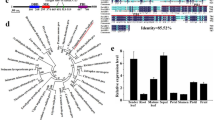
Abstract
Recent research revealed that TERMINAL FLOWER 1 (TFL1) homologues are involved in the critical developmental process of floral initiation in several plant species. In this study, the functions of three putative TFL1 homologues (JcTFL1a, JcTFL1b and JcTFL1c) in the biofuel plant Jatropha curcas were analysed using the transgenic approach. JcTFL1b and JcTFL1c, but not JcTFL1a, could complement the TFL1 function and rescue early flowering and determinate inflorescence phenotype in tfl1-14 Arabidopsis mutant, thus suggesting that JcTFL1b and JcTFL1c may be homologues of TFL1. Transgenic Jatropha overexpressing JcTFL1a, JcTFL1b or JcTFL1c showed late flowering, whereas only JcTFL1b and JcTFL1c overexpression delayed flowering in transgenic Arabidopsis. JcTFL1b-RNAi transgenic Jatropha consistently exhibited moderately early flowering phenotype. JcFT and JcAP1 were significantly downregulated in transgenic Jatropha overexpressing JcTFL1a, JcTFL1b or JcTFL1c, which suggested that the late flowering phenotype of these transgenic Jatropha may result from the repressed expression of JcFT and JcAP1. Our results indicate that these three JcTFL1 genes play redundant roles in repressing flowering in Jatropha.
Similar content being viewed by others
Introduction
Floral initiation, a key developmental process in higher plant life, involves transition from vegetative to reproductive growth. It is controlled by genetic pathways that integrate environmental cues such as temperature and day length as well as endogenous signals such as hormones, regulation of genes and the developmental status of the plant1,2.
The proper timing of flowering is the most critical aspect to ensure reproductive success3. In the model plant Arabidopsis thaliana, complex genetic networks for flowering are mainly regulated by five major pathways: the photoperiod, vernalisation, gibberellin, autonomous and age pathways4,5,6. In Arabidopsis, the floral induction signals from these five major flowering pathways are transmitted via floral integrator genes, such as FLOWERING LOCUS T (FT), SUPPRESSOR OF OVEREXPRESSION OF CONSTANS1 (SOC1) and FLOWERING LOCUS C (FLC), to the floral meristem identity genes LEAFY (LFY) and APETALA1 (AP1) at the apical meristem6,7,8,9.
FT belongs to the FT/TFL1 gene family, which is similar to the phosphatidyl ethanolamine-binding protein (PEBP) gene family2,7,10. The FT/TFL1 gene family includes three major subfamilies—FT-like, TERMINAL FLOWER 1 (TFL1)-like and MOTHER OF FT AND TFL1 (MFT)-like—in the plant2,10. In Arabidopsis, FT and TFL1 act as key flowering regulators playing antagonistic roles in the flowering transition2,11,12.
FT promotes flowering initiation, whereas TFL1 acts as a repressor for floral initiation and maintains the inflorescence meristem through suppression of the expression of LFY and AP112,13,14,15. The Arabidopsis tfl1 mutants produce fewer leaves and flower earlier than wild-type (WT) plants and the inflorescence meristem converts to a terminal flower12. In contrast, overexpression of TFL1 genes from Arabidopsis and other plant species results in a lengthening of the vegetative phase, increased secondary inflorescence production and a delayed flowering in transgenic Arabidopsis16,17,18,19,20.
Study of TFL1 homologues in various plant species has revealed similar as well as distinctive functions. For instance, constitutive expression of CsTFL1 in chrysanthemums resulted in extremely late flowering and prevented upregulation of floral meristem identity genes in shoot tips and leaves3. Ectopic expression of the maize TFL1 homologues produced similar phenotypes, including delayed flowering and altered inflorescence architecture21. Moreover, the overexpression of two TFL1-like genes, CorfloTFL1 and CorcanTFL1, cloned from Cornus florida and C. canadensis, extended vegetative growth and delayed flowering in transgenic Arabidopsis, indicating functional conservation of TFL1 homologues in control of transition to flowering22.
Overexpression of a homologue of TFL1 (StTFL1) in transgenic potato resulted in a marked increase in tuber number in the transgenic plants compared to the wild-type plants, which suggested that StTFL1 was involved in the regulation of tuberisation23. A 2-bp deletion in the coding region of theTFL1 homologue was responsible for continuous flowering behaviour in woodland strawberry24. The transgenic apples and pears expressing antisense transcripts of apple TFL1 homologue (MdTFL1) displayed early flowering phenotypes and extreme reduction of the juvenile phase25. These studies therefore suggest that TFL1 homologues have unique features in flowering time and plant architecture in each plant species18,26.
Jatropha curcas (physic nut), which belongs to the Euphorbiaceae family, is widely recognised as a potential bioenergy crop in tropical and subtropical regions due to its high oil content seeds and its adaptability to marginal land27,28. Indeed, the oil content of Jatropha seeds ranges from 30 to 50% by weight, and its oil exhibits excellent physicochemical properties such as low acidity, good stability, low viscosity and good cold flow properties. Therefore it is suitable for biodiesel production and industrial applications29,30. However, unstable and poor flowering causes low seed yield in Jatropha31. Jatropha shows diverse flowering behaviour in different cultivation areas. Normally, Jatropha flowers during the rainy season (one or two flowering peaks), but in permanently humid regions, flowering occurs throughout the year32. Also, the harvest of Jatropha seeds is time-consuming, and the harvest efficiency is low because of its continuous flowering33.
Molecular breeding will be an effective genetic improvement method to modify the flowering behaviour of Jatropha to obtain high-yielding Jatropha cultivars. The function analysis of JcTFL1 genes is required for the comprehensive understanding of the molecular mechanism of flowering in Jatropha, which would be helpful for the genetic modification and improvement of Jatropha with optimal flowering behaviour.
We previously isolated six members of the FT/TFL1 gene family and analysed their expression patterns throughout the vegetative and reproductive developmental stages in Jatropha. Three TFL1 homologues were identified and named JcTFL1a, JcTFL1b and JcTFL1c, respectively34. Also, we analysed the function of an FT orthologue (JcFT) in Jatropha and found that JcFT acts as a flowering promoter35.
Chua et al.36 isolated two Jatropha TFL1 homologues, JcTFL1-1 and JcTFL1-2 corresponding to JcTFL1a and JcTFL1b and preliminarily analysed their functions by overexpressing them in transgenic Arabidopsis and Jatropha. In their research, overexpression of JcTFL1-1 or JcTFL1-2 resulted in earlier flowering in transgenic Arabidopsis, which was different from our findings. JcTFL1-1 overexpression Jatropha showed early flowering and self-pruning phenotypes, and JcTFL1-2 overexpression Jatropha showed multiple shoots phenotype36, which were not found in our transgenic Jatropha.
To elucidate the biological functions of three JcTFL1 genes in the floral initiation of Jatropha using a transgenic approach, we overexpressed JcTFL1 genes in transgenic Arabidopsis and Jatropha, and downregulated JcTFL1b in transgenic Jatropha in this study. Our results show JcTFL1 genes play important roles in the regulation of flowering behaviour in Jatropha, which would be helpful to improve seed yield of Jatropha.
Results
Function analysis of JcTFL1 genes in transgenic Arabidopsis
To determine the effects of JcTFL1 genes on flowering time and inflorescence architecture, JcTFL1a, JcTFL1b and JcTFL1c cDNA driven by the constitutive 35S promoter were transformed into WT Arabidopsis plants, respectively. Consequently, we obtained >30 independent transgenic lines for 35S::JcTFL1a, 35S::JcTFL1b and 35S::JcTFL1c, respectively. For most 35S::JcTFL1b and 35S::JcTFL1c transgenic Arabidopsis lines, bolting occurred significantly later than in WT under long-day (LD) conditions (Fig. 1A,B). On the other hand, all transformants with 35S::JcTFL1a normally bolted like WT plants (Fig. 1A,B). Also, 35S::JcTFL1b and 35S::JcTFL1c transgenic plants had an increased rosette leaf number (Fig. 1A,B); in contrast, 35S::JcTFL1a transgenic Arabidopsis plants produced approximately the same rosette leaf number as WT plants (Fig. 1A,B). We also analysed expression levels of some flowering-related genes in WT and transgenic Arabidopsis by quantitative reverse transcriptase-polymerase chain reaction (qRT-PCR) experiment (Supplementary Fig. S2). Expression levels of all the flowering-related genes that we analysed was not significantly affected in 35S::JcTFL1a transgenic Arabidopsis lines (Supplementary Fig. S2D,H). On the other hand, AtAP1 was significantly downregulated in the 35S::JcTFL1b and 35S::JcTFL1c transgenic lines (Supplementary Fig. S2E), which was consistent with the late flowering phenotype. However, the expression levels of flowering-related genes AtLFY and AtSOC1 were not significantly affected in the transgenic lines (Supplementary Fig. S2F,G), which was not in accordance with the findings that AtTFL1 repressed the expression of AtLFY12,14. We also analysed the expression levels of AtTFL1and AtFT, and unexpectedly found that the expression of AtTFL1 and AtFT were downregulated in the 35S::JcTFL1b and 35S::JcTFL1c transgenic lines (Supplementary Fig. S2D,H).
(A) Growth of wild-type and JcTFL1 overexpression transgenic lines in the Col-0 background under long-day (LD) conditions 40 days after germination. (B) Days and leaves to bolting for transgenic Arabidopsis lines in the Col-0 background and wild-type plants grown under LD conditions. Bar represents 5 cm. Values are means ± SD of the results from ten plants of each line. Student’s t-test was used to determine significant differences between the transgenic and WT plants. Significance level: **P < 0.01.
We also found that more cauline branches were produced in the severe transgenic plants overexpressing JcTFL1b and JcTFL1c during the late development stage compared to WT plants (Fig. 2A–E), and 35S::JcTFL1b and 35S::JcTFL1c transgenic plants showed abnormality in the development of inflorescence, flowers and siliques. The shoot tips of severe transformants with 35S::JcTFL1b and 35S::JcTFL1c could not produce normal flower buds like WT plants, but instead a flower bud-like structure at the same position (Fig. 2F,G,I) which was similar to the phenotype overexpressing apple TFL1 homologue in transgenic Arabidopsis18. Sometimes the tip consisted of small compact clusters of flower buds surrounded by leaves (Fig. 2H,J), and most of the flower buds could not develop into normal siliques. In addition, some unexpected phenotypes were observed in some of the transgenic Arabidopsis lines with 35S::JcTFL1c where a new inflorescence emerged from the position that should be pistil in WT plants (Fig. 2K,L,M), and the top portion of siliques in some 35S::JcTFL1c plants became abnormal (Fig. 2N,O,P). These data indicated that JcTFL1b and JcTFL1c might function as flowering repressors and JcTFL1a did not affect flowering time in transgenic Arabidopsis.
Whole plant (A), inflorescence (F), flowers (K) and siliques (N) of wild-type Arabidopsis; whole plant (B and C) and inflorescence buds (G and H) of 35S::JcTFL1b transgenic Arabidopsis; whole plant (D and E), inflorescence buds (I and J), inflorescences (L and M) and siliques (O and P) of 35S::JcTFL1c transgenic Arabidopsis. Bars in A–E represent 5 cm, and bars in F–P represent 1 mm.
To investigate whether JcTFL1 homologues function equivalently to Arabidopsis TFL1, we introduced the 35S::JcTFL1a, 35S::JcTFL1b and 35S::JcTFL1c vector into Arabidopsis tfl1-14 mutant plants, which showed early flowering and determinate inflorescence phenotypes under LD conditions.
Consistent with the results overexpressing JcTFL1 genes in WT Arabidopsis background, tfl1-14 mutant plants transformed with 35S::JcTFL1b and 35S::JcTFL1c rescued the early flowering phenotype and maintained their indeterminate flowering behaviour similar to WT Arabidopsis plants (Fig. 3A–D,F; Supplementary Fig. S3). Moreover, some severe 35S::JcTFL1b and 35S::JcTFL1c transformants of tfl1-14 background exhibited a similar phenotype in the shoot tip to these severe 35S::JcTFL1b and 35S::JcTFL1c transformants of WT background (Figs 2G–J,3E,3G).
(A) Growth of mutant tfl1-14 and JcTFL1 overexpression transgenic lines in the tfl1-14 background under long-day (LD) conditions 23 days after germination. (B) Days and leaves to bolting for transgenic Arabidopsis lines in the tfl1-14 background and mutant tfl1-14 plants grown under LD conditions. (C) Inflorescence of mutant tfl1-14. (D and E) Inflorescences of transgenic Arabidopsis overexpressing JcTFL1b. (F and G) Inflorescences of transgenic Arabidopsis overexpressing JcTFL1c. Values are means ± SD of the results from ten plants of each line. Student’s t-test was used to determine significant differences between the transgenic and tfl1-14 mutant plants. Significance level: **P < 0.01. Bar in A represents 5 cm, and bars in C-G represent 1 mm.
On the other hand, overexpressing JcTFL1a could not rescue the early flowering and determinate inflorescence phenotype of tfl1-14 mutant plants (Fig. 3A,B). These results supported the theory that JcTFL1b and JcTFL1c might be the orthologues of TFL1 in Jatropha and that they affected flowering time and the inflorescence architecture in Arabidopsis.
Taken together, these findings demonstrated that ectopic expression of JcTFL1b and JcTFL1c in Arabidopsis delayed flowering and caused the changes in inflorescence architecture, while overexpression of JcTFL1a did not affect the flowering time and inflorescence architecture in transgenic Arabidopsis.
Overexpression of JcTFL1 genes affected flowering time and plant morphology in Jatropha
To determine the effects of JcTFL1 genes on flowering time and plant morphology in Jatropha, we generated transgenic Jatropha plants with JcTFL1a, JcTFL1b and JcTFL1c under the control of the 35S promoter, respectively. Consistent with the results overexpressing JcTFL1b and JcTFL1c in Arabidopsis, transgenic Jatropha with 35S::JcTFL1b and 35S::JcTFL1c showed the extremely late flowering phenotype.
WT Jatropha flowered about 9 months after planting, and its height in the first flowering stage was about 1.5 m (Fig. 4D,H), while the 35S::JcTFL1b and 35S::JcTFL1c transgenic Jatropha flowered about 1.5 years after planting and reached a height of about 2.0 m. When we wrote this paper (3 years after planting), the extremely severe phenotype of 35S::JcTFL1b and 35S::JcTFL1c transgenic Jatropha had still not flowered (Fig. 4B,C,F,G).
Transgenic Jatropha overexpressing JcTFL1a (A), JcTFL1b (B) and JcTFL1c (C) have not flowered yet 3 years after plantation in the field. Wild-type (WT) Jatropha (D) has set fruits about 10 months after plantation in the field; E, F and G correspond to the developmental stages of shoot apical of transgenic Jatropha shown in (A), (B) and (C), respectively. (H) The fruits of WT Jatropha shown in (D). The red circle indicates fruits. Bars represent 20 cm.
Unexpectedly, 35S::JcTFL1a transgenic Jatropha also exhibited late flowering phenotype, which did not agree with the result in Arabidopsis—and the severest phenotype was similar to the 35S::JcTFL1b and 35S::JcTFL1c transgenic Jatropha, which had not flowered after 3 years (Fig. 4A,E). The heights of these transgenic plants are shown in Supplementary Fig. S4. Because of the no-flowering behaviour in these severe transgenic Jatropha, they grew no branches in the first year after transplanting to the soil (Supplementary Fig. S5). Meanwhile, WT Jatropha usually branched in the position where they flowered in the first year (Fig. 4D).
To determine whether JcTFL1 gene overexpression in the transgenic Jatropha changed the expression of some flowering-related genes—such as AP1, LFY, FT and SOC1 homologues in Jatropha—qRT-PCR analysis was performed by using the shoot tip of the transgenic and WT Jatropha 2 years later after planting to soil (Fig. 5). Overexpression of each of three JcTFL1 genes did not affect the transcripts of the other two JcTFL1 genes (Fig. 5A–C).
(A) to (G) are expression levels of JcTFL1a, JcTFL1b, JcTFL1c, JcAP1, JcLFY, JcSOC1and JcFT, respectively. The qRT-PCR results were obtained from two independent biological replicates with three technical replicates each. Levels of the detected amplicons were normalized using the amplified products of JcActin.
As expected, JcAP1 was significantly downregulated in three transgenic lines (Fig. 5D), which was consistent with the findings that TFL1 represses the expression of AP1 in Arabidopsis37. However, the expression of JcLFY and JcSOC1 was not apparently affected in these transgenic Jatropha (Fig. 5E,F), whereas AtTFL1 repressed the expression levels of AtLFY in Arabidopsis12,14,15.
Meanwhile, we also detected the expression level of the florigen gene JcFT in transgenic Jatropha and found JcFT was significantly downregulated in these JcTFL1 overexpressed transgenic Jatropha (Fig. 5G), which suggested that JcTFL1 might repress the expression of JcFT.
Silencing of the JcTFL1b gene moderately accelerated flowering in transgenic Jatropha
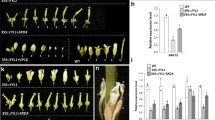
We previously found that among the three genes (JcTFL1a, JcTFL1b, and JcTFL1c) only JcTFL1b transcripts are abundantly accumulated in the flower buds of mature plants34. At the beginning of this study, we found that overexpression of JcTFL1b or JcTFL1c, but not JcTFL1a, delayed the flowering time in transgenic Arabidopsis plants (Fig. 1) and rescued the early flowering phenotype of tfl1-14 mutant plants (Fig. 3). Therefore, we first chose JcTFL1b for further functional analysis by RNA interference (RNAi) in this study. We generated transgenic Jatropha containing the JcTFL1b-RNAi construct. These transgenic plants showed moderately early flowering, and produced flowers about 6 months later after planting to soil, which was 3 months earlier than that of WT Jatropha (Fig. 6E). The height of the first flowering JcTFL1b RNAi Jatropha was about 0.8 m (Fig. 6A,C), and the WT Jatropha remained as vegetative growth at the same time (Fig. 6B,D). In addition, we also detected the expression levels of some flowering-related genes in JcTFL1b RNAi Jatropha and found JcAP1 was remarkably upregulated (Fig. 6F). These findings further demonstrated that JcTFL1b acts as a flowering repressor in Jatropha.
The whole plant (A) and flower buds (C) of JcTFL1b RNAi transgenic Jatropha. The whole plant (B) and leaf buds (D) of WT Jatropha. The photos were imaged about 6 months later after transplanting to soil. Bars represent 10 cm. (E) Flowering time of JcTFL1b RNAi transgenic Jatropha. Flowering time was scored by the number of days from transplantation to soil to the day of first inflorescence emergence. (F) Quantitative RT-PCR analysis of several flowering-related genes in wild-type and JcTFL1b RNAi transgenic Jatropha. The qRT-PCR results were obtained from two independent biological replicates with three technical replicates each. Levels of the detected amplicons were normalized using the amplified products of JcActin.
Discussion
FT and TFL1 are thought to be molecular switches for vegetative growth to reproductive development in Arabidopsis. Moreover, FT promotes flowering and TFL1 represses flowering2,6,22.
In the present study, we analysed the function of three JcTFL1 genes using the transgenic approach. We found that JcTFL1b and JcTFL1c could delay flowering in transgenic Arabidopsis and Jatropha and that both of them could also rescue early flowering and determinate inflorescence phenotype of Arabidopsis tfl1-14 mutant (Figs 1,3,4; Supplementary Fig. S3). However, while JcTFL1a did not affect the flowering time in transgenic Arabidopsis in both WT and tfl1-14 background, it could delay flowering in Jatropha (Figs 1,3,4), which might be the results of ectopic expression in Arabidopsis. In addition, JcTFL1b RNAi Jatropha showed moderately early flowering phenotype (Fig. 6), which was similar to MdTFL1 RNAi apple that displayed early flowering phenotypes and extreme reduction of the juvenile phase25.
These findings suggested that three JcTFL1 genes act as flowering repressors in Jatropha and that they might act redundantly in repressing flowering time. Chua et al. also analysed the functions of JcTFL1-1and JcTFL1-2 corresponded to our JcTFL1a and JcTFL1b, respectively36. They found early flowering phenotypes in transgenic Arabidopsis plants overexpressing JcTFL1-1 and JcTFL1-2, early flowering and self-pruning phenotypes in JcTFL1-1 overexpression Jatropha and multiple shoots phenotype in JcTFL1-2 overexpression Jatropha36. We did not find these phenotypes in our transgenic Arabidopsis and Jatropha. Chua et al. considered that JcTFL1-1and JcTFL1-2 also functioned as flowering activator or florigen like JcFT in Jatropha, which was totally opposite to our results. The function analysis of the TFL1 homologues in apple18, chrysanthemum3, Cornus florida and C. canadensis22, gentian38, orchid39 and other plant species demonstrated that TFL1 homologues act as flowering repressors, which is consistent with our results.
The JcTFL1 genes also affected the plant morphology in Arabidopsis and Jatropha. Transgenic Arabidopsis plants overexpressing JcTFL1b and JcTFL1c showed more cauline branches phenotype (Fig. 2A,E), but transgenic Jatropha plants overexpressing each of three JcTFL1 genes showed no branches in the first year after planting (Supplementary Fig. S5), which may be the result of different plant species that generated new branches through different mechanisms. Transgenic Arabidopsis plants overexpressing JcTFL1c exhibited abnormal flowers and siliques (Fig. 2L–M,O–P), which suggested that JcTFL1c might have multiple functions in plant development.
To better understand the roles of JcTFL1 genes, we detected the expression levels of some flowering-related genes in transgenic Arabidopsis and Jatropha, and found that the expression patterns of the flowering-related genes in transgenic Arabidopsis exhibiting late flowering were similar to that of transgenic Jatropha. AtAP1and JcAP1 were significantly downregulated in overexpression transgenic Arabidopsis (Supplementary Fig. S2E) and Jatropha (Fig. 5D), which was consistent with the result that AtTFL1 represses the expression levels of AtAP1 in Arabidopsis3,37. At the same time, JcAP1 was remarkably upregulated in JcTFL1b RNAi transgenic Jatropha (Fig. 6F), which further demonstrated that JcTFL1b functioned as a flowering inhibitor in Jatropha. The expression levels of AtLFY were not affected in the transgenic Arabidopsis (Supplementary Fig. S2F), and the expression levels of JcLFY gene, the homologue of LFY in Jatropha, were also not obviously affected in transgenic Jatropha overexpressing JcTFL1a, JcTFL1b, or JcTFL1c and JcTFL1b-RNAi transgenic Jatropha (Figs 5E,6F), which was not agreed with the finding that TFL1 also repressed the expression levels of LFY in Arabidopsis37. In addition, the florigen gene AtFT and JcFT was significantly downregulated in transgenic Arabidopsis exhibiting late flowering phenotype (Supplementary Fig. S2H) and Jatropha overexpressing JcTFL1a, JcTFL1b, or JcTFL1c (Fig. 5G). Therefore, delayed flowering caused by overexpressing JcTFL1 in transgenic Arabidopsis and Jatropha might result from the repressed expression of AtAP1 and AtFT, JcAP1 and JcFT, respectively.
Materials and Methods
Plant materials and growth conditions
Mature Jatropha seeds were collected from Xishuangbanna Tropical Botanical Garden of the Chinese Academy of Sciences, Mengla County, Yunnan Province, China. All control and transgenic Jatropha were also grown in Mengla County. WT Arabidopsis thaliana ecotype Columbia (Col-0), the tfl1-14 mutant, and the transgenic lines were grown in peat soil in plant growth chambers at 22 ± 2 °C under a 16/8 h (light/dark) photoperiod, with cool-white fluorescent lamps used for lighting.
Transgenic Arabidopsis in the T2 or T3 generation were selected to examine flowering time and other phenotypes. For each genotype, 10 plants were used for characterisation; the number of leaves was counted along with the number of days between sowing and when the first flower bud was visible. All tissues prepared for qRT-PCR were immediately frozen in liquid nitrogen (N2) and stored at −80 °C until needed.
Plant expression vector construction and Arabidopsis and Jatropha transformation
Total RNA was extracted from the shoot tip of Jatropha using the protocol described by Ding et al.40 First-strand cDNA was synthesised using M-MLV-reverse transcriptase from TAKARA (Dalian, China) according to the manufacturer’s instructions. The full-length JcTFL1a, JcTFL1b and JcTFL1c cDNA were obtained by PCR using the primer pairs JcTFL1a F and JcTFL1a R, JcTFL1b F and JcTFL1b R and JcTFL1c F and JcTFL1c R, which introduced BamH I and Sac I recognition sites, respectively. The PCR products were subsequently cloned into the pMD19-T and sequenced. Primers used in this study are listed in Supplementary Table S1.
To construct the plant overexpression vector 35S::JcTFL1a, the JcTFL1a sequence was excised from the pMD19-T simple vector using the restriction enzymes BamH I and Sac I and then cloned into the pBI121 vector containing the cauliflower mosaic virus 35S (35S) promoter. Plant overexpression vector 35S::JcTFL1b and 35S::JcTFL1c were also constructed following the steps described above. A JcTFL1b RNA interference (RNAi) vector was constructed for silencing the JcTFL1b transcripts. A 123-bp coding region of the JcTFL1b cDNA was selected and amplified by the PCR using the primers listed in Supplementary Table S1. The PCR products were cloned into a pJL10 vector41 in opposing orientations on either side of a pdk intron to create an invert repeat driven by the 35S promoter. A schematic representation of the plant transformation vectors used in this study is shown in Supplementary Fig. S1. The fidelity of the constructs was confirmed by PCR and restriction digestion.
Transformation of WT Col-0 and tfl1-14 mutant plants with Agrobacterium strain EHA105 carrying the recombinant constructs was performed using the floral dip method42. Transgenic seedlings were selected for kanamycin resistance and confirmed by genomic PCR and RT-PCR.
Transformation of Jatropha with Agrobacterium strain LBA4404 carrying the overexpression construct was performed according to the protocol described by Fu et al.43. Transgenic Jatropha plants were confirmed by genomic PCR and RT-PCR.
Expression analysis by qRT-PCR
Arabidopsis and Jatropha total RNAs were extracted from frozen tissue using TRIzol reagent (Transgene, China). Synthesis of the first-strand cDNA was performed following the methods described above. qRT-PCR was performed using SYBR® Premix Ex Taq™ II (Takara Bio) on a Roche 480 Real-Time PCR Detection System (Roche Diagnostics).
The primers used for qRT-PCR are listed in lSupplementary Table S1. qRT-PCR was performed using two independent biological replicates and three technical replicates for each sample. Data were analysed using the 2−ΔΔCT method as described by Livak and Schmittgen44. The transcript levels of specific genes were normalised using Arabidopsis Actin2 or Jatropha Actin45.
Additional Information
How to cite this article: Li, C. et al. Three TFL1 homologues regulate floral initiation in the biofuel plant Jatropha curcas. Sci. Rep. 7, 43090; doi: 10.1038/srep43090 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Song, Y. H., Ito, S. & Imaizumi, T. Flowering time regulation: photoperiod- and temperature-sensing in leaves. Trends Plant Sci. 18, 575–583 (2013).
Wickland, D. P. & Hanzawa, Y. The FLOWERING LOCUS T/TERMINAL FLOWER 1 gene family: Functional evolution and molecular mechanisms. Mol. Plant 8, 983–997 (2015).
Higuchi, Y. & Hisamatsu, T. CsTFL1, a constitutive local repressor of flowering, modulates floral initiation by antagonising florigen complex activity in chrysanthemum. Plant Sci. 237, 1–7 (2015).
Srikanth, A. & Schmid, M. Regulation of flowering time: all roads lead to Rome. Cell Mol. Life Sci. 68, 2013–2037 (2011).
Fornara, F., de Montaigu, A. & Coupland, G. SnapShot: Control of flowering In Arabidopsis. Cell 141, 550–550. e552 (2010).
Teotia, S. & Tang, G. To bloom or not to bloom: Role of microRNAs in plant flowering. Mol. Plant 8, 359–377 (2015).
Pin, P. & Nilsson, O. The multifaceted roles of FLOWERING LOCUS T in plant development. Plant Cell Environ. 35, 1742–1755 (2012).
Wolabu, T. W. et al. Three FLOWERING LOCUS T-like genes function as potential florigens and mediate photoperiod response in sorghum. New Phytol. 210, 946–959 (2016).
Zhao, C. et al. A recessive allele for delayed flowering at the soybean maturity locus E9 is a leaky allele of FT2a, a FLOWERING LOCUS T ortholog. BMC Plant Biol. 16, 1–15 (2016).
Wang, Z. et al. Functional evolution of phosphatidylethanolamine binding proteins in soybean and Arabidopsis . Plant Cell 27, 323–326 (2015).
Corbesier, L. et al. FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis . Science 316, 1030–1033 (2007).
Ratcliffe, O. J. et al. A common mechanism controls the life cycle and architecture of plants. Development 125, 1609–1615 (1998).
Repinski, S., Kwak, M. & Gepts, P. The common bean growth habit gene PvTFL1y is a functional homolog of Arabidopsis TFL1 . Theor. Appl. Genet. 124, 1539–1547 (2012).
Bradley, D., Ratcliffe, O., Vincent, C., Carpenter, R. & Coen, E. Inflorescence commitment and architecture in Arabidopsis . Science 275, 80–83 (1997).
Conti, L. & Bradley, D. TERMINAL FLOWER1 is a mobile signal controlling Arabidopsis architecture. Plant Cell 19, 767–778 (2007).
Pillitteri, L. J., Lovatt, C. J. & Walling, L. L. Isolation and characterization of a TERMINAL FLOWER homolog and its correlation with juvenility in citrus. Plant Physiol. 135, 1540–1551 (2004).
Coelho, C. P., Minow, M. A., Chalfun-Júnior, A. & Colasanti, J. Putative sugarcane FT/TFL1 genes delay flowering time and alter reproductive architecture in Arabidopsis . Front. Plant Sci. 5, doi: 10.3389/fpls.2014.00221 (2014).
Mimida, N. et al. Four TFL1/CEN-like genes on distinct linkage groups show different expression patterns to regulate vegetative and reproductive development in apple (Malus×domestica Borkh.). Plant Cell Physiol. 50, 394–412 (2009).
Mohamed, R. Populus CEN/TFL1 regulates first onset of flowering, axillary meristem identity and dormancy release in Populus. Plant J. 62, 674–688 (2010).
Hanzawa, Y., Money, T. & Bradley, D. A single amino acid converts a repressor to an activator of flowering. Proc. Natl. Acad. Sci. USA 102, 7748–7753 (2005).
Danilevskaya, O. N., Meng, X. & Ananiev, E. V. Concerted modification of flowering time and inflorescence architecture by ectopic expression of TFL1-like genes in maize. Plant Physiol. 153, 238–251 (2010).
Liu, X. et al. Analysis of two TFL1 homologs of dogwood species (Cornus L.) indicates functional conservation in control of transition to flowering. Planta 243, 1129–1141 (2016).
Guo, J.-L. et al. Cloning and characterization of a potato TFL1 gene involved in tuberization regulation. Plant Cell Tiss. Org. 103, 103–109 (2010).
Iwata, H. et al. The TFL1 homologue KSN is a regulator of continuous flowering in rose and strawberry. Plant J. 69, 116–125 (2012).
Kotoda, N., Iwanami, H., Takahashi, S. & Abe, K. Antisense expression of MdTFL1, a TFL1-like gene, reduces the juvenile phase in apple. J. Am. Soc. Hortic. Sci. 131, 74–81 (2006).
Teo, Z. W. N., Song, S., Wang, Y.-Q., Liu, J. & Yu, H. New insights into the regulation of inflorescence architecture. Trends Plant Sci. 19, 158–165 (2014).
Ye, J. et al. The Jatropha FT ortholog is a systemic signal regulating growth and flowering time. Biotechnol. Biofuels 7, 91 (2014).
Ni, J. et al. Gibberellin promotes shoot branching in the perennial woody plant Jatropha curcas . Plant Cell Physiol. 56, 1655–1666 (2015).
Koh, M. Y. & Mohd Ghazi, T. I. A review of biodiesel production from Jatropha curcas L. oil. Renew. Sust. Energ. Rev. 15, 2240–2251 (2011).
Tao, Y. B., Luo, L., He, L. L., Ni, J. & Xu, Z. F. A promoter analysis of MOTHER OF FT AND TFL1 (JcMFT1), a seed-preferential gene from the biofuel plant Jatropha curcas . J. Plant Res. 127, 513–524 (2014).
Ghosh, A. et al. Paclobutrazol arrests vegetative growth and unveils unexpressed yield potential of Jatropha curcas . J. Plant Growth Regul. 29, 307–315 (2010).
Contran, N. et al. State-of-the-art of the Jatropha curcas productive chain: From sowing to biodiesel and by-products. Ind. Crop Prod. 42, 202–215 (2013).
Gaul, M. An analysis model for small-scale rural energy service pathways—applied to Jatropha-based energy services in Sumbawa, Indonesia. Energy Sustain. Dev. 16, 283–296 (2012).
Li, C., Luo, L., Fu, Q., Niu, L. & Xu, Z.-F. Identification and characterization of the FT/TFL1 gene family in the biofuel plant Jatropha curcas . Plant Mol. Biol. Rep. 33, 326–333 (2015).
Li, C., Luo, L., Fu, Q., Niu, L. & Xu, Z.-F. Isolation and functional characterization of JcFT, a FLOWERING LOCUS T (FT) homologous gene from the biofuel plant Jatropha curcas . BMC Plant Biol. 14, 125 (2014).
Chua, N.-H., Ye, J., Geng, Y.-F. & Zhang, B. Flowering modification in Jatropha and other plants. Publication No. WO 2013/130016 A1. patent (2013).
Liljegren, S. J., Gustafson-Brown, C., Pinyopich, A., Ditta, G. S. & Yanofsky, M. F. Interactions among APETALA1, LEAFY, and TERMINAL FLOWER1 specify meristem fate. Plant Cell 11, 1007–1018 (1999).
Imamura, T., Nakatsuka, T., Higuchi, A., Nishihara, M. & Takahashi, H. The gentian orthologs of the FT/TFL1 gene family control floral initiation in Gentiana. Plant Cell Physiol. 52, 1031–1041 (2011).
Hou, C.-J. & Yang, C.-H. Functional analysis of FT and TFL1 orthologs from orchid (Oncidium Gower Ramsey) that regulate the vegetative to reproductive transition. Plant Cell Physiol. 50, 1544–1557 (2009).
Ding, L.-W., Sun, Q.-Y., Wang, Z.-Y., Sun, Y.-B. & Xu, Z.-F. Using silica particles to isolate total RNA from plant tissues recalcitrant to extraction in guanidine thiocyanate. Anal. Biochem. 374, 426–428 (2008).
Shao, J.-L. The subcellular localization and the function of tobacco small GTP-binding protein NtRab5b Doctor thesis, Sun Yat-sen University (2009).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana . Plant J. 16, 735–743 (1998).
Fu, Q. et al. An efficient protocol for Agrobacterium-mediated transformation of the biofuel plant Jatropha curcas by optimizing kanamycin concentration and duration of delayed selection. Plant Biotechnol. Rep. 9, 405–416, doi: 10.1007/s11816-015-0377-0 (2015).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Zhang, L., He, L.-L., Fu, Q.-T. & Xu, Z.-F. Selection of reliable reference genes for gene expression studies in the biofuel plant Jatropha curcas using real-time quantitative PCR. Int. J. Mol. Sci. 14, 24338–24354 (2013).
Acknowledgements
This work was supported by the Natural Science Foundation of China (31370595 and 31300568) and the CAS 135 program (XTBG-T02). The authors gratefully acknowledge the Central Laboratory of the Xishuangbanna Tropical Botanical Garden for providing research facilities.
Author information
Authors and Affiliations
Contributions
C.L., Q.F. and Z.-F.X. conceived and designed the experiments. C.L., Q.F., L.N. and L.L. performed the experiments and analysed the data. J.C. contributed materials. C.L. and Z.-F.X. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, C., Fu, Q., Niu, L. et al. Three TFL1 homologues regulate floral initiation in the biofuel plant Jatropha curcas. Sci Rep 7, 43090 (2017). https://doi.org/10.1038/srep43090
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep43090
This article is cited by
-
MsTFL1A delays flowering and regulates shoot architecture and root development in Medicago sativa
Plant Reproduction (2024)
-
Genes Involved in the Transition and Floral Sexual Differentiation of Jatropha curcas L
Plant Molecular Biology Reporter (2024)
-
Functional characterization of three TERMINAL FLOWER 1-like genes from Platanus acerifolia
Plant Cell Reports (2023)
-
JcSEUSS1 negatively regulates reproductive organ development in perennial woody Jatropha curcas
Planta (2023)
-
An ortholog of the MADS-box gene SEPALLATA3 regulates stamen development in the woody plant Jatropha curcas
Planta (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.