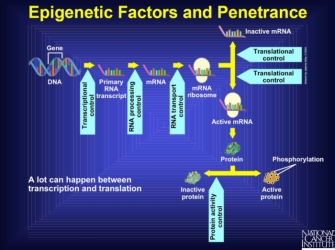
 Figure 1: Epigenetic factors.
Figure 1: Epigenetic factors.



























