Abstract
FOREVER YOUNG FLOWER (FYF) has been reported to play an important role in regulating flower senescence/abscission. Here, we functionally analyzed five Arabidopsis FYF-like genes, two in the FYF subgroup (FYL1/AGL71 and FYL2/AGL72) and three in the SOC1 subgroup (SOC1/AGL20, AGL19, and AGL14/XAL2), and showed their involvement in the regulation of flower senescence and/or abscission. We demonstrated that in FYF subgroup, FYF has both functions in suppressing flower senescence and abscission, FYL1 only suppresses flower abscission and FYL2 has been converted as an activator to promote flower senescence. In SOC1 subgroup, AGL19/AGL14/SOC1 have only one function in suppressing flower senescence. We also found that FYF-like proteins can form heterotetrameric complexes with different combinations of A/E functional proteins (such as AGL6 and SEP1) and AGL15/18-like proteins to perform their functions. These findings greatly expand the current knowledge behind the multifunctional evolution of FYF-like genes and uncover their regulatory network in plants.
Similar content being viewed by others
Introduction
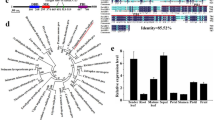
We have previously reported that Arabidopsis FOREVER YOUNG FLOWER (FYF) specifically regulates flower organ senescence and abscission by suppressing the downstream genes of ethylene signaling EDF1/2/3/4 and abscission-associated genes BOP1/2 and IDA1,2. Conserved functions in regulating flower organ senescence and abscission have been reported for FYF orthologs identified in different species of orchids3,4,5. Based on sequence homology and phylogenetic analysis, six putative closely related FYF-like genes could be subjected to two subgroups: (1) the FYF group composed of FYF, AGL71, and AGL72 and (2) the SOC1 group composed of AGL20 (SOC1), AGL19, and AGL14 (XAL2) in the Arabidopsis genome6.
It has been reported that additional new genes could be generated in the genome through gene duplication, which was thought to play an important role in organisms during evolution7,8,9. Phylogenetic analysis revealed that three duplication events from an FYF-like ancestor may have occurred (two within subgroups) to generate six Arabidopsis FYF-like genes6. Functional analysis indicated that the majority of the duplicate FYF-like gene pairs (FYF/AGL71/AGL72/SOC1/AGL19/AGL14) may retain overlap of the original ancestral function, such as the regulation of flowering time10,11,12,13,14,15,16,17,18,19,20,21,22,23, or have specific subsets of the original ancestral function, such as the regulation of root24,25 and ovule26 development as seen for AGL14 (XAL2), which was described as subfunctionalization9,27,28,29,30. Although different putative functions were uncovered for these Arabidopsis FYF-like genes, exploration of the function in regulating flower organ senescence and abscission was investigated for only the FYF gene and has never been reported for other five Arabidopsis FYF-like genes. It is therefore not clear whether the FYF gene evolved to have this unique function in regulating flower organ senescence and abscission from its ancestor or whether some of the other FYF-like genes also harbored this function during evolution. To uncover these questions, we comprehensively functionally characterized all putative Arabidopsis FYF-like genes for their involvement in regulating flower organ senescence and abscission. Furthermore, we used a FRET-based strategy to investigate the possible heterotetrameric protein complexes formed by the interactions of FYF-like and other MADS box proteins to further verify the regulatory networks for these duplicate FYF-like gene pairs in Arabidopsis.
Here, we show that all other five Arabidopsis FYF-like genes have a function in regulating flower senescence and/or abscission similar to FYF. In this work, we also found that FYF-like proteins can interact with different combinations of A/E functional proteins (such as AGL6 and SEP1) and AGL15/18-like proteins (such as AGL15/18) to form heterotetrameric complexes in regulating flower senescence and abscission.
Results
Characterization of two FYF closely related genes, FYL1 and FYL2
Two Arabidopsis thaliana MADS box genes, known to be closely related to FYF (AT5G62165), named FYF-like 1, 2 (FYL1/AGL71, FYL2/AGL72), in the FYF group were analyzed. FYL1 (AT5G51870) encodes a protein containing 219 amino acids that showed 40.3% identity to FYF, with 86.2% (50/58) of the amino acids being identical in the MADS box domain (Supplementary Fig. 1). FYL2 (AT5G51860) encodes a protein containing 202 amino acids that showed 43.7 and 65% identity to FYF and FYL1, respectively, with 86.2% (50/58) and 94.8% (55/58) of the amino acids being identical in the MADS box domain (Supplementary Fig. 1), respectively. The amino acid identity and the phylogenetic tree relationship6 indicated that FYL1 was more closely related to FYL2 than to FYF (Supplementary Fig. 2).
The distinct expression patterns of FYL1 and FYL2
To investigate the expression patterns of the FYL1 and FYL2 genes, FYL1/2 expression was further analyzed in flowers at different developmental stages. The results indicated that higher FYL1/2 expression was observed during early flower development (before stage 9) than during late developmental stages (after stage 12) (Supplementary Fig. 3a, b), which was similar to the spatial and temporal expression pattern of FYF during flower development (Supplementary Fig. 3c)1. When FYL1::GUS and FYL2::GUS constructs were generated and transformed into Arabidopsis, GUS staining was exclusively detected in the abscission zone (AZ) of the sepals/petals of FYL1::GUS flowers (Fig. 1a–c) and shows a more extended pattern in the sepals/petals of FYL2::GUS flowers (Fig. 2a–d). Further RT-qPCR analysis indicated that FYL1 expression was highly detected whereas FYL2 expression was almost undetectable in the AZ (Supplementary Fig. 3d). This result is interesting since GUS staining was detected in both sepals/petals and in the abscission zones of FYF::GUS flowers1, suggesting that FYL1 and FYL2 might have different subfunctions of FYF in regulating flower senescence and abscission.
a GUS was strongly stained in the AZ (arrowed) of floral buds and mature flowers of FYL1::GUS Arabidopsis. The numbers indicate the different developmental stages of Arabidopsis flowers. Bar = 1 mm. Magnified view of stage 10 (b) and 13 (c) FYL1::GUS flowers. GUS was only strongly stained in the AZ (arrowed). s: sepal, p: petal. Bars = 0.5 mm. d Flowers along the inflorescences of 35S::FYL1 (first row), 35S::FYL1+SRDX (second row) and 35S::FYL1+VP16 (third row, left) and wild-type (WT) (third row, right) plants. The numbers indicate the positions of the flowers. Bar = 1.5 mm. Magnified view of the delayed senescent 35 S::FYL1 (e), 35S::FYL1+SRDX (f) and early senescent 35S::FYL1+VP16 (g) flowers. s: sepal, p: petal, st: stamen. Bars = 0.5 mm. Detection of SAG12 (h), EDF1-4 and ERF1 (i) and BOP1/2, IDA and HAESA (j) expression in 35S::FYL1 and 35S::FYL1+SRDX Arabidopsis. Error bars show ± SD. n = 3 biologically independent samples. The expression of each gene in the transgenic plants is given relative to that of the wild-type plant, which was set at 1. The letter “a”, “b” and “c” indicates significant difference from the wild-type (WT) value (a: P < 0.05, b: P < 0.01, and c: P < 0.001). The two-sided Student’s t-test was used. k Flowers along the inflorescence of 35S::FYL1 (first row), 35S::FYL1 + SRDX (second row), and wild-type (third row) plants after exposure to ethylene. Bar = 2 mm. Inflorescence of IDA::FYL1 plants (l) and flowers along the inflorescence (m) of IDA::FYL1 (bottom) and wild-type (WT, top) plants. The flower organs (arrowed) remained on the IDA::FYL1 flower and siliques. Bars = 2 mm. n Magnified view of an IDA::FYL1 flower with a senescent but not abscised phenotype from (m). s: sepal, p: petal, st: stamen. Bar = 0.5 mm.
a GUS was stained in the sepals/petals of flowers of FYL2::GUS Arabidopsis. GUS staining gradually decreased in the mature flowers during the late stage of flower development. The numbers indicate the different developmental stages of Arabidopsis flowers. Bar =1 mm. Magnified view of stage 7 (b), 11 (c), and 13 (d) FYL2::GUS flowers. GUS was strongly and relatively weakly stained in stage 7 young and 11 mature flower buds and was barely detected in stage 13 mature flowers. s: sepal, p: petal, st: stamen. Bars = 0,5 mm. e Flowers along the inflorescences of 35S::FYL2 (first row), 35S::FYL2+SRDX (second row), 35S::FYL2+VP16 (third row) and wild-type (WT) (fourth row) plants. The numbers indicate the positions of the flowers. Bar = 2 mm. Magnified view of early senescent 35S::FYL2 (f) and 35S::FYL2+VP16 (h) flowers and delayed senescent 35S::FYL2+SRDX (g) flowers. s: sepal, p: petal, st: stamen. Bars = 0.5 mm. Detection of SAG12 (i), EDF1-4 and ERF1 (j) and BOP1/2, IDA and HAESA (k) expression in 35S::FYL2+SRDX Arabidopsis. Error bars show ± SD. n = 3 biologically independent samples. The expression of each gene in the transgenic plants is given relative to that of the wild-type plant, which was set at 1. The letter “a”, “b” and “c” indicates significant difference from the wild-type (WT) value (a: P < 0.05, b: P < 0.01, and c: P < 0.001). The two-sided Student’s t-test was used. l Flowers along the inflorescence of 35S::FYL2+SRDX (top) and wild-type (bottom) plants after exposure to ethylene. Bar = 1 mm. m Flowers along the inflorescence of 35S::FYF, 35S::FYF/35S::FYL2, and wild-type (WT) plants. Magnified view of the flower organs that did not become senescent and did not undergo abscission (boxed) in the #7 35 S::FYF flower. s: sepal, p: petal. Bar = 2 mm. Analysis of the interaction of MADS proteins FYF, FYL1, FYL2, SOC1, AGL19, AGL15, and AGL18 fused with CFP to AGL6 (n) and SEP1 (o) fused with YFP through the FRET technique. CyPet- and YPet-fused protein pair fluorescence signals were detected in the nucleus expressed in tobacco leaves. CFP and YFP channels were excited with a 440 nm laser, and these two channels were used to calculate the raw FRET signal. FRET values were divided by CFP signals to calculate the FRET efficiency. The average FRET efficiency values were quantified in multiple samples (n > 4). Image frame = 20 × 20 µm2. N.D. indicates not determined. (Blue line: mean). p Analysis of the effect of FYL2 on FYF-SEP1 interactions. The FRET efficiency for the formation of FYF-CFP/SEP1-YFP complexes was analyzed in tobacco cells by adding different amounts (0, 25, 75, and 100%) of unlabeled FYL2 proteins. The average FRET efficiency values were quantified in multiple samples (n > 4). Image frame = 20 × 20 µm2. (Red line: mean).
FYL1 delayed flower senescence and abscission once ectopically expressed in transgenic Arabidopsis plants
To investigate function correlated to the expression pattern of FYL1, 35S::FYL1, 35S::FYL1+SRDX (containing a suppression motif), and 35S::FYL1-DR+VP16 (containing an activation VP16-AD) were transformed into Arabidopsis. The 35S::FYL1 plants showed a delay in both flower senescence and abscission (Fig. 1d, top and 1e), similar to what was observed in 35S::FYF plants1. Furthermore, enhancement of the delay of flower senescence/abscission was observed in 35S::FYL1+SRDX transgenic plants (Fig. 1d, middle and 1f), and an opposite promotion of flower senescence/abscission was produced in 35S::FYL1-DR+VP16 transgenic plants (Fig. 1d, bottom left and 1g), suggesting that FYL1 should act as a repressor in suppressing flower senescence/abscission, similar to FYF. Once it has been converted into an activator, FYL1-DR+VP16, an opposite dominant negative mutant phenotype will be observed. We also found that the expression of the senescence-associated gene SAG12, downstream genes in ethylene signaling EDF1-3, and ERF1 and abscission-associated genes BOP1/2, IDA, and HAESA were all downregulated in 35S::FYL1 and 35S::FYL1+SRDX plants (Fig. 1h–j). In addition, 35S::FYL1 and 35S::FYL1+SRDX flowers were insensitive to ethylene treatment (Fig. 1k). Our data suggest that FYL1 could have a prominent role like FYF in controlling floral senescence/abscission once ectopically expressed in Arabidopsis flowers. However, FYL1 should only have a partial role in controlling floral abscission in real life since its expression was restricted to the abscission zone (AZ) of the sepals/petals of flowers (Fig. 1a–c and Supplementary Fig. 3d).
To further confirm the relationship between FYL1 and sepal/petal abscission, the FYL1 gene driven by the IDA promoter (IDA::FYL1) was transformed into Arabidopsis. A clear delay in the abscission of the perianth organs was observed in IDA::FYL1 flowers (Fig. 1l–n). However, senescence of the perianth normally occurred from positions 4–5 in these flowers that were not abscised in the IDA:FYL1 transgenic plants (Fig. 1l–n). This phenotype was very similar to what has been observed in ida mutants31 and IDA::FYF plants1; thus, the result supported the role of FYL1 as a suppressor and its function together with FYF in suppressing IDA and sepal/petal abscission.
FYL2 promoted flower senescence and abscission when ectopically expressed in transgenic Arabidopsis plants
Similar to FYL1, 35S::FYL2, 35S::FYL2+SRDX, and 35S::FYL2-DR+VP16 Arabidopsis were generated. 35S::FYL2 plants surprisingly showed promotion of both flower senescence and abscission (Fig. 2e, first row and 2f). A similar promotion of flower senescence/abscission was observed in 35S::FYL2-DR+VP16 transgenic plants (Fig. 2e, third row and 2h), and an opposite delay of flower senescence/abscission was produced in 35S::FYL2+SRDX transgenic plants (Fig. 2e, second row and 2g), suggesting that FYL2 should act as an activator in promoting flower senescence/abscission, in contrast to FYF and FYL1. We also found that the expression of EDF1-4, ERF1, BOP1/2, IDA, and HAESA were all downregulated in 35S::FYL2+SRDX plants (Fig. 2i–k). By contrast, SAG12, EDF1-4, ERF1, IDA, and HAESA were all upregulated in 35S::FYL2 and 35S::FYL2-DR+VP16 transgenic plants (Supplementary Fig. 4). In addition, 35S::FYL2+SRDX flowers were insensitive to ethylene treatment (Fig. 2l). This result suggested that FYL2 should function opposite to FYF and could have the same role as FYF once it is converted into a repressor as FYL2+SRDX, and a dominant negative mutant phenotype in suppressing floral senescence/abscission will be observed. However, FYL2 should only have an opposite role to FYF in controlling floral senescence in real life since its expression was specifically in the sepals/petals of flowers (Fig. 2a–d).
To further confirm the relationship between FYL2 and FYF in regulating sepal/petal senescence, 35S::FYF and 35S::FYL2 were doubly transformed into Arabidopsis, and plants ectopically expressing FYF and FYL2 were generated simultaneously. A clear wild-type-like phenotype by senescence and abscission of the perianth organs at approximately positions 3–4 was observed in the 35S::FYF/35S::FYL2 flowers (Fig. 2m, right), which was earlier than that in the 35S::FYF flowers (Fig. 2m, left) and later than that in the 35S::FYL2 flowers (Fig. 2e, first row). This 35S::FYF/35S::FYL2 intermediate phenotype between 35S::FYF and 35S::FYL2 clearly indicated an antagonistic relationship between FYF and FYL2. Thus, the results supported a role for FYL2 as an activator with a function antagonistic to part of the FYF function in controlling senescence of the sepal/petal.
FYF can interact with AGL6 and SEP1 in regulating flower abscission/senescence
To investigate which proteins could possibly interact with FYF to form a complex in regulating flower organ abscission and senescence, two potential candidates, AGL6 and SEP1, which have been reported to be able to interact with FYF through yeast two-hybrid screen32, were identified. It is important to determine whether the spatial and temporal expression patterns of the AGL6 and SEP1 genes were correlated with FYF. Based on the AGL6::GUS assay, AGL6 expression could be detected in the basal parts and abscission zone of the floral organs during floral development (Supplementary Fig. 5a–c)33 and was detected to be more abundant in wild-type flowers before (BP) than after (AP) pollination (Supplementary Fig. 5d), which overlapped with the expression pattern of FYF. SEP1 has been reported to be expressed in all four whorls of flower organs, is more abundant during early morphological differentiation than in mature flowers, and was not reported in the abscission zone during floral development34,35. These results suggested that FYF might interact with AGL6 to form a complex in regulating sepal/petal abscission. In addition, FYF can interact separately with SEP1 and AGL6 to form a complex in regulating sepal/petal senescence.
We further performed FRET analyses by using FYF-CFP and AGL6/SEP1-YFP to observe physical interactions of FYF and AGL6/SEP1 protein complexes in tobacco cells36. The results confirmed that FYF/AGL6 formed heterodimers with high efficiency in the nucleus (Fig. 2n, column 1, Fig. 3a). The same result was observed for FYF/SEP1 (Fig. 2o, column 1). These results suggested that FYF is able to interact with AGL6 and SEP1 to form complexes in regulating flower senescence and/or abscission.
The steady state of dimerization of the protein complexes FYF-AGL6 (a), FYL1-AGL6 (b), AGL15-AGL6 (c), FYF-AGL15 (d), and FYL1-AGL15 (e) is revealed in scatter diagrams showing pFRET and FRET efficiency. The black dots show the independent cell nuclei, and the yellow boxes indicate the steady-state FRET efficiency range for the protein complex. The mean value of FRET efficiency in the steady state is shown at the top of the schematic model, which is the baseline. The protein fused with CFP/YFP was attached by blue/yellow spots. f Schematic model of the protein interactions in the Arabidopsis abscission complexes (1) FYF-AGL6-AGL6-AGL15 and (2) FYL1-AGL6-AGL6-AGL15. Scatter diagram of the raw FRET (pFRET) and FRET efficiency values of the dimer pairs in adjacent lines AGL6-FYF (g) and AGL6-AGL15 (h) and diagonal lines AGL6-AGL6 (i) and FYF-AGL15 (j) in the abscission complex FYF-AGL6-AGL6-AGL15, with a different number of cell nuclei measured. The green dotted lines indicate the overlapping distribution range at the steady state. The yellow boxes (in g, h, j) indicate the baselines obtained for the dimer pairs. The protein fused with CFP/YFP was attached by blue/yellow spots. k Schematic model and FRET efficiency of four different pairs (two adjacent lines and two diagonal lines) of the protein interactions in the Arabidopsis stable abscission complex FYF-AGL6-AGL6-AGL15. The two adjacent lines (black) show similar FRET efficiencies (31%/30%), and the two diagonal lines (blue) show similar FRET efficiencies (29%/24%). Scatter diagram of the raw FRET (pFRET) and FRET efficiency values of the dimer pairs in adjacent lines AGL6-FYL1 (l) and AGL6-AGL15 (m) and diagonal lines AGL6-AGL6 (n) and FYL1-AGL15 (o) in the abscission complex FYL1-AGL6-AGL6-AGL15, with a different number of cell nuclei measured. The green dotted lines indicate the overlapping distribution range at the steady state. The yellow boxes (in l, m, o) indicate the baselines obtained for the dimer pairs. The protein fused with CFP/YFP was attached by blue/yellow spots. p Schematic model and FRET efficiency of four different pairs (two adjacent lines and two diagonal lines) of the protein interactions in the Arabidopsis stable abscission complex FYL1-AGL6-AGL6-AGL15. The two adjacent lines (black) show similar FRET efficiencies (32%/32%), and the two diagonal lines (blue) show similar FRET efficiencies (36%/34%).
FYL1 can interact with AGL6 in regulating flower abscission
To further investigate whether FYL1 could also interact with proteins similar to FYF in regulating flower organ abscission, FRET analyses were performed. When FYL1-CFP and AGL6/SEP1-YFP were used to observe the physical interactions of FYL1 and AGL6/SEP1, FYL1/AGL6 can form heterodimers with similar efficiency to FYF/AGL6 in the nucleus of tobacco cells (Fig. 2n, column 2, Fig. 3b). However, the efficiency for the formation of FYL1/SEP1 (Fig. 2o, column 2) was clearly lower than that for FYF/SEP1 (Fig. 2o, column 1). These results suggested that FYL1 is able to interact with AGL6 in a more stable manner than with SEP1 to form complexes. Since FYL1 was only expressed in the AZ of the sepals/petals and could only regulate flower organ abscission, it is reasonable to believe that FYL1 can only physically interact with AGL6 in the AZ to regulate abscission of the sepals/petals. Although they can interact, the FYL1/SEP1 complex should not exist during Arabidopsis flower development since these two genes have no overlapping expression pattern.
The ability of AGL6 to interact with FYF and FYL1 to form complexes reveals that its function should be related to FYF/FYL1. This assumption was further supported by the result that a similar delay in the flower senescence/abscission phenotype (Supplementary Fig. 5e–i) and downregulation of EDF1, BOP1/2, IDA, and HAESA (Supplementary Fig. 5j, k) were observed in 35S::AGL6+SRDX plants. This result revealed that FYF/FYL1 can interact with AGL6 to target similar downstream genes once expressed in the same places.
FYF and FYL1 can interact with AGL6/AGL6 and AGL15 proteins to form stable heterotetrameric abscission complexes
Based on the floral quartet model in which plant MADS-box proteins function as higher-order tetrameric complexes37, we hypothesized that FYF-AGL6 heterodimer proteins would further form heterotetrameric complexes with other MADS box proteins in the AZ to regulate sepal/petal abscission. It is interesting to note that two MADS box genes, AGL15 and AGL18, have been reported to be expressed in flower organs and in the AZ of flowers38,39 (Supplementary Fig. 6a–d), and a delay in senescence/abscission of flowers has also been observed in 35S::AGL15 and 35S::AGL18 Arabidopsis plants38,39. The similar delay of flower senescence/abscission (Supplementary Fig. 6e–h) and downregulation of EDF1-4, ERF1. BOP1/2, IDA, and HAESA (Supplementary Fig. 6i–j) in 35S::AGL15+SRDX Arabidopsis supports the notion that AGL15/AGL18 function similarly to FYF/FYL1 as repressors39 in regulating flower organ abscission. To further explore whether AGL15/AGL18 could also interact with proteins similar to FYF/FYL1 in regulating flower organ abscission, FRET analyses were performed. The results indicated that AGL15/AGL6 are able to form heterodimers with similar efficiency to FYF/AGL6 in the nucleus of tobacco cells (Fig. 2n, column 6, Fig. 3c). In contrast, AGL18 showed a very weak interaction with AGL6 (Fig. 2n, column 7). This result revealed that AGL15 can form a complex with AGL6, whereas AGL18 might form different protein complexes to perform the redundant function in regulating flower organ abscission.
To test whether FYF (FYL1), AGL6, and AGL15 could form heterotetrameric abscission complexes, a strategy using in vivo FRET based on the distance change and distance symmetry of a stable tetrameric complex in tobacco leaf cells was performed40. Since FYF is unlikely to interact with AGL15 (Fig. 3d) in the FRET analysis, we analyzed the possible abscission tetramer containing FYF/AGL6 and AGL15/AGL6 heterodimers. In the FYF-AGL6-AGL6-AGL15 complex (Fig. 3f–l), the FRET efficiency of the coexpression admixture of AGL6:CFP/FYF:YFP/AGL15 (Fig. 3g) was similar to that of AGL6:CFP/AGL15:YFP/FYF (Fig. 3h) (31%/30%) (Fig. 3k), and the distribution range overlapped. The FRET efficiency of the coexpression admixture of AGL6:CFP/AGL6:YFP/FYF/AGL15 (Fig. 3i) was similar to that of FYF:CFP/AGL15:YFP/AGL6/AGL6 (Fig. 3j) (29%/24%) (Fig. 3k), and the distribution range overlapped.
Similarly, FYL1 was unlikely to interact with AGL15 (Fig. 3e), and the FRET efficiency of the coexpression admixture of AGL6:CFP/FYL1:YFP/AGL15 (Fig. 3l) was similar to that of AGL6:CFP/AGL15:YFP/FYL1 (Fig. 3m) (32%/34%) (Fig. 3p), and the distribution range overlapped. Similarly, the FRET efficiency of the coexpression admixture of AGL6:CFP/AGL6:YFP/FYL1/AGL15 (Fig. 3n) was similar to that of AGL15:CFP/FYL1:YFP/AGL6/AGL6 (Fig. 3o) (36%/34%) (Fig. 3p), and the distribution range overlapped. These overlapped patterns for the distribution range observed for FYF-AGL6-AGL6-AGL15 and FYL1-AGL6-AGL6-AGL15 heterotetrameric complexes were similar to that for the most stable heterotetrameric complexes PI-AP3-AG-SEP3 described in our previous study40. These results indicated that FYF-AGL6-AGL6-AGL15 and FYL1-AGL6-AGL6-AGL15 are likely stable heterotetrameric complexes in regulating flower organ abscission. In addition, FYF-AGL6-AGL6-AGL15 is also likely a stable heterotetrameric complex in regulating flower organ senescence.
FYL2 can interact with AGL6 and SEP1 in regulating flower senescence
Similarly, the investigation of whether FYL2 could also interact with proteins similar to FYF in regulating flower organ senescence was also performed using FRET analyses. When FYL2-CFP and AGL6-YFP or SEP1-YFP were used to observe the physical interactions of FYL2 and AGL6 or SEP1, a lower efficiency of FYL2/AGL6 heterodimer formation than that of FYF/AGL6 (Fig. 2n, column 1) in the nucleus of tobacco cells was observed (Fig. 2n, column 3). The efficiency for the formation of FYL2/SEP1 (Fig. 2o, column 3) was also lower than that for FYF/SEP1 (Fig. 2o, column 1). These results suggested that similar to FYF, FYL2 can also interact with AGL6 and SEP1 to form senescence complexes, although at a lower efficiency. Since FYL2 was only expressed in the flower organs of sepals/petals, which overlapped with part of AGL6 and SEP1 expression, these data revealed that FYL2 can physically interact with AGL6 and SEP1 during Arabidopsis flower development to regulate sepal/petal senescence.
Since we have already shown that FYL2 functions opposite to FYF in controlling sepal/petal senescence, FYL2 might compete to bind the interacting protein to form a functional complex. To examine this assumption, FRET efficiency for the formation of FYF-CFP/SEP1-YFP complexes was examined in tobacco cells by adding different amounts of unlabeled FYL2 proteins. The results indicated that the efficiency for FYF-CFP to interact with SEP1-YFP (Fig. 2p, column 1) was clearly decreased by the presence of 25–75% of the FYL2 proteins (Fig. 2p, columns 2 and 3). The ability of FYF-CFP to interact with SEP1-YFP was almost completely competed for by the presence of 100% FYL2 protein (Fig. 2p, column 4). Thus, FYL2 competes with FYF to interact with SEP1, performing opposite functions in controlling sepal/petal senescence.
FYF-like genes AGL19/14 and SOC1 are complementary to FYF in regulating flower senescence
We found a possible mechanism involving three FYF-like genes (FYF and FYL1/2) in regulating flower organ senescence and abscission. It is interesting to note that three other genes in the SOC1 subgroup, AGL19, AGL14 (XAL2), and AGL20 (SOC1), were also closely related to the FYF/FYL1/FYL2 genes (Supplementary Figs. 1, 2)6,16. Do these three genes also harbor similar functions to FYF/FYL1/FYL2 in regulating flower senescence/abscission? Interestingly, similar to that observed in 35S::FYF and 35S::FYF+SRDX Arabidopsis, a strong delay in flower senescence and abscission (Fig. 4a–c), insensitivity to ethylene treatment (Fig. 4d–i) and downregulation of EDF1-4, ERF1, BOP2, IDA, and HAESA (Fig. 4j, k) were observed in 35S::AGL19 and 35S::AGL19+SRDX Arabidopsis. In contrast to AGL19, only 35S::AGL14+SRDX and 35S::SOC1+SRDX caused a strong delay in flower senescence/abscission and a downregulation of senescence/abscission-related genes (Supplementary Figs. 7a–d, 8a–c), whereas no or a reduced effect was seen in 35S::AGL14 and 35S::SOC1 plants. These results indicated that AGL19 and AGL14/SOC1 functioned as strong and weak repressors, respectively, and that part of their function complemented FYF in suppressing flower organ senescence/abscission.
a Flowers along the inflorescence of 35S::AGL19 plants. The numbers indicate the positions of the flowers. Bars = 2 mm. b Magnified view of the flower organs of a 35S::AGL19 flower that were not senescent/abscised from (a). s: sepal, p: petal, st: stamen. Bars = 0.5 mm. c Flowers along the inflorescence of 35S::AGL19+SRDX plants. The numbers indicate the positions of the flowers. Bars = 2 mm. Flowers along the inflorescence of wild-type (d, e), 35S::AGL19 (f, g), and 35S::AGL19+SRDX (h, i) plants after exposure to ethylene. Wild-type flowers were senescent (arrowed in d, e), whereas 35S::AGL19 and 35S::AGL19+SRDX flowers were not senescent/abscised. Bars = 2 mm. Detection of EDF1–4 and ERF1 (j) and BOP1/2, IDA and HAESA (k) expression in 35S::AGL19 and 35S::AGL19+SRDX Arabidopsis. Error bars show ± SD. n = 3 biologically independent samples. The expression of each gene in the transgenic plants is given relative to that of the wild-type plant, which was set at 1. The letter “a”, “b” and “c” indicates significant difference from the wild-type (WT) value (a: P < 0.05, b: P < 0.01, and c: P < 0.001). The two-sided Student’s t-test was used. l Detection of AGL19 expression before (BP) and after (AF) pollination. m, n GUS was stained in the sepals/petals of flowers of AGL19::GUS Arabidopsis. GUS was strongly stained in stage 8–10 young flower buds and gradually decreased in the mature flowers during the late stage (after stage 12) of flower development. The numbers indicate the different developmental stages of Arabidopsis flowers. n is the magnified view from (m). s: sepal, p: petal, st: stamen. Bars = 1 mm. Detection of FYL1/FYL2/AGL19/AGL14/SOC1 expression in 35S::FYF Arabidopsis (o) and FYF/FYL1/FYL2/AGL14/SOC1 expression in 35S::AGL19 Arabidopsis (p). Error bars show ± SD. n = 3 biologically independent samples. The expression of each gene in the transgenic plants is given relative to that of the wild-type plant, which was set at 1. The letter “a”, “b” and “c” indicates significant difference from the wild-type (WT) value (a: P < 0.05, b: P < 0.01, and c: P < 0.001). The two-sided Student’s t-test was used.
Similar to the expression pattern of FYF/FYL2, higher AGL19/AGL14/SOC1 expression was observed during early flower development (before stage 9) than during late developmental stages (after stage 12) (Fig. 4l and Supplementary Figs. 7e, 8d), which further revealed possible similar and overlapping functions of FYF and AGL19/AGL14/SOC1. GUS staining was detected in sepal/petal organs and was absent in the AZ of AGL19::GUS (Fig. 4m, n) and SOC1::GUS flowers (Supplementary Fig. 8e–g), suggesting that AGL19/14 and SOC1 might have part of the functions of FYF in regulating flower senescence but not abscission. Although SOC1 and AGL19 might also be involved in regulating flower senescence, they have very low or complete no interactions with AGL6 (Fig. 2n, columns 4, 5) or SEP1 (Fig. 2o, columns 4, 5). This result indicated that the FYF/FYL1/FYL2 and AGL19/14/SOC1 groups might have evolved to have their own interacting partners in regulating flower senescence/abscission.
FYF activates FYL1 and AGL19/14/SOC1 expression to enhance the regulation of flower abscission and senescence
Since FYF has the same function as FYL1 and AGL19/14/SOC1 to regulate abscission and senescence of the sepal/petal, respectively, we were also interested in determining how they work together. When the expression pattern of endogenous FYL1 and AGL19/14/SOC1 was analyzed in 35 S::FYF flowers, we found that the expression of all three genes was clearly upregulated (Fig. 4o). Our results revealed that FYL1 was activated by FYF in the AZ during flower development and could enhance the function of the FYF gene in suppressing sepal/petal abscission, whereas AGL19/14/SOC1 were activated by FYF in sepals/petals during flower development, which could enhance the function of the FYF gene in suppressing sepal/petal senescence. Interestingly, we found that FYF, FYL1, AGL14, and SOC1 expression was also upregulated in 35 S::AGL19 flowers (Fig. 4p). This result suggested that FYF and other FYF-like genes with the strong repressor role, such as AGL19, could reciprocally activate each other to enhance the suppression of sepal/petal senescence.
FYF activates FYL2 expression to regulate flower senescence possibly through a feedback loop
Exploring how FYF competes with FYL2 to oppositely regulate sepal/petal senescence is interesting. When the expression pattern of endogenous FYL2 was analyzed in 35S::FYF flowers, we found that FYL2 expression was upregulated (Fig. 4o). Our results revealed that FYL2 was activated by FYF during flower development and that FYL2 could possibly form a feedback loop to contend with endogenous FYF function and to more appropriately control flower senescence. This assumption was further supported by the downregulation of FYL2 expression in fyf/agl15 double mutants (Supplementary Fig. 9a). We also found that FYL2 expression was upregulated in 35S::AGL19 flowers (Fig. 4p). This result suggested that FYF and AGL19 might control sepal/petal senescence by regulating FYL2 in a similar way.
Discussion
The Arabidopsis MADS box gene FYF can regulate flower organ senescence and abscission1. Ectopic expression of FYF caused a delay of senescence and a deficiency of abscission in flowers of transgenic Arabidopsis and Eustoma grandiflorum1. This study further showed that two tandem repeat FYF-like genes, FYL1, and FYL2, and three other FYF-like genes, AGL19/14 and SOC1, in Arabidopsis were also involved in the regulation of flower organ abscission and/or senescence, and their functions were complementary or antagonistic to FYF.
FYL1 was found to act as a repressor to suppress abscission in sepals/petals with a complementary function to FYF (Fig. 5a). Unexpectedly, FYL2 could function as an activator and antagonize FYF in promoting the senescence of sepals/petals (Fig. 5a). The functions of FYL1/2 are correlated with their expression pattern since FYL1 was specifically expressed in the AZ of sepals/petals, whereas FYL2 expression was detected in the organs of sepals/petals. The amino acid identity and the phylogenetic tree relationship6 revealed that FYL1/2 were possibly the result of two duplication events. The first event generated an FYL1/2 ancestor from FYF, and the second event produced the two tandem repeats FYL1 and FYL2 from this FYL1/2 ancestor. The conserved role of FYF/FYL1/FYL2 during evolution in regulating flower senescence and/or abscission was further supported by their ability to interact with the same MADS box proteins AGL6 and SEP1. The original FYF gene must have contained both regulatory elements in its promoter or introns1,33,41,42,43,44,45, which are required for its expression in the organs and AZ of sepals/petals. In the AZ, FYF specifically interacts with AGL6 to suppress abscission of the sepals/petals (Fig. 5a, b). In the sepal/petal organs, FYF can interact with either AGL6 or SEP1 to suppress senescence (Fig. 5a, c). In the tandem repeat genes FYL1 and FYL2, the subfunctional alteration of the regulatory elements during evolution resulted in restriction of the expression of FYL1 in the AZ and FYL2 in sepal/petal organs. FYL1 should maintain the conserved ability of FYF to interact with AGL6 together to suppress the abscission of sepals/petals (Fig. 5a, b). Conversely, FYL2 only retained the conserved ability of FYF to interact with AGL6 and SEP1 in regulating sepal/petal senescence (Fig. 5a, c). However, FYL2 evolved into a role antagonistic to FYF and possibly helped to control FYF activity through a feedback loop to protect the flower buds from senescence and ensure the final senescence of the mature flowers (Fig. 5a, c).
a In Arabidopsis, six FYF-like genes in two subgroups (FYF/FYL1/FYL2 and AGL19/AGL14/SOC1) were all involved in regulating flower senescence and/or abscission. In the FYF subgroup, FYF acts as a repressor (R) in suppressing both flower senescence (indicated by a blue box) and abscission (indicated by a pink box), and FYL1 acts as a repressor (R) and only suppresses flower abscission (indicated by a pink box). FYL2 functions as an activator (A) in promoting flower senescence (indicated by a gray box). In the SOC1 subgroup, AGL19, AGL14, and SOC1 function as repressors (R) and have only one function in suppressing flower senescence (indicated by a light blue box), with the effect of AGL19 being stronger than that of AGL14/SOC1. In addition, the A/E functional genes AGL6/SEP1 and AGL15/18-like gene AGL15 (as a repressor) can regulate senescence (indicated by a blue box) by interacting with FYF/FYL2, whereas AGL6 and AGL15 can also regulate abscission (indicated by a pink box) by interacting with FYF/FYL1. The size of the letter R in the box correlated with the strength of the repressor for the MADS box proteins. b In the AZ of the perianth, FYF and FYL1 complement each other by forming two identified heterotetrameric abscission complexes, FYF/AGL6/AGL6/AGL15 and FYL1/AGL6/AGL6/AGL15, suppressing flower abscission through the downregulation (⊣) of BOP1/2 and IDA/HAESA expression. c In sepals/petals, FYF, AGL19, AGL14, and SOC1 functioned antagonistically to FYL2 in suppressing flower senescence. An identified FYF/AGL6/AGL6/AGL15 together with FYF/SEP1/Y, AGL19/X/Y, AGL14/X/Y, and SOC1/X/Y heterotetrameric senescence complexes suppressed sepal/petal senescence through the downregulation (⊣) of ethylene downstream gene expression. In contrast, FYL2/AGL6/Y and FYL2/SEP1/Y heterotetrameric complexes promoted sepal/petal senescence by the activation (→) of ethylene downstream gene expression, possibly through a negative feedback loop to FYF/X/Y, AGL19/X/Y, AGL14/X/Y, and SOC1/X/Y. d In these cases from (c), the heterotetrameric complexes are composed of FYF-like, X (in red) and Y (in blue) proteins. FYF-like can be either one of the FYF/FYL1/FYL2/AGL19/AGL14/SOC1, X can be AGL6, SEP1, or any unidentified A/E proteins, whereas Y can be AGL15, AGL18, or any unidentified AGL15/18-like proteins.
In addition to FYL1/2, three putative FYF-like genes in the SOC1 subgroup, AGL19, AGL14, and SOC1, which are closely related to FYF/FYL1/FYL2 genes based on the phylogenetic tree relationship6,16, were also characterized. Based on the results of the functional analysis and the expression patterns of these three genes, we found that AGL19, AGL14, and SOC1 were all involved in the regulation of senescence but not abscission of flower organs (Fig. 5a, c). AGL19 might play a stronger repressive role than AGL14/SOC1, which functions similarly to FYF in suppressing flower senescence (Fig. 5a). Although AGL19/14 and SOC1 were found to be involved in regulating flower senescence, similar to FYF/FYL2, they seemed to perform their function in a different way in terms of finding interacting partners to form functional complexes. For example, FYF/FYL1/FYL2 can interact sufficiently with AGL6/SEP1, whereas AGL19/14/SOC1 can not interact with AGL6/SEP1. This finding indicated that the AGL19/14/SOC1 subgroup might have their own interacting partners in regulating flower senescence that differ from those of FYF/FYL1/FYL2 during evolution.
One interesting finding is that these FYF-like genes could regulate the expression of each other. For example, FYF could positively regulate the expression of FYL1 and AGL19/14/SOC1 to enhance suppression of the abscission and senescence of flower organs, respectively, whereas AGL19 could positively regulate the expression of FYF and AGL14/SOC1 to enhance the suppression of flower organ senescence. This positive reciprocal regulatory network among the FYF-like genes should provide a mechanism to ensure the suppression of senescence/abscission during the early stage of flower organ development (Fig. 5b, c). In addition, we also found a possible feedback loop regulatory mechanism between FYF and its opposite functional activator FYL2. FYF could activate the expression of FYL2, which possibly sequentially antagonized the activity of FYF. In this case, FYF activity will be countervailed at an appropriate level by FYL2, which is high in early and low in late flower development and ensures that sepal/petal senescence will occur after flower maturation and will not occur in the flower bud stage (Fig. 5c).
In addition to the FYF-like genes studied, we also found that one other MADS box protein, AGL15, which has also been characterized to be able to regulate flower organ senescence/abscission (Fig. 5a)38,39, was able to interact with AGL6 to form stable heterotetrameric abscission complexes (FYF-AGL6-AGL6-AGL15 and FYL1-AGL6-AGL6-AGL15) and a senescence complex (FYF-AGL6-AGL6-AGL15) with FYF/FYL1 (Fig. 5b, c). Interestingly, AGL18, a closely related gene to AGL1538,39, has no interaction with AGL6 to form further heterotetrameric abscission/senescence complexes with FYF/FYL1. Thus, despite redundant functions, AGL15 and AGL18 may interact with their specific partners to form different protein complexes to regulate flower organ abscission/senescence (Fig. 5d).
A scenario was proposed to elucidate the evolutionary modification of the functions and complicated network of FYF-like genes in regulating flower senescence/abscission. In Arabidopsis, an FYF-like ancestor duplicated into two subgroups of genes (FYF/FYL1/FYL2 and AGL19/AGL14/SOC1) and eventually evolved to divergent functions through subfunctionalization in regulating various flower development processes, including senescence/abscission. In the FYF subgroup, FYF has both functions in suppressing flower senescence and abscission, whereas FYL1 only suppresses flower abscission, and FYL2’s function is converted into the activation of flower senescence (Fig. 5a). In the SOC1 subgroup, AGL19, AGL14, and SOC1 showed only one function in suppressing flower senescence, with the effect of AGL19 being stronger than that of AGL14/SOC1 (Fig. 5a). FYF-like proteins can form heterotetrameric complexes with A/E functional proteins (AGL6, SEP1, and unidentified X) and AGL15/18-like proteins (AGL15/18 and unidentified Y) to perform their functions (Supplementary Fig. 2 and Fig. 5d). In the AZ, FYF/AGL6/AGL6/AGL15 and FYL1/AGL6/AGL6/AGL15 were two identified heterotetrameric abscission complexes that suppressed flower abscission (Fig. 5b). In sepals/petals, we identified FYF/AGL6/AGL6/AGL15 together with FYF/SEP1/Y, AGL19/X/Y, AGL14/X/Y, and SOC1/X/Y heterotetrameric senescence complexes in suppressing flower senescence (Fig. 5c).
Our findings reveal the potential immense complexity of the different combinations of FYF-like, A/E functional, and AGL15/18-like proteins in forming heterotetrameric abscission/senescence complexes (Fig. 5d). This complicated gene redundancy might explain why it is difficult to identify the senescence/abscission mutant phenotype in a single gene mutation for these genes. In an attempt to mutate FYF (key gene in FYF-like) and AGL15 (key gene in AGL15/18-like) simultaneously, T-DNA mutants for each gene were crossed to generate fyf/agl15 double mutations. Very interestingly, early senescence and abscission of the flowers was observed in these fyf/agl15 double mutants (Supplementary Fig. 9). This result strongly supported our assumption that abscission/senescence heterotetrameric complexes are at least composed of different combinations of FYF-like and AGL15/18-like proteins. Simultaneous mutations in FYF and AGL15 proteins will disrupt the functions of various combinations of the complexes and result in early senescence/abscission mutant phenotypes. In conclusion, our findings not only greatly expand the current knowledge concerning the multifunctional evolution of FYF-like genes in regulating flower senescence/abscission but also provide an excellent example for the study of diverse functionalizations of duplicate gene pairs in plants.
Methods
Plant materials and growth conditions
The T-DNA insertion mutants of FYF (fyf, SALK_047915) and AGL15 (agl15, SALK_076234C) mutants Arabidopsis seeds were obtained from the Arabidopsis Biological Resource Center, Ohio State University, Columbus, OH, USA. Seeds for Arabidopsis were germinated and grown as described previously1,2,46. Arabidopsis seeds were sterilized and placed on agar plates containing 1/2 X Murashige & Skoog medium47 at 4 °C for 2 days. Before being transplanted to soil, the seedlings were grown in growth chambers under long-day conditions (16 h light/8 h dark) at 22 °C for 10 days. The light intensity of the growth chambers was 150 μE m−2 s−1.
Cloning of the cDNA for FYL1, FYL2, AGL6, AGL19, AGL14, SOC1, and AGL15 from Arabidopsis
For 35S::MADS constructs, the cDNAs for FYL1, FYL2, AGL6, AGL19, AGL14, SOC1, and AGL15 were obtained by PCR amplification using gene-specific 5′ and 3′ primers. The primers contained the XbaI and KpnI recognition sites to facilitate the cloning of the cDNAs. The XbaI-KpnI fragment containing the cDNA was cloned into the binary vector pEpyon-22K1 under the control of the CaMV 35S promoter and used for plant transformation. Sequences for the primers are listed in the Supplementary Table 1.
Cloning of the promoter DNA fragment from Arabidopsis
For the FYL1::GUS and FYL2::GUS constructs, the promoter regions which included the 5′UTR and first intron for FYL1 (2.65 kb) and FYL2 (2.43 kb) were obtained by PCR amplification using specific primer pairs from the genomic DNA followed by cloning into the pGEM-T easy vector (Promega, Madison, WI, USA). These promoter fragments were then subcloned into the linker region before the β-Glucuronidase (GUS) coding region in the binary vector pEpyon01k1,2. For the AGL6::GUS, AGL15::GUS, AGL19::GUS, and SOC1::GUS constructs, the promoter regions which included the 5’UTR and first intron for AGL6 (4.96 kb), AGL15 (0.94 kb), AGL19 (4.59 kb) and SOC1 (5.68 kb) were obtained by PCR amplification and these promoter fragments were subcloned into the linker region before the β-Glucuronidase (GUS) coding region in the binary vector pEpyon01k in the same manner as FYL1/2::GUS. Sequences for the primers are listed in Supplementary Table 1. For the IDA::FYL1 construct, the IDA promoter (1.43 kb) was obtained by PCR amplification as described previously1. The cDNA for FYL1 was obtained by PCR amplification. The IDA promoter and the cDNA for FYL1 were subcloned into the modified binary vector pEpyon-12K1. Sequences for the primers are listed in Supplementary Table 1.
Construction of the MADS + SRDX constructs
For the 35S::FYL1+SRDX, 35S::FYL2+SRDX, 35S::AGL6+SRDX, 35S::AGL15+SRDX, 35S::AGL19+SRDX, 35S::AGL14+SRDX, 35S::SOC1+SRDX constructs, the cDNAs for FYL1/FYL2/AGL6/AGL15/AGL19/AGL14/SOC1 were obtained by PCR amplification and cloned into the pEpyon-2aK plasmid upstream of the SRDX (LDLDLELRLGFA*) sequence, under the control of the CaMV 35S promoter as described previously1,2. The sequences for the primers are listed in Supplementary Table 1.
Construction of the MADS + VP16 constructs
For the 35S::FYL1+VP16, and 35S::FYL2+VP16 construct, the cDNAs for FYL1/FYL2 were obtained by PCR amplification and cloned into the pEpyon-2bK plasmid upstream of the VP16-AD fragment sequence, under the control of the CaMV 35S promoter as described previously1,2. The sequences for the primers were listed in Supplementary Table 1.
Plant transformation and transgenic plant analysis
A floral dip method as described elsewhere48 was used to introduce constructs made in this study in the Agrobacterium tumefaciens strain GV3101 into Arabidopsis plants. PCR and RT-PCR analyses were used to verified the transformants that survived in medium containing kanamycin (50 µg/ml). To generate 35S::FYF/35S::FYL2 Arabidopsis, constructs of 35S::FYL2 which contained hygromycin resistant gene were co-transformed with 35S::FYF (kanamycin resistant) into Arabidopsis plants. Transformants that survived in medium containing both kanamycin (50 µg/ml) and hygromycin (5 µg/ml) were selected for further analysis. To generate fyf/agl15 double mutant Arabidopsis, homozygous fyf were crossed with the agl15 T-DNA mutants in the Columbia background and F1 plants were used to further generate the F2 generation. One quarter of the F2 plants were fyf/agl15 and were further verified and selected for further analysis.
Histochemical GUS assay
Histochemical staining was performed under standard method described previously2,49,50. Samples were incubated in solution (0.05 mM Potassium ferricyanide, 0.05 mM Potassium ferrocyanide, 100 mM Phosphate buffer, pH 7.0) containing 2 mM X-Gluc (5-bromo-4-chloro-3-indolyl ß-D-glucuronic acid) for several hours at 37 °C. The sample was examined under a dissecting microscope.
Real-time PCR analysis
For real-time quantitative RT-PCR, the reaction was performed on a MJ Opticon system (MJ Research, Waltham, MA) using SYBR® Green Real-time PCR Master Mix (TOYOBO Co., LTD.). The amplification condition was 95 °C for 10 min, followed by 40 cycles of amplification (95 °C for 15 s, 58 °C for 15 s, 72 °C for 30 s and then plate reading) and melted (50–95 °C with plate readings every 1 °C) as described previously1,2. Sequences for the primers used for real-time quantitative RT-PCR for FYF, FYL1, FYL2, AGL6, AGL15, AGL19, AGL14, SOC1, EDF1, EDF2, EDF3, EDF4, ERF1, SAG12, BOP1, BOP2, IDA, and HAESA, were listed in Supplementary Table 1. The housekeeping gene UBQ10 was used as normalization control with the following primers: RT-UBQ10-1 and RT-UBQ10-251. Data were analyzed using Gene Expression Macro software (version 1.1, Bio-Rad).
Ethylene responses
As described previously1,52, wild-type and transgenic Arabidopsis plants were sealed in plastic chambers and gassed with air or air containing 10 ppm ethylene for 3 days in a 16 h light/8 h dark cycle and phenotype analyzed.
FRET analysis
The procedure used to prepare FRET-associated fusion constructs was described in previous studies36,53. To fuse FYF/FYL1/FYL2/AGL6/SEP1/AGL15/AGL18/AGL19/SOC1 with CFP or YFP, the cDNAs for Arabidopsis FYF/FYL1/FYL2/AGL6/SEP1/AGL15/AGL18/AGL19/SOC1 were obtained by PCR amplification using gene-specific primers and cloned into the pEpyon-36K and pEpyon-37K vectors upstream of the CFP or YFP sequence under the control of the CaMV 35S promoter. The following gene-specific primers were used: FYF, FYF-FRET-F-PstI (5′-CTGCAGATGGTTAGAGGAAAGATAGAGATGAAG-3′) and FYF-FRET-R-ns-SalI (5′-GTCGACGCAGTTTCTATTTGGCAAACCG-3′); FYL1, FYL1-FRET-F-XbaI (5′-TCTAGAATGGTGAGAGGGAAGATCGAGATC-3′) and FYL1-FRET-R-ns-kpnI (5′-GGTACCTAGCCGAGTCACGGGCAATC-3′); FYL2, FYL2-FRET-F-XbaI (5′-TCTAGACTGCAGATGGGAAGGGGAAGAGTTGA-3′) and FYL2-FRET-R-ns-kpnI (5′-GGTACCTGGTCGGTTCTTCAGAAATCC-3′); AGL6, AGL6-FRET-F-XbaI (5′-TCTAGAATGGGAAGAGGGAGAGTGG-3′) and AGL6-FRET-R-ns-kpnI (5′-GGTACCAAGAACCCAACCTTGGACG-3′); SEP1, SEP1-FRET-F-pstI (5′-CTGCAGATGGGAAGAGGAAGAGTAGAGCTGAAGAG-3′) and SEP1-FRET-R-ns-SalI (5′-GTCGACGAGCATCCACCCCGGGATGT-3′); AGL15, AtAGL15-FRET-F-XbaI (5′-TCTAGAATGGGTCGTGGAAAAATCGAG-3′) and AtAGL15-FRET-R-ns-kpnI (5′-GGTACCAACAGAGAACCTTTGTCTTTTGGC-3′); AGL18, AtAGL18-FRET-F-XbaI (5′-TCTAGAATGGGTCGGGGAAAGATAGA-3′) and AtAGL18-FRET-R-ns-kpnI (5′-GGATCCATCAGAAGCCACTTGACTCC-3′); AGL19, AtAGL19-FRET-F-XbaI (5′-TCTAGAATGGTGAGGGGCAAAACG-3′) and AtAGL19-FRET-R-ns-kpnI (5′-GGTACCATTTTGAGGAGGGAATTTTTTGGATTGTC-3′); SOC1, AtSOC1-FRET-F-XbaI (5′-TCTAGAATGGTGAGGGGCAAAACTC-3′) and AtSOC1-FRET-R-ns-kpnI (5′-GGTACCCTTTCTTGAAGAACAAGGTAACCCAATG-3′). These constructs were transformed into the Agrobacterium strain C58C1. Different ectopic proteins were expressed in tobacco cells and a confocal microscope was used to detect the fluorescence signals in the nucleus. To perform the subcellular localization assay, Agrobacterium-infiltrated N. benthamiana leaves were vacuum infiltrated in 10 mM MgCl2 at room temperature until immersed. As previously described36,52, an Olympus FV1000 confocal microscope (Olympus FV1000, Tokyo, Japan) and the FV-ASW 3.0 software were used to visualize fluorophores and to calculate the raw FRET and FRET efficiency values. To evaluate the variation in protein interaction distances among different protein complexes (n > 4), the mean value of FRET efficiency in the nucleus was calculated.
Statistics and reproducibility
Data in the analysis of gene expression in various transgenic plants were analyzed using the two-sided Student’s t-test and represented as the mean ± SD. In these cases, n = 3 biologically independent samples. The letter “a”, “b” and “c” indicates significant difference from the wild-type (WT) value (a: P < 0.05, b: P < 0.01, and c: P < 0.001).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data supporting the findings of this work are available within the paper and the Supplementary Information files. The data sets generated and analyzed during this study are available from the corresponding author upon request.
References
Chen, M. K. et al. The MADS box gene, FOREVER YOUNG FLOWER, acts as a repressor controlling floral organ senescence and abscission in Arabidopsis. Plant J. 68, 168–185 (2011).
Chen, W. H., Li, P. F., Chen, M. K., Lee, Y. I. & Yang, C. H. FOREVER YOUNG FLOWER negatively regulates ethylene response DNA-binding factors by activating an ethylene-responsive factor to control Arabidopsis floral organ senescence and abscission. Plant Physiol. 168, 1666–1683 (2015).
Chen, M. K., Lee, P. F. & Yang, C. H. Delay of flower senescence and abscission in Arabidopsis transformed with an FOREVER YOUNG FLOWER homolog from Oncidium orchid. Plant Signal. Behav. 6, 1841–1843 (2011).
Chen, W. H., Lee, Y. I. & Yang, C. H. Ectopic expression of two FOREVER YOUNG FLOWER Orthologues from Cattleya orchid suppresses ethylene signaling and DELLA results in delayed flower senescence/abscission and reduced flower organ elongation in Arabidopsis. Plant Mol. Biol. Report. 36, 710–724 (2018).
Chen, W. H., Jiang, Z. Y., Hsu, H. F. & Yang, C. H. Silencing of FOREVER YOUNG FLOWER like genes from Phalaenopsis orchids promotes flower senescence and abscission. Plant Cell Physiol. 62, 111–124 (2021).
Parenicová, L. et al. Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis new openings to the MADS world. Plant Cell 15, 1538–1551 (2003).
Ohno, S. Evolution by Gene Duplication (Springer-Verlag, 1970).
Xu, G., Guo, C., Shan, H. & Kong, H. Divergence of duplicate genes in exon–intron structure. Proc. Natl Acad. Sci. USA 109, 1187–1192 (2012).
Kuzmin, E. et al. Exploring whole-genome duplicate gene retention with complex genetic interaction analysis. Science 368, 1446 (2020).
Cao, S. et al. Genetic architecture underlying light and temperature mediated flowering in Arabidopsis, rice, and temperate cereals. N. Phytologist 230, 1731–1745 (2021).
Gómez-Soto, D. et al. Overexpression of a SOC1-related gene promotes bud break in Ecodormant Poplars. Front. Plant Sci. 12, 670497 (2021).
Lee, H. et al. The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev. 14, 2366–2376 (2000).
Lee, J. & Lee, I. Regulation and function of SOC1, a flowering pathway integrator. J. Exp. Bot. 61, 2247–2254 (2010).
Lee, S., Kim, J., Han, J. J., Han, M. J. & An, G. Functional analyses of the flowering time gene OsMADS50, the putative SUPPRESSOR OF OVEREXPRESSION OF CO 1/AGAMOUS-LIKE 20 (SOC1/AGL20) ortholog in rice. Plant J. 38, 754–764 (2004).
Preston, J. C., Jorgensen, S. A. & Jha, S. G. Functional characterization of duplicated SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1-Like genes in petunia. PLOS ONE 9, e96108 (2014).
Dorca-Fornell, C. et al. The Arabidopsis SOC1-like genes AGL42, AGL71, and AGL72 promote flowering in the shoot apical and axillary meristems. Plant J. 67, 1006–1017 (2011).
Schönrock, N. et al. Plycomb-group proteins repress the floral activator AGL19 in the FLC-independent vernalization pathway. Genes Dev. 20, 1667–1678 (2006).
Liang, S. et al. Transcriptional regulations on the low-temperature-induced floral transition in an Orchidaceae species, Dendrobium nobile: An expressed sequence tags analysis. Comp. Funct. Genomics 2012, 757801 (2012).
Kim, W., Latrasse, D., Servet, C. & Zhou, D. X. Arabidopsis histone deacetylase HDA9 regulates flowering time through repression of AGL19. Biochem. Biophys. Res. Commun. 432, 394–398 (2013).
Kang, M. J., Jin, H. S., Noh, Y. S. & Noh, B. Repression of flowering under a noninductive photoperiod by theHDA9-AGL19-FT module in Arabidopsis. N. Phytol. 206, 281–294 (2015).
Liu, X. R. et al. Overexpression of an orchid (Dendrobium nobile) SOC1/TM3-Like ortholog, DnAGL19, in Arabidopsis regulates HOS1-FT expression. Front. Plant Sci. 7, 99 (2016).
Pe´rez-Ruiz, R. V. et al. XAANTAL2 (AGL14) Is an important component of the complex gene regulatory network that underlies Arabidopsis shoot apical meristem transitions. Mol. Plant. 8, 796–813 (2015).
Azpeitia, E. et al. Cauliflower fractal forms arise from perturbations of floral gene networks. Science 373, 192–197 (2021).
Garay-Arroyo, A. et al. The MADS transcription factor XAL2/AGL14 modulates auxin transport during Arabidopsis root development by regulating PIN expression. EMBO J. 32, 2884–2895 (2013).
Alvarez-Buylla, E. R. et al. MADS-box genes underground becoming mainstream: Plant root developmental mechanisms. N. Phytol. 223, 1143–1158 (2019).
Xu, G., Huang, J., Lei, S. K., Sun, X. G. & Li, X. Comparative gene expression profile analysis of ovules provides insights into Jatropha curcas L. ovule development. Sci. Rep. 9, 15973 (2019).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Semon, M. & Wolfe, K. H. Preferential subfunctionalization of slow-evolving genes after allopolyploidization in Xenopus laevis. Proc. Natl Acad. Sci. USA 105, 8333–8338 (2008).
Cusack, B. P. & Wolfe, K. H. When gene marriages don’t work out: Divorce by subfunctionalization. Trends Genet. 23, 270–272 (2007).
Wapinski, I., Pfeffer, A., Friedman, N. & Regev, A. Natural history and evolutionary principles of gene duplication in fungi. Nature 449, 54–61 (2007).
Butenko, M. A. et al. Inflorescence deficient in abscission controls floral organ abscission in Arabidopsis and identifies a novel family of putative ligands in plants. Plant Cell 15, 2296–2307 (2003).
de Folter, S. et al. Comprehensive interaction map of the Arabidopsis MADS box transcription factors. Plant Cell 17, 1424–1433 (2005).
Schauer, S. E. et al. Intronic regulatory elements determine the divergent expression patterns of AGAMOUS-LIKE6 subfamily members in Arabidopsis. Plant J. 59, 987–1000 (2009).
Flanagan, C. A. & Ma, H. Spatially and temporally regulated expression of the MADS-box gene AGL2 in wild-type and mutant Arabidopsis flowers. Plant Mol. Biol. 26, 581–595 (1994).
Rounsley, S. D., Ditta, G. S. & Yanofsky, M. F. Diverse roles for MADS box genes in Arabidopsis development. Plant Cell 7, 1259–1269 (1995).
Hsu, H. F. et al. Model for perianth formation in orchids. Nat. Plants 1, 15046 (2015).
Theissen, G. & Saedler, H. Plant biology. Floral quartets. Nature 409, 469–471 (2001).
Fernandez, D. E. et al. The embryo MADS domain factor AGL15 acts postembryonically: Inhibition of perianth senescence and abscission via constitutive expression. Plant Cell 12, 183–198 (2000).
Adamczyk, B. J., Lehti-Shiu, M. D. & Fernandez, D. E. The MADS domain factors AGL15 and AGL18 act redundantly as repressors of the floral transition in Arabidopsis. Plant J. 50, 1007–1019 (2007).
Mao, W. T., Hsu, W. H., Li, J. Y. & Yang, C. H. Distance-based measurement determines the coexistence of B protein hetero- and homodimers in lily tepal and stamen tetrameric complexes. Plant J. 105, 1357–1373 (2021).
Sieburth, L. E. & Meyerowitz, E. M. Molecular dissection of the AGAMOUS control region shows that cis elements for spatial regulation are located intragenically. Plant Cell 9, 355–365 (1997).
Deyholos, M. K. & Sieburth, L. E. Separable whorl-specific expression and negative regulation by enhancer elements within the AGAMOUS second intron. Plant Cell 12, 1799–1810 (2000).
Sheldon, C. C., Conn, A. B., Dennis, E. S. & Peacock, W. J. Different regulatory regions are required for the vernalization-induced repression of FLOWERING LOCUS C and for the epigenetic maintenance of repression. Plant Cell 14, 2527–2537 (2002).
Kooiker, M. et al. BASIC PENTACYSTEINE1, a GA binding protein that induces conformational changes in the regulatory region of the homeotic Arabidopsis gene SEEDSTICK. Plant Cell 17, 722–729 (2005).
Gu, R. et al. Functional Characterization of the promoter and second intron of CUM1 during flower development in cucumber (Cucumis sativus L.). Horticultural. Plant J. 4, 103–110 (2018).
Chang, Y. Y. et al. Characterization of the possible roles for B class MADS box genes in regulation of perianth formation in orchid. Plant Physiol. 152, 837–853 (2010).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant 15, 473–479 (1962).
Clough, S. J. & Bent, A. F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Chou, M. L., Haung, M. D. & Yang, C. H. EMF interact with late-flowering genes in regulating floral initiation genes during shoot development in Arabidopsis. Plant Cell Physiol. 42, 499–507 (2001).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Chen, W. H., Hsu, W. H., Hsu, H. F. & Yang, C. H. A tetraspanin gene regulating auxin response and affecting orchid perianth size and various plant developmental processes. Plant Direct 3, 1–20 (2019).
Chen, Q. G. & Bleecker, A. B. Analysis of ethylene signal-transduction kinetics associated with seedling-growth response and chitinase induction in wild-type and mutant Arabidopsis. Plant Physiol. 108, 597–607 (1995).
Hsu, W. H. et al. AGAMOUS-LIKE13, a putative ancestor for the E functional genes, specifies male and female gametophyte morphogenesis. Plant J. 77, 1–15 (2014).
Acknowledgements
This work was supported by grants to C.-H.Y. from the Ministry of Science and Technology, Taiwan, ROC, grant number: MOST 107-2313-B-005-018-MY3 and MOST 109-2326-B-005-001. This work was also financially supported (in part) by the Advanced Plant Biotechnology Center from The Featured Areas Research Center Program within the framework of the Higher Education Sprout Project by the Ministry of Education (MOE) in Taiwan.
Author information
Authors and Affiliations
Contributions
C.-H.Y. developed the overall strategy, designed experiments, and coordinated the project. W.-H.C., P.-T.L., C.-W.T., and M.-C.H. generated transgenic Arabidopsis plants, performed phenotypic and gene expression analyses. H.-F.H. and Y.-C.L. performed gene expression analyses. W.-H.H., C.-W.T., P.-T.L., and W.-T.M. performed FRET analyses. C.-H.Y. prepared and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Jazmin Abraham and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: José Estevez and Luke R. Grinham.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, WH., Lin, PT., Hsu, WH. et al. Regulatory network for FOREVER YOUNG FLOWER-like genes in regulating Arabidopsis flower senescence and abscission. Commun Biol 5, 662 (2022). https://doi.org/10.1038/s42003-022-03629-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-03629-w
This article is cited by
-
Exploring ethylene-related genes in Cannabis sativa: implications for sexual plasticity
Plant Reproduction (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.