Abstract
Perineuronal nets (PNNs), proteoglycan-rich extracellular matrix structures, are thought to be expressed around inhibitory neurons and contribute to critical periods of brain function and synaptic plasticity. However, in some specific brain regions such as the amygdala, PNNs were predominantly expressed around excitatory neurons. These neurons were recruited during auditory fear conditioning and memory retrieval. Indeed, the activation of PNN-expressing excitatory neurons predicted cognitive performance.
Similar content being viewed by others
Introduction
Some neurons are enwrapped by a specialized matrix structure called a perineuronal net (PNN), which acts as a diffusion barrier for ion channels and receptors, scaffold binding for signaling molecules, and an ionic buffer1. Therefore, PNNs modulate synaptic transmission and neuronal activity2. PNNs in the adult brain are expressed in a region- and cell-specific manner; in particular, they are expressed around parvalbumin (PV)-positive inhibitory neurons3,4, although some glutamatergic neurons express PNNs5. Recent studies using enzymes that degrade PNNs demonstrate that PNNs regulate synaptic plasticity, critical period closure, and fear memory6,7,8. Despite these important functions, it is unclear what differentiates neurons with PNNs from those without. Here, we show that PNNs are mainly expressed around excitatory neurons in brain regions related to emotional learning and memory. We then compared the expression of c-Fos, a marker of neuronal activity9, between neurons with and without PNNs after fear conditioning, which is a form of associative learning10.
Results
PNNs are predominantly expressed around excitatory neurons in specific brain regions, including the amygdala
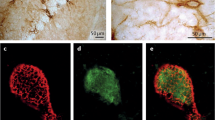
We investigated the overall distribution of PNN using Wisteria floribunda agglutinin (WFA), a ubiquitous marker for PNNs11, in GAD67-GFP knock-in adult mice in which GABAergic neurons are specifically labeled by GFP12. These mice have a normal GABA content in the adult brain and exhibit no abnormality of the brain at the macroscopic level12. WFA-positive PNNs were widely observed around GFP-positive inhibitory neurons throughout the adult brain, a finding consistent with those reported previously3,13 (Fig. 1A); however, we found it surprising that WFA-labeling and GFP fluorescence were mutually exclusive in some regions, including the dorsal tenia tecta, the piriform cortex, the lateral amygdala, the basolateral amygdala, the basomedial amygdala, the hippocampal CA2 region, the ventromedial hypothalamus, the entorhinal cortex, the temporal association cortex, and the ectorhinal cortex, even though these regions contained many GFP-positive GABAergic neurons (Fig. 1B). These PNN-expressing non-GABAergic neurons were labeled with calcium/calmodulin-dependent protein kinase II (CaMKII), a marker of excitatory neurons (Fig. 1C). Thus, PNNs are predominantly expressed around excitatory neurons rather than inhibitory neurons in these brain regions. The brain regions are associated with emotional memory formation14. Therefore, we reasoned that PNN-expressing excitatory neurons contribute to fear memory.
Merged images showing signals generated by WFA (red), GFP (green), or an anti-CaMKII antibody (cyan) in GAD67-GFP knock-in mice (A,B) or C57BL/6J mice (C). A subpopulation of GFP-expressing GABAergic neurons is enwrapped by WFA-positive PNNs (yellow arrows; A,B); however, WFA-positive PNNs also enwrap some neurons that do not express GFP (white arrowheads; B) or neurons that express CaMKII (white arrowheads; C).
PNN-expressing excitatory neurons are spatially clustered in the amygdala
To verify the abovementioned hypothesis, we first examined the types and distribution of PNN-expressing neurons in the lateral (LA) and basolateral (BA) nuclei of the amygdala, in which plasticity occurs when sensory cues are paired with an aversive stimulus10. All PNN-expressing GABAergic neurons contained PV (Fig. S1A and Table S1), and all PNN-expressing PV-negative (PNN-PV−) neurons expressed CaMKII (Fig. 2A,B and Table S1) and neurogranin15 (Fig. S1B), but showed no immunoreactivity for GABA (Fig. S1C). These data demonstrate that the population of PNN neurons in the LA and BA consists of two subpopulations: PNN-PV-positive (PV+) inhibitory neurons and PNN-PV− excitatory neurons. Therefore, an anti-PV antibody was used in the following experiments to distinguish between PNN-expressing inhibitory neurons and PNN-expressing excitatory neurons in the LA and BA. There were more PNN-PV− neurons than PNN-PV+ neurons in these regions, except for the rostral portions of the LA and BA (Fig. 2C,D). Moreover, a large percentage of PV+ neurons in the medial and caudal portions of the BA were not sheathed by PNNs (Fig. S2), in contrast to previous descriptions of PV+ neurons in the neocortex and the hippocampus16,17. Because the proteoglycan aggrecan is a major PNN component18, we next examined expression of aggrecan in WFA-labeled neurons in the LA and BA. Aggrecan immunoreactivity largely overlapped with WFA-labeling (Fig. 2E, upper). High-magnification images showed that PNN-CaMKII+ neurons exhibited immunoreactivity for aggrecan (Fig. 2E, lower). The pattern resembled that seen for WFA (Fig. 2E, lower).
(A) Low-magnification confocal microscopy image of cells double-labeled with WFA (red) and anti-PV antibody (green) in the caudal portions of the LA and BA. PNNs enwrap not only PV+ (yellow arrows) but also PV− (white arrowheads) neurons. Scale bar, 200 μm. (B) Triple fluorescence signals in the LA generated by WFA (red), an anti-PV antibody (green), and an anti-CaMKII antibody (cyan), as well as a merged image (Merge). Arrowheads indicate that WFA is detected in neurons expressing CaMKII, but not in neurons expressing PV. n = 6 (91/91 neurons in 12 slices). Scale bar, 20 μm. (C) The average density of PNN neurons that are PV− (red) or PV+ (green) in a single slice of one hemisphere throughout the rostrocaudal axis of the LA (left) and BA (right). Error bars indicate SEM. n = 9 mice, t-test, **P < 0.01, ***P < 0.001. (D) Percentage of PNN neurons throughout the rostrocaudal axis of the LA (upper) and BA (lower) that are PV− (red) or PV+ (green). n = 9 mice (224 neurons in 33 slices were analyzed). (E) Low-magnification confocal microscopy images of double-labeling (arrowheads) for WFA (left, red) and aggrecan (right, green) in the caudal LA and BA (upper). Triple-labeled fluorescence signals generated by WFA (red), anti-aggrecan (green), and anti-CaMKII antibodies (cyan) in the LA, and a merged image (Merge) (lower). WFA/CaMKII double-labeling is detected in a neuron expressing aggrecan. n = 3 mice. Scale bar, 200 μm (upper) or 20 μm (lower). (F) Distribution of PNN-PV− (red, 73 neurons) and PNN-PV+ (green, 33 neurons) neurons in the caudal LA and BA that were analyzed in (D). (G) Distance from a PNN-PV− neuron to its nearest neighbor in the LA (left) and BA (right) constructed from 129 neurons and 209 neurons in (F), respectively. Upper panel, representative image analysis.
Figure 2F summarizes the distribution of PNN-PV− and PNN-PV+ neurons in 12 brain slices obtained from nine animals. PNNs in the medial and caudal regions of the amygdala exhibited an unusual distribution pattern. Whereas a large proportion of PNN-PV+ neurons were localized in the dorsal LA, PNN-PV− neurons were spatially concentrated in the ventral LA and in the lateral portion of the BA (Fig. 2F). The mean distance  from a given PNN-PV− neuron to its nearest neighbor in the LA was 89.9 ± 7.5 μm, and that in the BA was 84.9 ± 4.4 μm (Fig. 2G). Assuming that PNN-PV− neurons are randomly distributed, the expected mean distance
from a given PNN-PV− neuron to its nearest neighbor in the LA was 89.9 ± 7.5 μm, and that in the BA was 84.9 ± 4.4 μm (Fig. 2G). Assuming that PNN-PV− neurons are randomly distributed, the expected mean distance  in the LA was 164.8 ± 11.4 μm and that in the BA was 169.1 ± 9.3 μm (LA: n = 8; t-test, t7 = 3.97, P = 0.0081; BA: n = 9; t-test, t8 = 6.04, P = 0.003; see Methods), suggesting that PNN-PV− neurons are spatially clustered.
in the LA was 164.8 ± 11.4 μm and that in the BA was 169.1 ± 9.3 μm (LA: n = 8; t-test, t7 = 3.97, P = 0.0081; BA: n = 9; t-test, t8 = 6.04, P = 0.003; see Methods), suggesting that PNN-PV− neurons are spatially clustered.
PNN-expressing excitatory neurons express c-Fos during fear conditioning
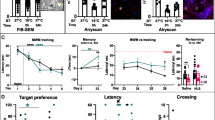
We next measured c-Fos expression in PNN-PV− neurons in the ventrolateral subdivision of the LA after fear conditioning and after a fear memory retrieval test (Fig. 3A). Mice were exposed to five pairings of a white-noise cue (conditioned stimulus, CS) either alone or combined with a mild foot shock (unconditioned stimulus, US), which resulted in an increase in freezing behavior with successive CS-US pairings, but not with CS-alone (Fig. 3B). Fear conditioning resulted in a significant increase in the total number of c-Fos+ neurons (51.3 ± 3.7 neurons/area) compared with that in the control groups (home cage: 12.2 ± 2.1 neurons/area, n = 6; one-way ANOVA, F3,19 = 32.03, P = 0.000000048 vs. CS-US; CS-alone: 30.3 ± 1.3 neurons/area, n = 6, P = 0.00026 vs. CS-US; immediate shock (IS): 34.4 ± 3.6 neurons/area, n = 6, P = 0.0025 vs. CS-US; Fig. 3C,D). The probability of c-Fos expression in the PNN-PV− neurons was 63.0% ± 2.9% (home cage: 13.8% ± 3.5%, n = 6; one-way ANOVA, F3,19 = 19.90, P = 0.0000021 vs. CS-US; CS-alone: 44.7% ± 4.1%, n = 6, P = 0.049 vs. CS-US; IS: 39.7% ± 6.0%, n = 6, P = 0.0096 vs. CS-US; Fig. 3E). Interestingly, there was no significant difference in the probability of c-Fos expression in PNN-PV+ neurons between all the groups (one-way ANOVA, F3,19 = 2.66, P = 0.077; Fig. S3). More PNN-CaMKII+ neurons than CaMKII+ neurons without PNNs expressed c-Fos after training (WFA+CaMKII+: 61.4% ± 6.3%, n = 8; WFA-CaMKII+: 9.65% ± 1.2%, n = 8; Wilcoxon paired signed-rank test, P = 0.000931; Fig. S4). On the other hand, c-Fos expression did not differ between PV+ neurons with and without PNNs (WFA+PV+: 33.5% ± 5.4%, n = 6; WFA-PV+: 26.6% ± 2.0%, n = 6; Wilcoxon paired signed-rank test, P = 0.31; Fig. S4). Therefore, PNN-CaMKII+ neurons are more likely to be recruited during fear conditioning.
(A) The experimental schedule. (B) Tone-induced freezing in mice that received either tone only (blue line, n = 16) or tone-shock pairing (red line, n = 17). Error bars indicate the SEM. (C) Representative images showing WFA-labeling (red) and expression of PV (cyan) and c-Fos (green) in mice exposed to their home cage (HC; upper), CS-alone (middle upper), CS-US pairing (middle lower), or IS (lower). Yellow arrows indicate PNN-PV− neurons expressing c-Fos. White arrowheads indicate PNN-PV− neurons that do not express c-Fos. Scale bar, 20 μm. (D) Total number of neurons expressing c-Fos in mice subjected to HC (black, n = 6), CS-alone (blue, n = 6), paired CS-US (red, n = 5), and IS (gray, n = 6). Fear conditioning by CS-US pairing (****P < 0.0001 vs. HC), CS-alone training (***P < 0.001 vs. HC), and IS (****P < 0.0001 vs. HC) increases the number of neurons expressing c-Fos. (E) The probability of c-Fos expression in PNN-PV− neurons in mice subjected to HC, CS-alone, paired CS-US, and IS. Compared with animals in their HC and those exposed to CS-alone and IS, c-Fos is localized to PNN-PV− neurons in mice subjected to fear conditioning. Colors as in (D). Error bars indicate the SEM. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; one-way ANOVA with Tukey’s post hoc test.
Activation of PNN-expressing neurons correlates positively with fear memory
Twenty-four hours after fear conditioning, mice were re-exposed to the training cue. Fear memory recall induced c-Fos expression in PNN-PV− neurons (Fig. 4A). The probability of c-Fos expression in PNN-PV− neurons was higher in the CS-US group than in the CS-alone group (CS-US: 52.5% ± 3.7%, n = 12; CS-alone: 38.4% ± 6.0%, n = 14; W = 44, P = 0.042; Fig. 4B) and also in the unpaired conditioned group (paired CS-US: 41.3% ± 4.5%, n = 4 unpaired CS-US: 17.6% ± 4.3%, n = 5; W = 19, P = 0.037; Fig. S5). To exclude the possibility that expression of c-Fos in PNN-PV− neurons was due to activation of neighboring neurons, the number of PNN-PV− neurons expressing c-Fos was normalized to the total number of c-Fos+ neurons. The normalized probability of c-Fos expression in PNN-PV− neurons during memory retrieval was negatively correlated with the activation of neighboring neurons in the CS-US group (Fig. S6A), but showed a weak but positive correlation in the CS-alone group (Fig. S6B). These findings indicate that PNN-PV− neurons are activated after fear memory retrieval, regardless of whether surrounding neurons are activated. Importantly, the normalized probability of c-Fos expression in PNN-PV− neurons correlated positively with freezing within the CS-US group (Fig. 4C), and even within both the CS-US and CS-alone groups (Fig. S6C), suggesting that greater activation of PNN-PV− neurons is linked to increased fear memory.
(A) WFA-labeling (red) and expression of PV (cyan) and c-Fos (green) during the cued recall test in mice that received fear conditioning (upper) or CS-alone training (lower). Yellow arrows indicate PNN-PV− neurons expressing c-Fos. White arrowheads indicate PNN-PV− neurons that do not express c-Fos. Scale bar, 20 μm. (B) c-Fos is expressed in PNN-PV− neurons during fear recall in the CS-US group (red; n = 12) compared with the CS-alone group (blue; n = 14). Error bars indicate the SEM. W = 44, *P < 0.05; Wilcoxon paired signed-rank test. (C) The normalized probability of neurons expressing PNNs correlates positively with freezing during fear recall in the CS-US group (red; n = 12, P < 0.01).
Discussion
To date, researchers have mainly examined the role of PNNs in inhibitory neurons19 since PNNs were barely detectable in association with excitatory neurons19,20,21,22. However, we found that PNNs, as determined by aggrecan expression and WFA-labeling, were present mainly in association with excitatory neurons in brain areas related to emotional memory function, suggesting that PNNs modulate the function of excitatory neurons.
PNNs limit plasticity in the adult central nervous system2,23. However, this function seems to differ depending on the brain region and animal species. For example, PNN degradation allows the induction of long-term potentiation in the hippocampal CA2 area where plasticity is normally limited5, and, likewise, the enhancement of long-term depression in the perirhinal cortex8. By contrast, the digestion of PNNs leads to the impairment of long-term potentiation at thalamo-LA synapses7 and the absence of enhanced plasticity in the feline visual cortex24. Since the types of PNN-expressing neurons differ according to the brain region, cell type-specific degradation of PNNs could provide insight into how PNNs play a role in plasticity, including fear memory.
We showed here that excitatory neurons expressing PNNs are functionally different from those lacking PNNs. PNN-PV− excitatory neurons are activated during fear conditioning, although the precise mechanism remains unclear. WFA recognizes 4-O-sulfation on chondroitin sulfate chains25. The deposition of chondroitin 4-sulfate on neurons causes membrane depolarization, which may help neurons reach the threshold for spike firing26, thereby lowering the threshold of c-Fos expression. Alternatively, since the meshwork of PNNs surrounds synaptic contacts at thalamocortical boutons25, synaptic inputs from both the thalamus and cortex may converge on PNN-PV− neurons. Our data do not rule out PNN-PV− neuron activation due to sensory stimuli because exposure to tone or foot shock alone resulted in the upregulation of c-Fos in these neurons. However, PNN-PV− neuron activation was affected by the activity of neighboring neurons in the CS-alone group, but not in the CS-US group. Furthermore, the population of PNN-PV− neurons activated during retrieval was higher in the paired CS-US group than in the unpaired CS-US group. Hence, these neurons may encode CS-US associations. While almost nothing is known about the participation of PNN-PV− neurons in memory traces, our data suggest that a memory engram is distributed in a subpopulation of neurons, PNN+ neurons, in the LA.
Methods
Animals
Male C57BL/6J mice (aged 5–10 weeks; SLC, Hamamatsu, Japan) and GAD67-GFP knock-in mice12 were used in this study. The mice were housed in plastic cages under a 12 h light/dark cycle at 24 °C and had free access to water and food. All procedures were approved by the animal experiment ethics committee of Hoshi University and performed in accordance with the Hoshi University guidelines for the care and use of laboratory animals. All experimental procedures minimized the number and suffering of the animals.
Immunohistochemistry
Mice were anesthetized using intraperitoneal urethane (1.25 g/kg) and transcardially perfused with phosphate-buffered saline (PBS; pH 7.4) followed by 4% paraformaldehyde in PBS. The brains were post-fixed overnight at 4 °C in the same fixative solution, transferred to 30% sucrose in PBS for 48 h, and then cut into 30 μm coronal sections using a cryostat (CM1510-11, Leica Co., Wetzlar, Germany). The free-floating sections were blocked with 5% bovine serum albumin (BSA) and 0.3% Triton X-100 in PBS for 1 h at room temperature and incubated overnight at 4 °C with the following antibodies: mouse monoclonal anti-parvalbumin (1:5,000; P3088, Sigma-Aldrich, St. Louis, MO, USA), rabbit polyclonal anti-parvalbumin (1:3,000; PV-Rb-Af750, Frontier Institute Co., Ltd. Hokkaido, Japan), mouse monoclonal anti-CaMKIIα (1:400, Cat. no. 05-532, clone 6G9, Merck Millipore), rabbit polyclonal anti-neurogranin (1:1,000; AB5620, Merck Millipore), rabbit polyclonal anti-GABA (1:4,000; A2052, Sigma-Aldrich), rabbit polyclonal anti-aggrecan (1:1,000; AB1031, Merck Millipore), and rabbit polyclonal anti-c-Fos (1:5,000; Cat. no. 226 003, Synaptic Systems, Goettingen, Germany). The following secondary antibodies used in the study were all purchased from Thermo Fisher Scientific (Waltham, MA, USA): Alexa 488-conjugated donkey anti-mouse IgG (1:1,000; A21202), Alexa 488-conjugated donkey anti-rabbit IgG (1:1,000; A21206), Alexa 594-conjugated donkey anti-rabbit IgG (1:1,000; A21207), Alexa 647-conjugated donkey anti-mouse IgG (1:1,000; A31571), and Alexa 647-conjugated donkey anti-rabbit IgG (1:1,000; A31573). To visualize PNNs, sections were incubated overnight at 4 °C with biotin-conjugated WFA (1:3,000; BA-3101, EY laboratories, Inc., San Mateo, CA, USA), followed by Alexa 594-conjugated streptavidin (0.67 μg/ml; S11227, Thermo Fisher Scientific). Labeled sections were mounted onto microscope slides with Prolong Diamond Antifade Reagent containing DAPI (Thermo Fisher Scientific).
Analysis of distance to the nearest neighbor
The mean distance  from each individual to its nearest neighbor in the LA and BA is represented as
from each individual to its nearest neighbor in the LA and BA is represented as  , where n and ri denote the total number of PNN-PV− neurons in each region and the distance of each individual to its nearest neighbor, respectively. If this population is randomly distributed, the expected mean distance
, where n and ri denote the total number of PNN-PV− neurons in each region and the distance of each individual to its nearest neighbor, respectively. If this population is randomly distributed, the expected mean distance  can be formulated by the following equation: where S is the area of the LA or BA27.
can be formulated by the following equation: where S is the area of the LA or BA27.
Behavioral procedures
The day before fear conditioning, male C57BL/6J mice were allowed 10 min to freely explore the conditioning chamber (box A), which consisted of a transparent plastic box with a stainless steel grid floor (16 × 14 × 12 cm). For training, mice were placed in box A, and, after 2 min, they were presented with five pairings of a 30 s white-noise CS (65 dB, 1 Hz) that co-terminated with a 1 s foot shock US (0.3 mA) at a variable interval of 20–120 s. Mice in the unpaired group received explicitly unpaired training with the same number and specifications of CS and US stimuli. Thirty seconds after the final CS-US for the paired group or the final CS for the unpaired group, mice were returned to their home cage. The CS-alone group was exposed to the same protocol without the US. Mice in the IS group were given three consecutive 1 s foot shocks (0.3 mA) at an interval of 1 s immediately after placement in the conditioning chamber and quickly returned to their home cage. The next day, mice were transported to a novel chamber (box B), which consisted of a white plastic box (17 × 10 × 10 cm), and tested for freezing behavior when re-exposed to the CS. All sessions were video-recorded for automatic scoring of freezing (TimeFZ software, O’HARA & Co., LTD., Tokyo, Japan), which was defined as a period of immobility, except for respiratory-related movements, lasting for at least 2 s.
Analysis of c-Fos
Mice were euthanized for immunohistochemical analysis 90 min after completing the behavioral task. The c-Fos-immunostained sections from −1.6 mm to −2.0 mm posterior to bregma were imaged at magnification ×20 and ×40 using an FV1200 confocal laser scanning microscope (Olympus, Tokyo, Japan). The c-Fos+ neurons in the ventrolateral subdivision of the LA were counted bilaterally using three slices from each mouse. The total number of c-Fos neurons in the LA was determined in an area of 59,862 ± 1,814 μm3 using ImageJ software (version 1.48; National Institutes of Health, Bethesda, MD, USA) and averaged within each mouse. The background fluorescence cut-off for all images was equal. The percentage of c-Fos+ neurons in PNN-PV− neurons was calculated as the percentage of c-Fos+ neurons per total PNN-PV− neurons in the LA, and was also normalized by the total number of c-Fos+ neurons in the same region.
Statistical analysis
The results are expressed as the mean ± SEM. Statistical comparisons were analyzed with Student’s t-test (paired data, normal distribution), Wilcoxon paired signed-rank test (non-normal distribution), or one-way ANOVA followed by Tukey’s post hoc test (comparisons of multiple groups). Correlations were calculated using a linear correlation. No statistical methods were used to predetermine sample sizes, but the sample sizes used were similar to those generally reported in the field for similar experiments9,17. The criterion for statistical significance was set at P < 0.05.
Additional Information
How to cite this article: Morikawa, S. et al. Activation of perineuronal net-expressing excitatory neurons during associative memory encoding and retrieval. Sci. Rep. 7, 46024; doi: 10.1038/srep46024 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Bitanihirwe, B. K. & Woo, T. U. Perineuronal nets and schizophrenia: the importance of neuronal coatings. Neurosci. Biobehav. Rev. 45, 85–99 (2014).
Wang, D. & Fawcett, J. The perineuronal net and the control of CNS plasticity. Cell. Tissue Res. 349, 147–160 (2012).
Hartig, W., Brauer, K. & Bruckner, G. Wisteria floribunda agglutinin-labelled nets surround parvalbumin-containing neurons. Neuroreport 3, 869–872 (1992).
Seeger, G., Brauer, K., Hartig, W. & Bruckner, G. Mapping of perineuronal nets in the rat brain stained by colloidal iron hydroxide histochemistry and lectin cytochemistry. Neuroscience 58, 371–388 (1994).
Carstens, K. E., Phillips, M. L., Pozzo-Miller, L., Weinberg, R. J. & Dudek, S. M. Perineuronal Nets Suppress Plasticity of Excitatory Synapses on CA2 Pyramidal Neurons. J. Neurosci. 36, 6312–6320 (2016).
Pizzorusso, T. et al. Reactivation of ocular dominance plasticity in the adult visual cortex. Science 298, 1248–1251 (2002).
Gogolla, N., Caroni, P., Luthi, A. & Herry, C. Perineuronal nets protect fear memories from erasure. Science 325, 1258–1261 (2009).
Romberg, C. et al. Depletion of perineuronal nets enhances recognition memory and long-term depression in the perirhinal cortex. J. Neurosci. 33, 7057–7065 (2013).
Reijmers, L. G., Perkins, B. L., Matsuo, N. & Mayford, M. Localization of a stable neural correlate of associative memory. Science 317, 1230–1233 (2007).
LeDoux, J. E. Emotion circuits in the brain. Annu. Rev. Neurosci. 23, 155–184 (2000).
Bruckner, G. et al. Perineuronal nets provide a polyanionic, glia-associated form of microenvironment around certain neurons in many parts of the rat brain. Glia 8, 183–200 (1993).
Tamamaki, N. et al. Green fluorescent protein expression and colocalization with calretinin, parvalbumin, and somatostatin in the GAD67-GFP knock-in mouse. J. Comp. Neurol. 467, 60–79 (2003).
Mulligan, K. A., van Brederode, J. F. & Hendrickson, A. E. The lectin Vicia villosa labels a distinct subset of GABAergic cells in macaque visual cortex. Vis. Neurosci. 2, 63–72 (1989).
LaBar, K. S. & Cabeza, R. Cognitive neuroscience of emotional memory. Nat. Rev. Neurosci. 7, 54–64 (2006).
Singec, I., Knoth, R., Ditter, M., Volk, B. & Frotscher, M. Neurogranin is expressed by principal cells but not interneurons in the rodent and monkey neocortex and hippocampus. J. Comp. Neurol. 479, 30–42 (2004).
Bruckner, G. et al. Cortical areas are revealed by distribution patterns of proteoglycan components and parvalbumin in the Mongolian gerbil and rat. Brain. Res. 658, 67–86 (1994).
Yamada, J., Ohgomori, T. & Jinno, S. Perineuronal nets affect parvalbumin expression in GABAergic neurons of the mouse hippocampus. Eur. J. Neurosci. 41, 368–378 (2015).
Giamanco, K. A., Morawski, M. & Matthews, R. T. Perineuronal net formation and structure in aggrecan knockout mice. Neuroscience 170, 1314–1327 (2010).
Berretta, S., Pantazopoulos, H., Markota, M., Brown, C. & Batzianouli, E. T. Losing the sugar coating: potential impact of perineuronal net abnormalities on interneurons in schizophrenia. Schizophr. Res. 167, 18–27 (2015).
Alpar, A., Gartner, U., Hartig, W. & Bruckner, G. Distribution of pyramidal cells associated with perineuronal nets in the neocortex of rat. Brain. Res. 1120, 13–22 (2006).
Hendry, S. H., Jones, E. G., Hockfield, S. & McKay, R. D. Neuronal populations stained with the monoclonal antibody Cat-301 in the mammalian cerebral cortex and thalamus. J. Neurosci. 8, 518–542 (1988).
Seeger, G., Luth, H. J., Winkelmann, E. & Brauer, K. Distribution patterns of Wisteria floribunda agglutinin binding sites and parvalbumin-immunoreactive neurons in the human visual cortex: a double-labelling study. J. Hirnforsch. 37, 351–366 (1996).
McRae, P. A. & Porter, B. E. The perineuronal net component of the extracellular matrix in plasticity and epilepsy. Neurochem. Int. 61, 963–972 (2012).
Vorobyov, V., Kwok, J. C., Fawcett, J. W. & Sengpiel, F. Effects of digesting chondroitin sulfate proteoglycans on plasticity in cat primary visual cortex. J. Neurosci. 33, 234–243 (2013).
Miyata, S., Komatsu, Y., Yoshimura, Y., Taya, C. & Kitagawa, H. Persistent cortical plasticity by upregulation of chondroitin 6-sulfation. Nat. Neurosci. 15, 414–422, S411-412 (2012).
Vigetti, D. et al. Chondroitin sulfates act as extracellular gating modifiers on voltage-dependent ion channels. Cell. Physiol. Biochem. 22, 137–146 (2008).
Clark, P. J. & Evans, F. C. Ecology 35, 445–453 (1954).
Acknowledgements
The GAD67 knock-in mice were provided by Dr. Yanagawa (Gunma University, Maebashi, Japan). The research was supported by the Lotte Shigemitsu Prize and by JSPS KAKENHI (Grant Number: JP15K06784).
Author information
Authors and Affiliations
Contributions
S.M. performed the experiments; S.M. and H.T. contributed to the figures and analyzed the data; Y.I. and H.T. designed the experiments; Y.I., M.N., and H.T. wrote the manuscript. All authors discussed the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Morikawa, S., Ikegaya, Y., Narita, M. et al. Activation of perineuronal net-expressing excitatory neurons during associative memory encoding and retrieval. Sci Rep 7, 46024 (2017). https://doi.org/10.1038/srep46024
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep46024
This article is cited by
-
Acan downregulation in parvalbumin GABAergic cells reduces spontaneous recovery of fear memories
Molecular Psychiatry (2023)
-
Zinc-a2-Glycoprotein Acts as a Component of PNN to Protect Hippocampal Neurons from Apoptosis
Molecular Neurobiology (2023)
-
Child abuse associates with increased recruitment of perineuronal nets in the ventromedial prefrontal cortex: a possible implication of oligodendrocyte progenitor cells
Molecular Psychiatry (2022)
-
The extracellular matrix and perineuronal nets in memory
Molecular Psychiatry (2022)
-
Differential regulation of alcohol consumption and reward by the transcriptional cofactor LMO4
Molecular Psychiatry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.