Key Points
-
Recent progress has expanded our knowledge about the metabolism of the model bacterial pathogens Listeria monocytogenes, Shigella flexneri (and the closely related enteroinvasive Escherichia coli (EIEC)), Salmonella enterica subsp. enterica serovar Typhimurium and Mycobacterium tuberculosis when living inside the host cell.
-
Differences in the metabolic characteristics of these four pathogens have been elucidated in the context of the metabolism of host cell lines used for in vitro infection.
-
There are several tools available to study the metabolism of these intracellular pathogens, and differential gene expression profiling (DGEP) and 13C isotopologue analysis (13C-IPA) have been particularly fruitful; however, there are both strengths and weaknesses for these techniques.
-
Models have been suggested (mainly on the basis of data from DGEP and 13C-IPA studies) for the metabolic pathways and fluxes used by the four pathogens when replicating in their specific intracellular compartments (the cytosol or specific phagosomal vacuoles of the host cell). Each pathogen adapts specifically to the host cell environment but exhibits a surprisingly high metabolic flexibility in response to altered metabolic conditions.
-
There is limited experimental evidence for interference by the metabolism of these intracellular bacteria with the expression of virulence genes that are required for their intracellular lifestyles.
-
There is an urgent need for improved in vivo systems and more sensitive analytical tools for studying the metabolism of the bacterial pathogens in real target cells and animal models.
Abstract
New technologies such as high-throughput methods and 13C-isotopologue-profiling analysis are beginning to provide us with insight into the in vivo metabolism of microorganisms, especially in the host cell compartments that are colonized by intracellular bacterial pathogens. In this Review, we discuss the recent progress made in determining the major carbon sources and metabolic pathways used by model intracellular bacterial pathogens that replicate either in the cytosol or in vacuoles of infected host cells. Furthermore, we highlight the possible links between intracellular carbon metabolism and the expression of virulence genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ogawa, M. & Sasakawa, C. Intracellular survival of Shigella. Cell. Microbiol. 8, 177–184 (2006).
Russell, D. G. Who puts the tubercle in tuberculosis? Nature Rev. Microbiol. 5, 39–47 (2007).
Haraga, A., Ohlson, M. B. & Miller, S. I. Salmonellae interplay with host cells. Nature Rev. Microbiol. 6, 53–66 (2008).
Tsolis, R. M., Young, G. M., Solnick, J. V. & Bäumler, A. J. From bench to bedside: stealth of enteroinvasive pathogens. Nature Rev. Microbiol. 6, 883–892 (2008).
Cossart, P. & Toledo-Arana, A. Listeria monocytogenes, a unique model in infection biology: an overview. Microbes Infect. 10, 1041–1050 (2008).
Ray, K., Marteyn, B., Sansonetti, P. J. & Tang, C. M. Life on the inside: the intracellular lifestyle of cytosolic bacteria. Nature Rev. Microbiol. 7, 333–340 (2009). This recent review gives a good overview of the mechanisms enabling bacteria to live in the host cell cytosol.
Flannagan, R. S., Cosio, G. & Grinstein, S. Antimicrobial mechanisms of phagocytes and bacterial evasion strategies. Nature Rev. Microbiol. 7, 355–366 (2009).
Smith, H. Questions about the behaviour of bacterial pathogens in vivo. Philos. Trans. R. Soc. Lond. B 29, 551–564 (2000).
Muñoz-Elías, E. J. & McKinney, J. D. Carbon metabolism of intracellular bacteria. Cell. Microbiol. 8, 10–22 (2006). The first review emphasizing our limited knowledge of the metabolism of intracellular bacterial pathogens.
Brown, S. A., Palmer, K. L. & Whiteley, M. Revisiting the host as a growth medium. Nature Rev. Microbiol. 6, 657–666 (2008). An interesting discussion on the role of in vivo carbon sources and carbon metabolism pathways as targets for antibiotic development.
Beuzón, C. R. et al. Salmonella maintains the integrity of its intracellular vacuole through the action of SifA. EMBO J. 19, 3235–3249 (2000).
Beuzón, C. R., Salcedo, S. P. & Holden, D. W. Growth and killing of a Salmonella enterica serovar Typhimurium sifA mutant strain in the cytosol of different host cell lines. Microbiology 148, 2705–2715 (2002). This article and reference 11 show that a S. enterica sifA mutant can actively replicate in the cytosol of certain host cells.
van der Wel, N. et al. M. tuberculosis and M. leprae translocate from the phagolysosome to the cytosol in myeloid cells. Cell 129, 1287–1298 (2007). The first evidence that M. tuberculosis might also be able to replicate in the cytosol of macrophages.
Macfarlane, G. T. & Macfarlane, S. Human colonic microbiota: ecology, physiology and metabolic potential of intestinal bacteria. Scand. J. Gastroenterol. Suppl. 222, 3–9 (1997).
Mao, X. J. et al. Interplay between CRP-cAMP and PII-Ntr systems forms novel regulatory network between carbon metabolism and nitrogen assimilation in Escherichia coli. Nucleic Acids Res. 35, 1432–1440 (2007).
Sonenshein, A. L. Control of key metabolic intersections in Bacillus subtilis. Nature Rev. Microbiol. 5, 917–927 (2007).
Deutscher, J., Francke, C. & Postma, P. W. How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol. Mol. Biol. Rev. 70, 939–1031 (2006).
Fujita, Y. Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci. Biotechnol. Biochem. 73, 245–259 (2009).
Dorman, C. J. Global regulators and environmental adaptation in Gram-negative pathogens. Clin. Microbiol. Infect. 15, 47–50 (2009).
Fisher, S. H. Regulation of nitrogen metabolism in Bacillus subtilis: vive la différence! Mol. Microbiol. 32, 223–232 (1999).
Görke, B. & Vogel. J. Noncoding RNA control of the making and breaking of sugars. Genes Dev. 22, 2914–2925 (2008).
De Lay, N. & Gottesman, S. The Crp-activated small noncoding regulatory RNA CyaR (RyeE) links nutrional status to group behavior J. Bacteriol. 191, 461–476 (2009).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nature Rev. Microbiol. 6, 613–624 (2008). An excellent overview of the complex regulatory circuit of carbon catabolite repression and its impact on the regulation of virulence genes.
Dorman, C. J. Nucleoid-associated proteins and bacterial physiology. Adv. Appl. Microbiol. 67, 47–64 (2009).
Vazquez-Torres, A. & Fang, F. C. Cellular routes of invasion by enteropathogens. Curr. Opin. Microbiol. 3, 54–59 (2000).
Disson, O. et al. Conjugated action of two species-specific invasion proteins for fetoplacental listeriosis. Nature 455, 1114–1118 (2008).
Phalipon, A. & Sansonetti, P. J. Shigella's ways of manipulating the host intestinal innate and adaptive immune system: a tool box for survival? Immunol. Cell Biol. 85, 119–129 (2007).
Disson, O. et al. Modeling human listeriosis in natural and genetically engineered animals. Nature Protoc. 4, 799–810 (2009). This article demonstrates the power of using transgenic animals to study the pathogenesis of human-adapted pathogens.
Giacomodonato, M. N. et al. SipA, SopA, SopB, SopD and SopE2 effector proteins of Salmonella enterica serovar Typhimurium are synthesized at late stages of infection in mice. Microbiology 153, 1221–1228 (2007).
Russell, D. G. Who puts the tubercle in tuberculosis? Nature Rev. Microbiol. 5, 39–47 (2007).
Pieters, J. Mycobacterium tuberculosis and the macrophage: maintaining a balance. Cell Host Microbe 12, 399–407 (2008).
Hwang, C., Sinskey, A. J. & Lodish, H. F. Oxidized redox state of glutathione in the endoplasmic reticulum. Science 257, 1496–1502 (1992).
De Domenico, I., McVey Ward, D., Kaplan, J. Regulation of iron acquisition and storage: consequences for iron-linked disorders. Nature Rev. Mol. Cell. Biol. 9, 72–81 (2008).
Goetz, M. et al. Microinjection and growth of bacteria in the cytosol of mammalian host cells. Proc. Natl Acad. Sci. USA 98, 12221–12226 (2001).
Slaghuis, J., Goetz, M., Engelbrecht, F. & Goebel, W. Inefficient replication of Listeria innocua in the cytosol of mammalian cells. J. Infect. Dis. 198, 393–401 (2004).
Köhler, S. et al. The intramacrophagic environment of Brucella suis and bacterial response. Vet. Microbiol. 90, 299–309 (2002).
Garcia-del Portillo, F., Nunez-Hernandez, C. Eisman, B. & Ramos-Vivas, J. Growth control in the Salmonella-containing vacuole. Curr. Opin. Microbiol. 11, 46–52 (2008).
Jin, Q. et al. Genome sequence of Shigella flexneri 2a: insights into pathogenicity through comparison with genomes of Escherichia coli K12 and O157. Nucleic Acids Res. 30, 4432–4441 (2002).
Glaser, P. et al. Comparative genomics of Listeria species. Science 294, 849–852 (2001).
Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
McClelland, M. et al. Complete genome sequence of Salmonella enterica serovar Typhimurium LT2. Nature 413, 852–856 (2001).
Eisenreich, W. et al. 13C isotopologue perturbation studies of Listeria monocytogenes carbon metabolism and its modulation by the virulence regulator PrfA. Proc. Natl Acad. Sci. USA 103, 2040–2045 (2006).
Joseph, B. & Goebel, W. Life of Listeria monocytogenes in the host cells' cytosol. Microbes Infect. 9, 1188–1195 (2007).
Premaratne, R. J., Lin, W. J. & Johnson, E. A. Development of an improved chemically defined minimal medium for Listeria monocytogenes. Appl. Environ. Microbiol. 57, 3046–3048 (1991).
Niederweis, M. Nutrient acquisition by mycobacteria. Microbiology 154, 679–692 (2008).
Van der Geize, R. et al. A gene cluster encoding cholesterol catabolism in a soil actinomycete provides insight into Mycobacterium tuberculosis survival in macrophages. Proc. Natl Acad. Sci. USA 104, 1947–1952 (2007). A discussion of the importance of cholesterol as a carbon source for the intracellular growth of M. tuberculosis.
Zhao, F. Q. & Keating, A. F. Functional properties and genomics of glucose transporters. Curr. Genomics 8, 113–128 (2007).
Cheeseman, C. Role of intestinal basolateral membrane in absorption of nutrients. Am. J. Physiol. 263, R482–R488 (1992).
Wright, E. M. et al. Active sugar transport in eukaryotes. J. Exp. Biol. 196, 197–212 (1994).
Wood, I. S. & Trayhurn, P. Glucose transporters (GLUT and SGLT): expanded families of sugar transport proteins. Br. J. Nutr. 89, 3–9 (2003).
Harris, A. L. Hypoxia: a key regulatory factor in tumour growth. Nature Rev. Cancer 2, 38–47 (2002).
Van der Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg Effect: the metabolic requirements of cell proliferation. Science. 324, 1029–1033 (2009).
Maxwell, P. H., Pugh, C. W. & Ratcliffe, P. J. Activation of the HIF pathway in cancer. Curr. Opin. Genet. Dev. 11, 293–299 (2001).
Rupp, J. et al. Chlamydia pneumoniae directly interferes with HIF-1α stabilization in human host cells. Cell. Microbiol. 9, 2181–2191 (2007).
Dehne, N. & Brüne, B. HIF-1 in the inflammatory microenvironment. Exp. Cell. Res. 315, 1791–1797 (2009).
Saenz, H. L. & Dehio, C. Signature-tagged mutagenesis: technical advances in a negative selection method for virulence gene identification. Curr. Opin. Microbiol. 8, 612–619 (2005).
Jansen, A. & Yu, J. Differential gene expression of pathogens inside infected hosts. Curr. Opin. Microbiol. 9, 138–142 (2006).
Sauer, U. Metabolic networks in motion: 13C-based flux analysis. Mol. Syst. Biol. 2, 62 (2006). This review provides a comprehensive overview on the potential of 13C-based methods for unraveling metabolic networks and carbon fluxes.
Eylert, E. et al. Carbon metabolism of Listeria monocytogenes growing inside macrophages. Mol. Microbiol. 69, 1008–1017 (2008). The first application of the 13C-IPA method for studying the carbon metabolism of an intracellular pathogen in host cells.
Zamboni, N., Fendt, S. Ruhl, M. & Sauer, U. 13C-based metabolic flux analysis. Nature Protoc. 4, 878–892 (2009).
Munger, J. et al. Systems-level metabolic flux profiling identifies fatty acid synthesis as a target for antiviral therapy. Nature Biotech. 26, 1179–1186 (2008).
Nielsen, J. & Oliver, S. The next wave in metabolome analysis. Trends Biotechnol. 23, 544–546 (2005).
Nicholson, J. K & Lindon, J. C. Metabonomics. Nature 455, 1054–1056 (2008).
Breitling, R., Vitkup, D. & Barrett, M. P. New surveyor tools for charting microbial metabolic maps. Nature Rev. Microbiol. 6, 156–161 (2008).
Swain, R. J. & Stevens, M. M. Raman microspectroscopy for non-invasive biochemical analysis of single cells. Biochem. Soc. Trans. 35, 544–549 (2007).
Wagner, M., Single-cell ecophysiology of microbes as revealed by Raman microspectroscopy or secondary ion mass spectrometry imaging. Ann. Rev. Microbiol. 63, 411–429 (2009).
Melican, K., Richter-Dahlfors, A. Real-time live imaging to study bacterial infections in vivo. Curr. Opin. Microbiol. 12, 31–36 (2009).
Bumann, D. & Valdivia, R. H. Identification of host-induced pathogen genes by differential fluorescence induction reporter systems. Nature Protoc. 2, 770–777 (2007).
Runyen-Janecky, L. J. & Payne, S. M. Identification of chromosomal Shigella flexneri genes induced by the eukaryotic intracellular environment. Infect. Immun. 70, 4379–4388 (2002).
Eriksson, S., Lucchini, S., Thompson, A., Rhen, M. & Hinton, J. C. Unravelling the biology of macrophage infection by gene expression profiling of intracellular Salmonella enterica. Mol. Microbiol. 47, 103–118 (2003).
Schnappinger, D. et al. Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: insights into the phagosomal environment. J. Exp. Med. 198, 693–704 (2003). The first comprehensive transcriptome analysis of M. tuberculosis in resting and activated macrophages, with a discussion of the metabolic implications.
Lucchini, S., Liu, H., Jin, Q., Hinton, J. C. & Yu, J. Transcriptional adaptation of Shigella flexneri during infection of macrophages and epithelial cells: insights into the strategies of a cytosolic bacterial pathogen. Infect. Immun. 73, 88–102 (2005). The first comprehensive comparative transcriptome analysis of intracellularly replicating S. flexneri.
Joseph, B. et al. Identification of Listeria monocytogenes genes contributing to intracellular replication by expression profiling and mutant screening. J. Bacteriol. 188, 556–568 (2006).
Chatterjee, S. S. et al. Intracellular gene expression profile of Listeria monocytogenes. Infect. Immun. 74, 1323–1338 (2006). This article and reference 73 show the first comprehensive transcriptome analyses of intracellularly replicating L. monocytogenes.
Hautefort, I. et al. During infection of epithelial cells Salmonella enterica serovar Typhimurium undergoes a time-dependent transcriptional adaptation that results in simultaneous expression of three type 3 secretion systems. Cell. Microbiol. 10, 958–984 (2008). A thorough transcriptome analysis of intracellularly replicating S. enterica.
La, M. V., Raoult, D. & Renesto, P. Regulation of whole bacterial pathogen transcription within infected hosts. FEMS Microbiol. Rev. 32, 440–460 (2008).
Chico-Calero, I. et al. Hpt, a bacterial homolog of the microsomal glucose-6-phosphate translocase, mediates rapid intracellular proliferation in Listeria. Proc. Natl Acad. Sci. USA 99, 431–436 (2002). This article demonstrates that a metabolic gene partially needed for the metabolism of intracellular L. monocytogenes is co-regulated with other virulence genes.
Goetz, A., Eylert, E., Eisenreich, W. & Goebel, W. Carbon metabolism of enterobacterial human pathogens growing in epithelial colorectal adenocarcinoma (Caco-2) cells. PLoS ONE (in the press).
Stoll, R. & Goebel, W. The major PEP-phosphotransferase systems (PTS) for glucose, mannose and cellobiose of Listeria monocytogenes and their significance for extra-and intracellular growth. Microbiology 156, 1069–1083 (2010).
Joseph, B. et al. Glycerol metabolism and PrfA activity in Listeria monocytogenes. J. Bacteriol. 190, 5412–5430 (2008). This study shows the essential role of PTS-dependent and PTS-independent transport of carbon substrates for the modulation of PrfA activity.
Schaer, J. et al. Pyruvate carboxylase plays a crucial role in carbon metabolism of extra- and intracellularly replicating Listeria monocytogenes. J. Bacteriol. 192, 1774–1784 (2010).
Marquis, H., Bouwer, H. G., Hinrichs, D. J. & Portnoy, D. A. Intracytoplasmic growth and virulence of Listeria monocytogenes auxotrophic mutants. Infect. Immun. 61, 3756–3760 (1993).
Stritzker, J. et al. Growth, virulence, and immunogenicity of Listeria monocytogenes aro mutants. Immun. Infect. 72, 5622–5629 (2004).
Schauer, K., Stolz, J., Scherer, S. & Fuchs, T. M. Both thiamine uptake and biosynthesis of thiamine precursors are required for intracellular replication of Listeria monocytogenes. J. Bacteriol. 191, 2218–2227 (2009).
O'Riordan, M., Moors, M. A. & Portnoy, D. A. Listeria intracellular growth and virulence require host-derived lipoic acid. Science 302, 462–464 (2003).
Keeney, K., Colosi, L., Weber, W. & O'Riordan, M. Generation of branched-chain fatty acids through lipoate-dependent metabolism facilitates intracellular growth of Listeria monocytogenes. J. Bacteriol. 191, 2187–2196 (2009).
Noriega, F. R. et al. Engineered ΔguaB-A ΔvirG Shigella flexneri 2a strain CVD 1205: construction, safety, immunogenicity, and potential efficacy as a mucosal vaccine. Infect. Immun. 64, 3055–3061 (1996).
Cersini, A., Martino, M. C., Martini, I., Rossi, G. & Bernardini, M. L. Analysis of virulence and inflammatory potential of Shigella flexneri purine biosynthesis mutants. Infect. Immun. 71, 7002–7013 (2003).
Bumann, D. System-level analysis of Salmonella metabolism during infection. Curr. Opin. Microbiol. 12, 1–9 (2009).
Bowden, S. D., Rowley, G., Hinton, J. C. D. & Thompson, A. Glucose and glycolysis are required for the successful infection of macrophages and mice by Salmonella enterica serovar Typhimurium. Infect. Immun. 77, 3117–3126, (2009).
García-del Portillo, F., Núñez-Hernández, C., Eisman, B. & Ramos-Vivas, J. Growth control in the Salmonella-containing vacuole. Curr. Opin. Microbiol. 11, 46–52 (2008).
Tchawa Yimga, M. et al. Role of gluconeogenesis and the tricarboxylic acid cycle in the virulence of Salmonella enterica serovar Typhimurium in BALB/c mice. Infect. Immun. 74, 1130–1140 (2006).
Mercado-Lubo, R., Leatham, M. P., Conway, T. & Cohen, P. S. Salmonella enterica serovar Typhimurium mutants unable to convert malate to pyruvate and oxaloacetate are avirulent and immunogenic in BALB/c mice. Infect. Immun. 77, 1397–1405 (2009). An interesting recent study on the carbon metabolism of S . Typhimurium in an animal model.
Fang, F. C., Libby, S. J., Castor, M. E. & Fung, A. M. Isocitrate lyase (AceA) is required for Salmonella persistence but not for acute lethal infection in mice. Infect. Immun. 73, 2547–2549 (2005).
Fields, P. I., Swanson, R. V., Haidaris, C. G. & Heffron, F. Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Natl Acad. Sci. USA 83, 5189–5193 (1986).
McFarland, W. C & Stocker, B. A. Effect of different purine auxotrophic mutations on mouse-virulence of a Vi-positive strain of Salmonella dublin and of two strains of Salmonella typhimurium. Microb. Pathog. 3, 129–141 (1987).
Leung, K. Y. & Finlay, B. B. Intracellular replication is essential for the virulence of Salmonella typhimurium. Proc. Natl Acad. Sci. USA 88, 11470–11474 (1991).
Tailleux, L. et al. Probing host pathogen cross-talk by transcriptional profiling of both Mycobacterium tuberculosis and infected human dendritic cells and macrophages. PLoS ONE 3, e1403 (2008).
Cappelli, G. et al. Profiling of Mycobacterium tuberculosis gene expression during human macrophage infection: upregulation of the alternative sigma factor G, a group of transcriptional regulators, and proteins with unknown function. Res. Microbiol. 157, 445–455 (2006).
Talaat, A. M, Lyons, R., Howard, S.T & Johnston, S. A. The temporal expression profile of Mycobacterium tuberculosis infection in mice. Proc. Natl Acad. Sci. USA 101, 4602–4607 (2004).
Talaat, A. M. et al. Mycobacterial bacilli are metabolically active during chronic tuberculosis in murine lungs: insights from genome-wide transcriptional profiling. J. Bacteriol. 189, 4265–4274 (2007).
Kendall, S. L. et al. A highly conserved transcriptional repressor controls a large regulon involved in lipid degradation in Mycobacterium smegmatis and Mycobacterium tuberculosis. Mol. Microbiol. 65, 684–699 (2007).
Boshoff, H. I. & Barry, C. E. 3rd. Tuberculosis — metabolism and respiration in the absence of growth. Nature Rev. Microbiol. 3, 70–80 (2005). A critical review of the metabolism of intracellular M. tuberculosis , with emphasis on energy metabolism in the absence of mycobacterial growth.
Casali, N. & Riley, L. W. A phylogenomic analysis of the Actinomycetales mce operons. BMC Genomics 8, 60 (2007).
Joshi, S. M. et al. Characterization of mycobacterial virulence genes through genetic interaction mapping. Proc. Natl Acad. Sci. USA 103, 11760–11765 (2006).
Santangelo, M. P. et al. Mce3R, a TetR-type transcriptional repressor, controls the expression of a regulon involved in lipid metabolism in Mycobacterium tuberculosis. Microbiology 155, 2245–2255 (2009).
Gioffré, A. et al. Mutation in mce operons attenuates Mycobacterium tuberculosis virulence. Microbes Infect. 7, 325–334 (2005).
Pandey, A. K. & Sassetti, C. M. Mycobacterial persistence requires the utilization of host cholesterol. Proc. Natl Acad. Sci. USA 105, 4376–4380 (2008).
McKinney, J. D. et al. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature 406, 735–738 (2000). A classic study showing the importance of the glyoxylate shunt and fatty acid catabolism in M. tuberculosis in the persistent stage of infection in mice.
Muñoz-Elías, E. J. & McKinney, J. D. Mycobacterium tuberculosis isocitrate lyases 1 and 2 are jointly required for in vivo growth and virulence. Nature Med. 11, 638–644 (2005).
Muñoz-Elías, E. J., Upton, A. M., Cherian, J. & McKinney, J. D. Role of the methylcitrate cycle in Mycobacterium tuberculosis metabolism, intracellular growth, and virulence. Mol. Microbiol. 60, 1109–1122 (2006).
Gould, T. A., van de Langemheen, H., Muñoz-Elías, E. J., McKinney, J. D. & Sacchettini, J. C. Dual role of isocitrate lyase 1 in the glyoxylate and methylcitrate cycles in Mycobacterium tuberculosis. Mol. Microbiol. 61, 940–947 (2006).
Rengarajan, J., Bloom, B. R. & Rubin, E. J. Genome-wide requirements for Mycobacterium tuberculosis adaptation and survival in macrophages. Proc. Natl Acad. Sci. USA 102, 8327–8332 (2005).
Liu, K., Yu, J. & Russell, D. G. pckA-deficient Mycobacterium bovis BCG shows attenuated virulence in mice and in macrophages. Microbiology 149, 1829–1835 (2003).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003).
Keating, L. A. et al. The pyruvate requirement of some members of the Mycobacterium tuberculosis complex is due to an inactive pyruvate kinase: implications for in vivo growth. Mol. Microbiol. 56, 163–174 (2005).
Tran Van Nhieu, G., Bourdet-Sicard, R., Duménil, G., Blocker, A. & Sansonetti, P. J. Bacterial signals and cell responses during Shigella entry into epithelial cells. Cell. Microbiol. 2, 187–193 (2000).
Dussurget, O. New insights into determinants of Listeria monocytogenes virulence. Int. Rev. Cell. Mol. Biol. 270, 1–38 (2008).
Grassl, G. A. & Finlay, B. B. Pathogenesis of enteric Salmonella infections. Curr. Opin. Gastroenterol. 24, 22–26 (2008).
McGhie, E. J., Brawn, L. C., Hume, P. J., Humphreys, D. & Koronakis, V. Salmonella takes control: effector-driven manipulation of the host. Curr. Opin. Microbiol. 12, 117–124 (2009).
Parsot, C. Shigella type III secretion effectors: how, where, when, for what purposes? Curr. Opin. Microbiol. 12, 110–116 (2009).
Mehrotra, J. & Bishai, W. R. Regulation of virulence genes in Mycobacterium tuberculosis. Int. J. Med. Microbiol. 291, 171–182 (2001).
Houben, E. N. et al. Differential expression of a virulence factor in pathogenic and non-pathogenic mycobacteria. Mol. Microbiol. 72, 41–52 (2009).
Poncet, S. et al. in Bacterial Sensing and Signaling. Contributions to Microbiology Vol. 16 (eds Collin, M. & Schuch, R.) 1–15 (Karger, Basel, 2009).
Hansen-Wester, I. & Hensel, M. Salmonella pathogenicity islands encoding type III secretion systems. Microbes Infect. 3, 549–559 (2001).
Jones, B. D. Salmonella invasion gene regulation: a story of environmental awareness. J. Microbiol. 43, 110–117 (2005).
Ellermeier, J. R. & Slauch, J. M. Adaptation to the host environment: regulation of the SPI1 type III secretion system in Salmonella enterica serovar Typhimurium. Curr. Opin. Microbiol. 10, 24–29 (2007).
Cirillo, D. M., Valdivia, R. H., Monack, D. M. & Falkow, S. Macrophage-dependent induction of the Salmonella pathogenicity island 2 type III secretion system and its role in intracellular survival. Mol. Microbiol. 30, 175–188 (1998).
Worley, M. J., Ching, K. H. & Heffron, F. Salmonella SsrB activates a global regulon of horizontally acquired genes. Mol. Microbiol. 36, 749–761 (2000).
Le Gall, T. et al. Analysis of virulence plasmid gene expression defines three classes of effectors in the type III secretion system of Shigella flexneri. Microbiology 151, 951–962 (2005).
Prosseda, G. et al. A role for H-NS in the regulation of the virF gene of Shigella and enteroinvasive Escherichia coli. Res. Microbiol. 149, 15–25 (1998).
Beloin, C. & Dorman, C. J. An extended role for the nucleoid structuring protein H-NS in the virulence gene regulatory cascade of Shigella flexneri. Mol. Microbiol. 47, 825–838 (2003).
Bajaj, V. Hwang, C. & Lee, C. A. hilA is a novel ompR/toxR family member that activates the expression of Salmonella typhimurium invasion genes. Mol. Microbiol. 18, 715–727 (1995).
Ahmer, B. M., van Reeuwijk, J., Watson, P. R., Wallis, T. S. & Heffron, F. Salmonella SirA is a global regulator of genes mediating enteropathogenesis. Mol. Microbiol. 31, 971–982 (1999).
Lawhon, S. D., Maurer, R., Suyemoto, M. & Altier, C. Intestinal short-chain fatty acids alter Salmonella typhimurium invasion gene expression and virulence through BarA/SirA. Mol. Microbiol. 46, 1451–1464 (2002).
Altier, C. Genetic and environmental control of Salmonella invasion J. Microbiol. 43, 85–92 (2005).
Santangelo, M. P. et al. Study of the role of Mce3R on the transcription of mce genes of Mycobacterium tuberculosis. BMC Microbiol. 8, 38 (2008).
Engelbrecht, F. et al. A new PrfA-regulated gene of Listeria monocytogenes encoding a small, secreted protein which belongs to the family of internalins. Mol. Microbiol. 21, 823–837 (1996).
Rajabian, T. et al. The bacterial virulence factor InlC disturbs apical cell junctions and promotes cell-to-cell spread of Listeria. Nature Cell Biol. 11, 1212–1218 (2009).
Scortti, M., Monzó, H. J., Lacharme-Lora, L., Lewis, D. A. & Vázquez-Boland, J. A. The PrfA virulence regulon. Microbes Infect. 9, 1196–1207 (2007).
Johansson, J. et al., A thermosensor controls expression of virulence genes in Listeria monocytogenes. Cell 110, 551–561 (2002).
Behari, J. & Youngman, P. A homolog of CcpA mediates catabolite control in Listeria monocytogenes but not carbon source regulation of virulence genes. J. Bacteriol. 180, 6316–6324 (1998).
Ermolaeva, S. et al. Negative control of Listeria monocytogenes virulence genes by a diffusible autorepressor. Mol. Microbiol. 52, 601–611 (2004).
Stoll, R., Mertins, S., Joseph, B., Müller-Altrock, S. & Goebel, W. Modulation of PrfA activity in Listeria monocytogenes upon growth in different culture media. Microbiology 154, 3856–3876 (2008).
Mertins, S. et al. Interference of components of the phosphoenolpyruvate phosphotransferase system with the central virulence gene regulator PrfA of Listeria monocytogenes. J. Bacteriol. 189, 473–490 (2007).
Freitag, N. E., Port, G. C. & Miner, M. D. Listeria monocytogenes — from saprophyte to intracellular pathogen. Nature Rev. Microbiol. 7, 623–628 (2009).
Szyperski, T. Biosynthetically directed fractional 13C-labeling of proteinogenic amino acids. An efficient analytical tool to investigate intermediary metabolism. Eur. J. Biochem. 232, 433–448 (1995).
Bacher, A. et al. Elucidation of biosynthetic pathways and metabolic flux patterns via retrobiosynthetic NMR analysis, FEMS Microbiol. Rev. 22, 567–598 (1999).
Fischer, E. & Sauer, U. Metabolic flux profiling of Escherichia coli mutants in central carbon metabolism using GC-MS. Eur. J. Biochem. 270, 880–891 (2003).
Holmes, H. Flux analysis and control of the central metabolic pathways in Escherichia coli. FEMS Microbiol. Rev. 19, 85–116 (1996). A classic paper and good overview of background information about the central metabolic pathways and fluxes in heterotrophic bacteria.
Acknowledgements
Work in the authors' laboratories is supported by grants from the German Research Foundation (DFG) (including grant numbers SPP1316, SFB479 and TR34). We thank E. Eylert for valuable suggestions and editorial help, and for allowing us to cite unpublished results. We thank A. Bacher, R. Gross and R. Haas for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (figure)
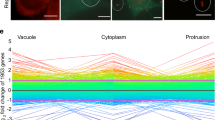
DGEP data compiled from published reports1,2, showing the Listeria monocytogenes genes encoding enzymes involved in the central metabolic pathways that are up-or down-regulated in L. monocytogenes grown in J774 macrophages and in BHI culture medium (up-regulated genes are marked with red and down-regulated ones with green arrows, respectively; blue arrows indicate non-differentially expressed genes. (PDF 189 kb)
Supplementary information S2 (figure)
a | DGEP data3 obtained with RNA derived from S. flexneri grown in U937 macrophages (numbers refer to RNA expression values after normalization) compared to RNA derived from S. flexneri grown in LB medium (for further details see3). b | DGEP data1 obtained with RNA derived from S. flexneri grown in HeLa cells (green numbers) compared to RNA derived from S. flexneri grown in LB medium (for further details see1). Up–regulated genes are indicated by red arrows and down–regulated ones by green arrows; blue arrows indicate non–differentially expressed genes. (PDF 201 kb)
Supplementary information S3 (figure)
a | DGEP data compiled from published report1, showing the up-and down-regulated genes, coding for enzymes involved in the central metabolic pathways of S. Typhimurium grown in J774 macrophages compared to RNA derived from S. Typhimurium grown in LB medium (for further details see1). b | DGEP data compiled from published report1, showing the up-and down-regulated genes, coding for enzymes involved in the central metabolic pathways of S. Typhimurium grown in J774 macrophages compared to RNA derived from S. Typhimurium grown in RPMI medium with glucose; for further details see1. Up–regulated genes are indicated by red arrows and down-regulated by green arrows, respectively; blue arrows indicate non–differentially expressed genes. Note that DGEF values in part b are dramatically different from those in part a for several genes. (PDF 208 kb)
Supplementary information S4 (figure)
DGEP data compiled from the published report5, showing the up-and down-regulated genes, coding for enzymes involved in the central metabolic pathways of M. tuberculosis. (PDF 193 kb)
Related links
Related links
DATABASES
Entrez Genome Project
Mycobacterium bovis bacille Calmette–Guérin
Salmonella enterica subsp. enterica serovar Typhimurium
FURTHER INFORMATION
Glossary
- Heterotroph
-
An organism that uses organic carbon compounds as energy sources and substrates for all carbon intermediates.
- Granuloma
-
A ball-shaped assembly of mononuclear cells that is formed when the immune system attempts to wall off substances that it perceives as foreign but is unable to eliminate.
- Prototrophic
-
The ability of an organism to synthesize all the essential organic compounds required for its growth itself.
- Anapleurotic reaction
-
A reaction that replenishes intermediates of the central metabolic pathways.
- Glyoxylate shunt
-
An anapleurotic pathway from some bacteria (and some higher plants), involving isocitrate lyase and malate synthase, which together convert isocitrate of the TCA cycle to malate or oxaloacetate.
- Anaerobiosis
-
The production of energy by an organism without the involvement of oxygen.
- Isotopologue
-
A molecular species that differs from another only in containing one or more heavier atoms (owing to these atoms having a different number of neutrons).
- Pathogenicity island
-
A discrete genetic unit (with a distinct GC content and a size ranging from 10 to 200 kb) in bacteria, often flanked by direct repeats and often inserted into tRNA genes. These islands usually carry genes that contribute to the virulence of the respective pathogen.
Rights and permissions
About this article
Cite this article
Eisenreich, W., Dandekar, T., Heesemann, J. et al. Carbon metabolism of intracellular bacterial pathogens and possible links to virulence. Nat Rev Microbiol 8, 401–412 (2010). https://doi.org/10.1038/nrmicro2351
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2351
This article is cited by
-
Nutrient and salt depletion synergistically boosts glucose metabolism in individual Escherichia coli cells
Communications Biology (2022)
-
Environment dependent expression of mycobacterium hormone sensitive lipases: expression pattern under ex-vivo and individual in-vitro stress conditions in M. tuberculosis H37Ra
Molecular Biology Reports (2022)
-
In vitro passage alters virulence, immune activation and proteomic profiles of Burkholderia pseudomallei
Scientific Reports (2020)
-
Identification of differentially regulated proteins of Caenorhabditis elegans during Salmonella enterica Serovar Typhi exposure using Mass Spectrometry
Journal of Proteins and Proteomics (2020)