Abstract
Leaf senescence is a complex biological process and defense responses play vital role for rice development, their molecular mechanisms, however, remain elusive in rice. We herein reported a rice mutant spotted leaf 32 (spl32) derived from a rice cultivar 9311 by radiation. The spl32 plants displayed early leaf senescence, identified by disintegration of chloroplasts as cellular evidence, dramatically decreased contents of chlorophyll, up-regulation of superoxide dismutase enzyme activity and malondialdehyde, as physiological characteristic, and both up-regulation of senescence-induced STAY GREEN gene and senescence-associated transcription factors, and down-regulation of photosynthesis-associated genes, as molecular indicators. Positional cloning revealed that SPL32 encodes a ferredoxin-dependent glutamate synthase (Fd-GOGAT). Compared to wild type, enzyme activity of GOGAT was significantly decreased, and free amino acid contents, particularly for glutamate and glutamine, were altered in spl32 leaves. Moreover, the mutant was subjected to uncontrolled oxidative stress due to over-produced reactive oxygen species and damaged scavenging pathways, in accordance with decreased photorespiration rate. Besides, the mutant showed higher resistance to Xanthomonas oryzae pv. Oryzae than its wild type, coupled with up-regulation of four pathogenesis-related marker genes. Taken together, our results highlight Fd-GOGAT is associated with the regulation of leaf senescence and defense responses in rice.
Similar content being viewed by others
Introduction
Leaf senescence, as the final stage of leaf development, is an important biological process and affected by intrinsic factors, such as organ age and environment conditions (like light, temperature and plant growth regulators)1,2,3,4,5. Lesion mimic mutant, termed as spotted leaf (spl) in rice, is often associated with early leaf senescence. Hence it provides a tool for dissecting molecular mechanisms about leaf senescence. For example, rapid leaf senescence occurs in rls1 (rapid leaf senescence 1) mutant, and RLS1 encodes an uncharacterized NB-ARM (nucleotide binding site-armadillo domain) protein regulating programed cell death (PCD) during leaf senescence6. SPL28, encoding a clathrin-associated adaptor protein complex 1, initiates leaf senescence through regulating vesicular trafficking7.
Besides displaying necrotic spots, lesion mimic mutants spontaneously activate plant defense response. Previous study reported defense response such as the resistance to bacterial blight inoculation was enhanced in 21 lesion mimic mutants isolated from IR64 rice mutant populations8, and similarly the resistance to rice blast fungus inoculation was increased in five lesion mimic mutants: spl5-2, spl12, spl13, spl14, and spl159. Along with elevated resistance, expressions of some pathogenesis-related (PR) genes were induced in rice lesion mimic mutants. For example, PR1 (pathogenesis-related 1) and PBZ1 (probenazole-inducible 1) were expressed abundantly in spl11, spl18 and HM47 mutants10,11,12,13. Hence, studying lesion mimic mutants will help us to elucidate the mechanism associated with leaf senescence and defense responses.
Glutamate synthase (GOGAT) and glutamine synthetase (GS) are known as key enzymes in the process of inorganic nitrogen assimilation14,15,16. Through coordinated action of GOGAT and GS, the inorganic nitrogen is transformed to organic nitrogen17. Ammonia, the main form of inorganic nitrogen, is assimilated though GS/GOGAT cycle to produce glutamine (Gln) and glutamate (Glu), both of which constitute the main form of organic nitrogen. In higher plants, there are two kinds of GOGAT18, viz. NADH-GOGAT (EC 1.4.7.1) and Fd-GOGAT (EC 1.4.1.14), using NADH and ferredoxin (Fd) as the electron donors respectively. NADH-GOGAT was found mainly in non-photosynthetic tissues such as roots and seeds19, while Fd-GOGAT plays a major role in reassimilating the large amounts of ammonia derived from photorespiration20. There are two expressed Fd-GOGAT genes in Arabidopsis, viz. AtGLU1 and AtGLU2. Analysis of gls mutants revealed that the major role of AtGLU1 is to reassimilate the large amounts of ammonia derived from photorespiration in leaves, while AtGLU2 functions in the assimilation of primary nitrogen in roots20. In another study, authors reported Arabidopsis Fd-GOGAT encoded by GLU1 is dual targeted to the chloroplasts and mitochondria, and plays an important role in photorespiration, largely consistent with the previous study21. In rice, Fd-GOGAT was revealed to locate mainly in mesophyll cells of leaves, chloroplast-containing cross-cells of grain pericarp, as well as in the apical meristem through immunocytological analysis22. Nevertheless, the function of Fd-GOGAT in leaf senescence and defense response was largely unknown in rice.
Here, we report a new spotted leaf mutant (spl32) derived from an indica cultivar 9311 by radiation. The spl32 mutant is covered with necrotic spots in leaves from tip to base in seedling stage, and spotted leaves emerge from bottom to top of the plant. Positional cloning indicated that SPL32 encodes an Fd-GOGAT protein. We further identified the mutant was subjected to oxidative stress due to suppressed reassimilation system of ammonia, thereby inducing increased resistance to Xanthomonas oryzae pv. Oryzae (Xoo). Our results reveal that Fd-GOGAT involves leaf senescence and defense response in rice.
Results
Phenotypic analyses of the spl32 mutant
Leaves of the spl32 mutant remained largely the same as the wild type before four-leaf stage (Fig. 1A). The necrotic spots were initiated from tip of older leaves in five-leaf stage (Fig. 1B) and then spread through whole leaf (Fig. 1C). From tillering to heading, the necrotic spots became more serious, accompanied by shorter culm and smaller grain, compared to the wild-type 9311 (Fig. 1D,E). Agronomic traits, such as tiller number, seed setting rate, and thousand kernel weight, were all significantly decreased in the spl32 mutant (Table 1).
(A,B,C) Phenotypes of 15 (A), 23 (B), 31 (C) days old wild-type 9311 and spl32 seedlings. The white arrows indicate the fourth real leaves of the wild-type 9311 and spl32 plants. The inserts show enlargement of the fourth real leaves of the wild-type 9311 and spl32 mutant (A–C). (D) Phenotypic comparison of the wild-type 9311 and spl32 mutant at heading stage. The insert shows flag leaves of the wild-type 9311 and spl32 plants. (E) Comparison of the wild-type 9311 and spl32 seeds (dehulled). Bars = 5 cm in A, B and C, 15 cm in D and 5 mm in (E). (F) Determination of the chlorophyll contents of the fourth leaves of the wild-type 9311 and spl32 mutant at three different developmental stages: 15, 23, 31d after sowing. Chla, Chlorophyll a; Chlb, chlorophyll b; Car, total carotenoids. Data represent means ± SD of three independent measurements. Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05. FW, fresh weight.
Environmental conditions could induce formation of lesion mimic symptom23, thus a shading experiment was conducted here. There was no difference between shaded leaf and non-shaded leaf in the wild type. Remarkably, non-shaded leaf in the spl32 mutant showed obvious lesion mimic phenotype, in contrast to shaded leaf showing no symptom (Supplemental Fig. S1). Thus the formation of lesion is induced by light in the spl32 mutant.
According to the severity of lesion spots in the spl32 mutant, we divided the phenotype into three stages, i.e. before appearance of spots (the first stage), spots emerge (the second stage), and serious spots present (the third stage). Chlorophyll contents were examined in the three stages. The result showed that the contents of chlorophyll a and chlorophyll b were both decreased during the second and third stages in the spl32 mutant relative to the wild-type 9311(Fig. 1F).
Chloroplast changes in leaves of the spl32 mutant
Transmission electron microscopy was used to compare ultrastructure of chloroplasts in the 4th leaves of wild-type and spl32 seedlings at the all three stages. The mutant leaf had same chloroplast morphology as the wild type at the first stage (Fig. 2A,B), that is, chloroplasts were well-developed with rich lamellae and a small number of osmiophilic bodies. Thus chloroplast development was supposed to be unaffected in the spl32 mutant. At the second stage, the lamellar structure of chloroplasts began to collapse and osmiophilic bodies were obviously increased in the mutant (Fig. 2C,D). Then the shrinking chloroplasts started to disintegrate and gradually little complete thylakoid structure was observed in the spl32 mutant. At the third stage, number and size of osmiophilic bodies in the mutant chloroplasts were significantly increased, relative to the wild type (Fig. 2E,F). This result suggested that chloroplasts in the spl32 mutant are normal during initial development but disintegrated early in later stages.
Gene expression of leaf senescence in the spl32 mutant
Leaf senescence is controlled by a number of genes. Among them, a senescence-induced gene, STAY GREEN (SGR), was reported to regulate chlorophyll degradation24. Analysis by quantitative RT-PCR (qRT-PCR) revealed that SGR transcript was dramatically upregulated in the spl32 mutant at tillering stage (Fig. 3A). Additionally, many transcription factors and senescence-associated genes (SAGs) were reported upregulated to induce leaf senescence25,26. To confirm senescence occurred in the spl32 mutant, we performed gene expression of OsWRKY23, OsWRKY72, Osl43 and Osl85 using qRT-PCR (Fig. 3A). The result showed that expression levels of all four genes were notably increased in the mutant at tillering stage (Fig. 3A), in agreement with the symptom of early leaf senescence.
The expression analysis of two senescence-associated transcription factors (OsWRKY23 and OsWRKY72), two senescence-associated genes (Osl43 and Osl85) SGR (A) and photosynthesis-associated genes (B) at tillering stage. Data represent means ± SD of three independent measurements. Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05. The accession numbers were listed as follows: OsWRKY23 BAG98560.1; OsWRKY23 BAG98549.1; Osl43 AAK82986; Osl85 AAL65398; SGR AAW82954.1; rbcS AAR19268.1; lhcA AAB65793.1; lhcB (Arabidopsis thaliana), AAA32760; rbcL NP_039391.1; psaA AJC99402.1; psbA AJC99383.1; petD AJC99432.1; ndhA AJC99458.1; atpA AJC99399.1.
A previous study showed photosynthesis-associated gene expressions were decreased when leaf senescence appeared5. Thus nine photosynthesis-associated genes were selected for analysis by qRT-PCR (Fig. 3B). Among them, three (rbcS, lhcA, and lhcB) belong to nuclear-encoded genes and the others (rbcL, psaA, psbA, petD, ndhA, and atpA) are chloroplast-encoded ones. The result revealed that all photosynthesis-related genes were down-regulated in spl32 mutant leaves (Fig. 3B).
Map-based Cloning of the SPL32 gene
Genetic analysis from reciprocal crosses between 9311 and the spl32 mutant showed that the lesion mimic phenotype was controlled by a single recessive nuclear locus (Supplemental Table S1). Molecular analysis of an F2 population and its derived F2:3 population from the cross spl32 × 02428 (a japonica rice variety) restricted the SPL32 locus between the markers RM118 and RM172 on chromosome 7 (Fig. 4A). Twelve insertion/deletion (InDel) markers were developed between the two markers (Supplemental Table S2). The SPL32 locus was further narrowed down to a 66-kb region (Fig. 4B), containing eight putative open reading frames (ORFs) (Fig. 4C). Sequencing all ORFs indicated a single base mutation (G into A) in the junction between the third exon and intron in Fd-GOGAT (Os07g0658400) (Fig. 4D) led to alternative splicing in mRNA. Thus a 57-bp sequence was missing in the complementary DNA (cDNA) in the mutant (Fig. 4F). A newly developed InDel marker zf-1 was used to distinguish the wild type from spl32 mutant genotypes by the size of amplified cDNA fragment (Supplemental Table S3). The cDNA of spl32 mutant has two forms: one is supposed to encode a truncated protein (ΔFd-GOGAT) (a strong band) lacking 19 amino acids; and the other one encodes a whole Fd-GOGAT protein (indicated by a weak band), just like wild type (Fig. 4F). Therefore the spl32 mutant is an Fd-GOGAT knockdown mutant.
(A) The SPL32 locus was mapped to the long arm of chromosome 7 between the markers RM118 and RM172. (B) Mapping of the SPL32 locus between markers zzy-21 and zzy-23 based on bacteriophage P1-derived artificial chromosome (PAC) or bacterial artificial chromosome (BAC) clone sequence (PAC1, P0047B07; BAC2, OJ1477_F01; PAC3, P0496C02). ‘n’ represents homozygous recessive individuals derived from an F2 population and an F2:3 population from the cross spl32 × 02428. The number of recombinants is indicated below the map. (C) the SPL32 locus was narrowed down to a 66-kb region; there are 8 putative open reading frames (ORFs). (D) The gene structure of 9311. The junction of the third exon and intron had a single base mutation (G into A) in ORF4 of spl32 mutant. (E) Complementation analysis of the spl32 mutant. The wild-type 9311 plant and homozygous recessive mutant derived from the cross spl32 × Nipponbare transformed with pFd-GOGAT show normal green leaves, whereas the spl32 mutant and homozygous recessive mutant of F2 population transformed with pΔFd-GOGAT show necrotic spots in leaves. (F) PCR analysis with primers Zf-1 showed the difference in the Fd-GOGAT cDNA in the wild-type 9311, spl32 mutant, ΔFd-GOGAT transgenic plants and Fd-GOGAT transgenic plants. Alternative splicing occurs in the coding region of Fd-GOGAT in spl32 mutant. (G) Phylogenetic analysis of Fd-GOGAT. Fd-GOGAT is most closely homologous to maize and sorghum Fd-GOGATs. The accession numbers were listed as follows: SiFd-GOGAT (Solanum lycopersicum), XP_004234830.1; StFd-GOGAT (Solanum tuberosum), XP_006363768.1; RcFd-GOGAT (Ricinus communis), XP_002526914.1; GmFd-GOGAT (Giycine max), XP_006576787.1; CsFd-GOGAT (Cucumis sativus), XP_004136778.1; OsFd-GOGAT (Oryza sativa), NP_001060520.1; ZmFd-GOGAT (Zea mays), NP_001105693.1; SbFd-GOGAT (Sorghumbicolor), XP_002463318.1; AtGLU1 (Arabidopsis thaliana), NP_850763.1; AtGLU2 (Arabidopsis thaliana), NP_181655.1.
To verify identity of Fd-GOGAT, we constructed the plasmid pFd-GOGAT, containing a 5-kb fragment with only Fd-GOGAT coding region of 9311. We also constructed the plasmid pΔFd-GOGAT, containing the 57 bp-deleted fd-gogat coding region of the mutant. Then, we introduced the two constructs into homozygous recessive individuals selected from the cross spl32 × Nipponbare. All transgenic lines containing pFd-GOGAT rescued the lesion mimic phenotype, whereas 10 independent lines transformed with vector pΔFd-GOGAT didn’t (Fig. 4E,F). The result indicated that the 57-bp missing in Fd-GOGAT cDNA was responsible for the necrotic spots in the spl32 mutant. Phylogenetic analysis showed that rice Fd-GOGAT was most closely related to maize and sorghum Fd-GOGATs (Fig. 4G).
Expression analyses and subcellular localization of Fd-GOGAT protein
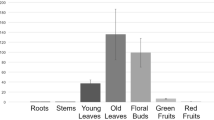
To investigate expression pattern of Fd-GOGAT in rice, we analyzed Fd-GOGAT expression in different tissues by qRT-PCR. We found that Fd-GOGAT was constitutively expressed in leaf, root, panicle and other tissues analyzed. The Fd-GOGAT was most pronounced in young leaves at seedling stage while relatively weak in roots (Fig. 5A), consistent with a previous study that Fd-GOGAT mainly expressed in photosynthetic tissues16. To detect expression levels of Fd-GOGAT at the three different developmental stages in the seedling period, we chose the fourth leaves of the spl32 mutant and 9311 at 15, 23, and 31 days after sowing (DAS) for qRT-PCR analysis. As the development of lesion spots, Fd-GOGAT expression showed a decreasing trend. At 31 DAS when the fourth leaves of spl32 mutant were filled with necrotic spots, Fd-GOGAT almost couldn’t be detected in the mutant (Figs 1C and 5B), indicating that the more severe necrotic spots, the lower expression of Fd-GOGAT in the spl32 mutant.
(A) The expression of Fd-GOGAT in different tissues. Leaf was selected from seedling of 30 days after sowing of the wild-type 9311; root, flag leaf, panicle, and leaf sheath were selected from the wild-type 9311 at heading stage. (B) The expression of Fd-GOGAT at three different developmental stages. The fourth leaves of 15, 23, 31 days after sowing in wild-type 9311 and spl32 mutant were selected. Data are means ± SD (n = 5). Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05. (C) GFP signals of the Fd-GOGAT-GFP fusion protein located in the chloroplasts of rice protoplasts. (D) GFP signals were dispersed in rice protoplasts. Bars = 10 μm.
To further investigate subcellular localization of Fd-GOGAT protein in rice, we use ChloroP27 and TargetP28 to search subcellular localization signals. We found that N-terminus of Fd-GOGAT protein comprised a putative chloroplast-targeting signal. Thus, we constructed a fusion protein with Fd-GOGAT and green fluorescent protein (GFP), and transformed it into rice protoplasts. Confocal microscopy was used to observe the fluorescent signals in 16–18 h after transformation. We observed that green fluorescent signal from Fd-GOGAT-GFP was colocalized with the autofluorescent signals of chlorophylls in the chloroplasts (Fig. 5C,D), suggesting Fd-GOGAT protein is located in the chloroplasts.
The reassimilating of ammonia is inhibited in the spl32 mutant
In photosynthetic green leaves and shoots of Arabidopsis, Fd-GOGAT accounts for more than 96% of total enzyme activity and the remaining is derived from NADH-GOGAT29. Here we measured GOGAT activity in 9311 and the spl32 mutant (Fig. 6A). The result revealed that GOGAT activities in mutant leaves were decreased by 13.2% in non-spotted stage, 12.7% in spot-appearing stage and 13.2% in serious-spotted stage, respectively. We also evaluated effect of the decreased GOGAT activity on GS expression by ELISA Kit (Fig. 6B). It showed that GS activity in the mutant had no difference from 9311 in the all three stages.
(A and B) Enzyme activity of GOGAT and GS in wild-type 9311 and spl32 mutant, non-spotted, spot-appearing and serious-spotted leaves of spl32 mutant and the corresponding sections of leaves in the wild-type 9311 at tillering stage. (C) Determination of NH4+ content in the wild-type 9311 and spl32 mutant at tillering stage. (D) Determination of net photosynthesis rate in atmospheric condition and hypoxia conditions using portable photosynthesis system LI-6400XTOPEN6.1. At tillering stage, new-fully expanded leaves of wild-type 9311 and spl32 mutant were chosen and net photosynthesis rate of mid-blade was measured before 12 o’clock. (E) Determination of GSH content in the wild-type 9311 and spl32 mutant at tillering stage. Data represent means ± SD of three independent measurements. Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05. FW, fresh weight.
Besides, GOGAT and GS were known as key enzymes involved in inorganic nitrogen assimilation14,15,16. To investigate amino acid changes in the spl32 mutant, we measured free amino acid contents in 9311 and the spl32 mutant at tillering stage (Table 2). Many amino acids such as Glu, Ser, Asp, Gly, Ala, Val, Met, Ile, Leu, Tyr, Phe, Lys, and His were significantly decreased, while Gln, Cys and Arg were dramatically increased in the spl32 mutant. Since Glu and Gln are produced from ammonia though GS/GOGAT cycle and both amino acids are significantly altered in the spl32 mutant, so we determined ammonia content at the tillering stage and found ammonium was noticeably increased in the mutant (Fig. 6C).
Previous study indicated Fd-GOGAT plays a major role in photorespiration by reassimilating ammonia20. To evaluate effect of the increased ammonia content on photorespiration, we measured photorespiration rates by supplying hypixic gas (<2%) to wild-type and mutant leaves, as described previously30. The result showed the spl32 mutant had much lower net photosynthetic rate than its wild type under hypoxic conditions, but no difference under atmospheric conditions (Fig. 6D).
Generally photorespiration can affect synthesis of glutathione (GSH), which is produced from condensation reaction of Gly and γ-glutamylcysteine31. Consistent with decreased Gly (Table 2), GSH content was reduced in the spl32 mutant (Fig. 6E).
Antioxidant enzyme activities and active oxygen burst in the spl32 mutant
GSH is an important active substance in plants and participates in formation of disulphide, sulfur ether and thioester. Meanwhile, it can remove free radicals to scavenge their poison in plants32. Hence we determined reactive oxygen species (ROS) content in plant cells by trypan blue dyeing. Leaves with no spots and serious spots in spl32 mutant and the corresponding parts of leaves in the wild type were stained at tillering stage (Fig. 7A). The result showed that no dyeing was observed on non-spotted leaves from either the mutant or wild type. In contrast, leaves with serious spots in spl32 mutant were dyed blue. This result indicated that severe cell necrosis occurred in spl32 mutant when spots emerged.
(A) non-spotted leaves and serious spotted leaves of spl32 mutant and the corresponding sections of leaves in the wild-type 9311 at tillering stage were stained by trypan blue and 3, 3-diaminobenzidine (DAB) respectively. Controls were not stained and decolorized directly by alcohol. (B) The contents of hydrogen peroxide (H2O2) and malonaldehyde (MDA) in the wild-type 9311 and spl32 mutant at tillering stage. FW, fresh weight. (C) The enzyme activities of catalase (CAT), superoxide dismutase (SOD), ascorbate peroxidase (POD), and peroxidase (APX) in wild-type 9311 and spl32 mutant at tillering stage. FW, fresh weight. (D) The enzyme activities of NADPH oxidase and polyamine oxidase (PAO) in wild-type 9311 and spl32 mutant at tillering stage. FW, fresh weight. Data represent means ± SD of three independent measurements. Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05. FW, fresh weight.
To determine accumulation level of hydrogen peroxide in the spl32 mutant, similar leaves in the above treatment were stained with 3,3-diaminobenzidine (DAB). No staining was detected in the non-spotted leaves in both the spl32 mutant and 9311; nevertheless, leaves with serious spots in the spl32 mutant were dyed bronzing (Fig. 7A). Moreover, we determined hydrogen peroxide contents at tillering stage, and found that spl32 mutant leaves accumulated much more hydrogen peroxide than the wild type (Fig. 7B).
Malonaldehyde (MDA) is one of the most important products of membrane lipid peroxidation, and its accumulation can aggravate membrane damage, indirectly reflecting the degree of cellular damage33. Hence, we measured MDA content at tillering stage (Fig. 7B). As expected, the spl32 mutant showed significantly increased MDA in leaves, compared to the wild type. Thus the spl32 mutant tissue could be susceptible to oxidative damage and started senescence more quickly than the wild type.
A previous study showed that enzymes involved in antioxidant systems were expressed in lesion mimic mutants34. Here, we detected activities of four oxidative stress related enzymes, including catalase (CAT), superoxide dismutase (SOD), ascorbate peroxidase (APX), and peroxidase (POD), respectively. As shown in Fig. 7C, activities of CAT and SOD were significantly increased while POD and APX activities were remarkably decreased in the mutant. Meanwhile, two other enzymes that catalyze ROS generation, viz. NADPH oxidase and polyamine oxidase (PAO), were also determined (Fig. 7D). It revealed that activity of PAO was increased while NADPH oxidase showed no change in the spl32 plants.
The spl32 mutant has enhanced disease resistance and up-regulated PR marker genes
The appearance of necrotic spots in the spl32 mutant resembles hypersensitive response (HR), which constitutes an important resistance mechanism in plants35. To evaluate response of the spl32 mutant to pathogen, bacterial blight pathogen Xoo strain Zhe173 was used for inoculation. As shown in Fig. 8A, the spl32 plants demonstrated significantly enhanced resistance compared to its wild type.
(A) Lesion lengths were determined after plant leaves inoculated by bacterial blight pathogen zhe173. Bars represent ± SD of six replicates. (B) The expression of pathogenesis-related (PR) marker genes at tillering stage. Bars represent ± SD of three measurements. The gene numbers of PR genes are as follows: PR1a, AJ278436; PR1b, B109D03; PR5, X68197; PR10, D38170. Asterisks indicate the statistical significance levels according to Student’s t test: **P < 0.01 and *P < 0.05.
Previous studies indicated that oxygen stress not only activated protection mechanism but also acted as a signal factor in undamaged tissues to enable systemic acquired resistance (SAR)36,37. So we detected expressions of four pathogenesis-related (PR) marker genes (PR1a, PR1b, PR5 and PR10) associated with defense response. The results revealed that all PR genes were consistently up-regulated in the spl32 mutant (Fig. 8B).
Discussion
Early leaf senescence occurred in spl32 mutant, confirmed by decreased pigment content and increased MDA content and SOD activity as physiological indicators, chloroplast degradation as cellular indicators, and both upregulation of senescence transcription factors (OsWRKY23 and OsWRKY72) and senescence-associated genes (Osl43 and Osl85) and downregulation of photosynthesis-related genes as molecular evidence. ROS plays an important role in early leaf senescence38. The main sources of ROS are from NADPH oxidase and PAO36. In the spl32 mutant, activity of NADPH oxidase showed no change but PAO activity was increased compared to the wild type. NADPH oxidase converts molecular oxygen into superoxide anion, and PAO interconverts polyamines, producing hydrogen peroxide. Overexpression of AtPAO3 resulted in an increased production of both hydrogen peroxide and superoxide anion39. Here, a large amount of hydrogen peroxide were accumulated, which thereafter induced CAT activity; while superoxide anion did not accumulate in the spl32 mutant by nitrotetrazolium blue chloride test (Supplemental Fig. S2), which might be that the increase of superoxide anion was promptly eliminated by SOD. Moreover, photorespiration provides Gly for GSH synthesis31 and plays a major role in the readjustment of redox homeostasis40. We found photorespiration was dramatically inhibited by over-accumulation of ammonium in the spl32 mutant. Meanwhile, contents of Gly and GSH were both decreased, accompanied by extremely decreased activities of POD and APX in the spl32 mutant. Therefore, the ROS accumulated in the mutant can probably be attributed to overproduction of hydrogen peroxide and damaged scavenging pathway.
OsWRKY23 played an important role in senescence and resistance, and overexpression of OsWRKY23 enhanced expression of PR related genes and increased resistance to the bacterial pathogen Pseudomanas syringae in Arabidopsis41. In the spl32 mutant, disease resistance to Xoo was significantly enhanced, consistent with a recently published study showing Fd-GOGAT plays an important role in broad spectrum bacterial blight resistance42. Moreover, we found four PR marker genes were up-regulated in the spl32 mutant. There are two major signaling pathways in plant disease resistance, viz. salicylic acid (SA) and jasmonic acid (JA)43. Since expressions of PR1a and PR1b were both enhanced in the mutant, suggesting the SPL32 might involve in SA and JA signal transduction.
Through coordinated action of GS and GOGAT, inorganic nitrogen is transformed to organic nitrogen20. GS catalyzes ammonium into Glu, thereby producing Gln. Subsequently, GOGAT transfers amide amino group of Gln to 2-oxoglutrate, forming two molecules of Glu again22. In this study, as contents of ammonium and Gln were remarkably increased, while Glu was decreased in the spl32 mutant, we concluded that Glu could be effectively catalyzed into Gln by functionally normal GS in the mutant. In contrast, decrease of GOGAT activity due to the spl32 mutation caused the over-accumulation of Gln. In addition to changed contents of Glu and Gln, we also found many other free amino acids, except Cys and Arg, were decreased in spl32 mutant, thus it was intriguing to investigate whether and how Fd-GOGAT involves in a complex cross-pathway regulatory mechanism in amino acid metabolism.
It is noteworthy that photorespiration plays an important role in protecting photosynthetic organs from damage through excessive absorption of light energy44. In the spl32 mutant, the lesion was induced by light and photorespiration rate was reduced, therefore light might be a trigger to accelerate oxidative damage in leaves, compared to the wild type.
In Arabidopsis, Fd-GOGAT contributes to major GOGAT enzyme activity, and the remaining is derived from NADH-GOGAT20. There are two NADH-GOGAT genes in rice19. OsNADH-GOGAT1 is mainly expressed in growing tissues, such as immature leaves, the early development of spikelet and roots, while OsNADH-GOGAT2 mRNA is only expressed in leaves and leaf sheathes. These isoforms have distinct functions: OsNADH-GOGAT1 is essential in assimilation of ammonia in roots45; OsNADH-GOGAT2 plays an important role in remobilization of nitrogen during senescence of rice leaves46. In this study, we found transcript levels of the Fd-GOGAT gene were dramatically reduced at 23 DAS and nearly abolished at 31 DAS in the spl32 mutant plants (Fig. 5C), whereas the GOGAT enzyme activity was only reduced constantly by 13% (Fig. 6A) during the period. The discrepancy between transcript levels and enzyme activities is possible due to following reasons: (1) potential post-transcriptional and post-translational regulation of Fd-GOGAT, such as alternative splicing, RNA stability, protein stability and modification, etc.; (2) compensation of another rice Fd-GOGAT, viz. osNADH-GOGAT. In the spl32 mutant, we indeed observed when expression level of Fd-GOGAT fell down to the lowest level at the most serious period of spots, NADH-GOGAT2 had a dramatic increase at the same stage (Supplemental Fig. S3). Therefore we deduce NADH-GOGAT2 might partially but not completely, compensate the function of Fd-GOGAT.
In conclusion, we report herein that SPL32 encodes an Fd-GOGAT that is highly expressed in young leaves. In the spl32 mutant, reassimilation of Gln and ammonia is inhibited due to Fd-GOGAT knockdown. Photorespiration rate and Gly content are decreased as well as glutathione content and some antioxidant enzyme activities, resulting in uncontrolled oxidative stress in the mutant. Moreover, defense response genes are activated and disease resistance is enhanced in the spl32 mutant. This provides a prospect in developing new rice resistant lines by manipulating SPL32 in gene engineering.
Materials and Methods
Plant materials and growth conditions
The rice lesion mimic mutant spl32 was selected from an M2 population of 9311 induced by 60Co radiation. M2 seeds were grown in a paddy field under natural growing conditions in Rice Station, Nanjing Agricultural University, Nanjing, China. The mutant was stabilized through backcrossing with 9311 and selection for three times.
SPL32 gene cloning
For genetic analysis, F2 populations were generated from two crosses of 9311/spl32 and spl32/9311. In both F2 populations, segregation of green-leaf and spotted-leaf individuals fitted a 3:1 ratio. To map the SPL32 gene, we constructed an F2 mapping population generated from a cross spl32 × 02428 (a japonica rice variety). Totally 330 SSR markers evenly spanning 12 rice chromosomes were used to rough map SPL32. The SPL32 was initially mapped to the markers between RM118 and RM172 on chromosome seven using 588 F2 mutant plants. Afterwards, the SPL32 locus was further narrowed down between markers zzy-21 and zzy-23 using 801 F2:3 mutant plants. The mapping primers were listed in Supplemental Table S2. The target region was analyzed with RiceGAAS (http://ricegaas.dna.affrc.go.jp) and eight ORFs were predicted. Then coding sequences of eight ORFs in both 9311 and spl32 mutant were amplified for sequencing.
Complementation of the spl32 mutant
To create complementation construct pFd-GOGAT, cDNA of Fd-GOGAT was amplified by RT-PCR from 9311 with the primer pairs listed in Supplemental Table S3. The resulting fragments were inserted into the vector 1300–221-FLAG in which the Fd-GOGAT gene was under the control of 35S promoter. Since transformation events with 9311 as receptor was difficult, then the construct was introduced into some F2 mutant plants selected from a cross between Nipponbare and spl32 mutant, by Agrobacteriun tumefaciens-mediated method as described previously47.
Determine of pigment contents
Pigment contents were measured according to the method described previously48. Fresh leaves of spl32 mutant plants and 9311 at three different developmental stages (15, 23, and 31 DAS) were collected. Absorption values were measured using UV/Vis spectrophotometer (Beckman DU800 USA).
Transmission electron microscopy
Samples of 9311 and spl32 mutant leaves were prepared for electron microscopy from the fourth leaves of seedling after growing 15, 23, 31 days under standard field conditions. The collected leaves were cut into small pieces and fixed in 2.5% glutaraldehyde in a phosphate buffer at 4 °C for 4 h, rinsed and incubated overnight in 1% OsO4 at 4 °C, dehydrated by an ethanol series, and infiltrated with a series of epoxy resin, and then embedded in Spurr’s medium prior to thin sectioning. Sections were stained again using uranyl acetate and measured with a JEM-1230 electron microscope.
RT-PCR and Real-Time RT-PCR
Total RNA of rice leaves was extracted with a RNA Prep Pure Plant kit (Tiangen) following the manufacturer’s instructions. Each RNA sample (2 μg) was reverse transcribed using the QuantiTect reverse transcription kit (Qiagen). Primer pairs were designed using Primer Express (Applied Biosystems) and listed in Supplemental Table S3. Additionally, the primer information of some genes including STAY GREEN (SGR), two senescence-associated transcription factors, senescence-associated genes and photosynthesis-associated genes were referred from a previous study49. Real-time PCR analysis was performed using ABI7300HT fast real-time PCR system with the SYBR Premix Ex Taq (TaKaRa; catalog no. RR041A). For each sample, we performed real-time PCR with three technical replicates on three biological replicates. ACTIN gene in rice was used as an internal control.
Amino acid sequence alignment and phylogenetic relationship
Searching and downloading the homologous proteins of Fd-GOGAT in different species were carried out in the NCBI website, and then homologous proteins of Fd-GOGAT were analyzed using biological software BioEdit7.0. Multiple sequence alignment was conducted using ClustalW program50, and phylogenetic analysis method described previously51.
Subcellular location of GFP protein
To investigate subcellular location of Fd-GOGAT, coding sequences of Fd-GOGAT were amplified by PCR using primer pairs listed in Supplemental Table S3. PCR products were cloned into pAN580 vector. The vector was transformed into rice protoplasts and incubated in the dark at 24 °C for 14–16 h according to the method described previously52. Finally, GFP fluorescence was observed using a confocal laser scanning microscopy (Zeiss LSM780).
Shading experiment and photorespiration determination
At heading stage, flag leaves (leaves were all green and no spots) in the spl32 mutant and 9311 were wrapped in tinfoil, then the plants were grown in the paddy filed for 10 days under natural light. Five flag leaves (each from mutant and wild type) were randomly selected for treatment. To investigate photorespiration in the spl32 mutant, we measured net photosynthesis rate in atmospheric and hypoxia condition using portable photosynthesis system LI-6400XTOPEN6.1. At tillering stage, we chose new-fully expanded leaves of the spl32 mutant and its wild type, and measured net photosynthesis rate of mid-blade before 12 o’clock. In both mutant and wild type, 10 plants were randomly selected. The gas access of instrument for measuring photosynthesis was input with 98% nitrogen and 2% oxygen to produce hypoxia conditions. Net photosynthesis rates were examined according to the manufacturer’s recommendation.
Determination of various antioxidant indexes
Commercial kits for assaying contents of GSH, hydrogen peroxide, and MDA, as well as enzyme activities for CAT, SOD, and POD were all purchased from Biological Engineering Institute of Nanjing JianchengTM. The enzyme activities of APX, PAO and NADPH oxidase were determined by corresponding Enzyme-Linked Immunosorbnent Assay (ELISA) Kit purchased from Biological Engineering Institute of Jiangsu lyuyeTM. Determination experiments were conducted at tillering stage. All determination protocols referred to manufacture’s instruction. In presence of peroxidases, DAB and hydrogen peroxide could react quickly and form brown polymer deposition. Integrity of cell membrane was examined by trypan blue. Dyeing experiments with trypan blue and DAB were performed according to Wang et al.53 using leaves with non-spotted and serious spotted leaves of spl32 mutant respectively, and their corresponding sections of normal leaves from 9311 at tillering stage. Finally, the leaves were photographed after the decolorization.
Assays of GOGAT and GS activities
GOGAT were determined by GOGAT ELISA Kit purchased from the company of Shanghai YanshengTM and GS activities by GS ELISA Kit from Biological Engineering Institute of Jiangsu lyuyeTM. We selected leaves from different sections of 9311 and spl32 plants (carrying no spots, newly-appearing spots, and serious spots) at tillering stage. Each sample was repeated three times according to the manufacturer’s instructions.
Inoculation with bacterial blight pathogen
Xoo strain Zhe173 (race IV) was provided by Jiangsu Academy of Agricultural Science. New-fully expanded leaves of six independent spl32 plants and 9311 at the tillering stage were inoculated with Zhe173 suspensions (optical density of 0.5 at 600 nm) using clipping leaf method54. Lesion lengths on inoculated plants were measured 21 days after inoculation.
Free amino acids and ammonium content analyses using automatic amino acid analyzer
Free amino acids and ammonium extraction method is as follows: At tillering stage, we chose new-fully expanded leaves of the spl32 mutant and 9311. Samples corresponding to ~300 mg fresh weight (FW) were collected and ground to powders in liquid nitrogen. We also set a blank control without samples. Aliquots of 100 mg FW and blank control were added into 10 mL of 0.1 mol/L HCl, then sealed the lid and properly soaked with shaking. The mixture was extracted by an ultrasonic equipment at temperature 40–50 °C. After 1–1.5 h, the samples were cooled and centrifuged (10 min at 8000 rpm). The proper amount of supernatant was added into equal volume of 10% sulfosalicylic acid. After cooling for an hour at 4 °C, the samples were centrifuged for 30 min at 15,000 rpm, then pH of the supernatant was adjusted with 8 mol/L NaOH to 2.0. Finally, the samples were filtered by 0.45 μm membrane filters and then determined by High Speed Amino Acid Analyzer (Hitachi L-8800). For ammonium content determination, absorbance was measured at 570 nm when retention time was 72.68 min.
Additional Information
How to cite this article: Sun, L. et al. Isolation and characterization of a spotted leaf 32 mutant with early leaf senescence and enhanced defense response in rice. Sci. Rep. 7, 41846; doi: 10.1038/srep41846 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Lutts, S., Kinet, J. M. & Bouharmont, J. NaCl-induced senescence in leaves of rice (Oryza sativa L.) cultivars differing in salinity resistance. Ann. Bot. 78, 389–398 (1996).
Pourtau, N. et al. Interactions of abscisic acid and sugar signaling in the regulation of leaf senescence. Planta. 219, 765–772 (2004).
Munns, R. Genes and salt tolerance: bringing them together. New Phytol. 167, 645–663 (2005).
Céline. M. D. et al. Genetic variation suggests interaction between cold acclimation and metabolic regulation of leaf senescence. Plant Physiol. 143, 434–446 (2007).
Lim, P. O., Kim, H. J. & Nam, H. G. Leaf senescence. Ann. Rev. Plant Biol. 58, 115–136 (2007).
Jiao, B. B. et al. A novel protein RLS1 with NB-ARM domains is involved in chloroplast degradation during leaf senescence in rice. Mol Plant. 5, 205–217 (2012).
Qiao, Y. L. et al. SPL28 encodes a clathrin-associated adaptor protein complex 1, medium subunit micro 1 (AP1M1) and is responsible for spotted leaf and early senescence in rice (Oryza sativa). New Phytol. 185, 258–274 (2010).
Wu, C. J. et al. Rice lesion mimic mutants with enhanced resistance to diseases. Mol Genet Genomics. 279, 605–619 (2008).
Mizobuchi, R. et al. Differential expression of disease resistance in rice lesion-mimic mutants. Plant Cell Rep. 21, 390–396 (2002).
Feng, B. H. et al. Characterization and genetic analysis of a novel rice spotted-leaf mutant HM47 with broad-spectrum resistance to Xanthomonas oryzae pv. oryzae . J Integr Plant Biol. 55, 473–483 (2013).
Kim, S. T. et al. Proteomics analysis of rice lesion mimic mutant (spl1) reveals tightly localized probenazole-induced protein (PBZ1) in cells undergoing programmed cell death. J Proteome Res. 7, 1750–1760 (2008).
Jung, Y. H. et al. Differential expression of defense/stress-related marker proteins in leaves of a unique rice blast lesion mimic mutant (blm). J Proteome Res. 5, 2586–2598 (2006).
Yin, Z. C. et al. Characterizing rice lesion mimic mutants and identifying a mutant with broad-spectrum resistance to rice blast and bacterial blight. Mol Plant Microbe Interact. 13, 869–876 (2000).
Funayama, K. et al. Cytosolic glutamine synthetase1;2 is responsible for the primary assimilation of ammonium in rice roots. Plant Cell Physiol. 54, 934–943 (2013).
Goodall, A. J., Kumar, P. & Tobin, A. K. Identification and expression analyses of cytosolic glutamine synthetase genes in barley (Hordeum vulgare L.). Plant Cell Physiol. 54, 492–505 (2013).
Lam, H. M., Coschigano, K. T., Oliveira, I. C., Melo-Oliveira, R. & Coruzzi, G. M. The molecular-genetics of nitrogen assimilation into amino acids in higher plants. Annu Rev Plant Physiol Plant Mol Biol. 47, 569–593 (1996).
Forde, B. G. & Lea, P. J. Glutamate in plants: metabolism, regulation, and signalling. J Exp Bot. 58, 2339–2358 (2007).
Matoh, T., Ida, S. & Takahashi, E. Isolation and characterization of NADH-glutamate synthase from pea (Pisum sativum L.). Plant Cell Physiol. 21, 1461–1474 (1980).
Tabuchi, M., Abiko, T. & Yamaya, T. Assimilation of ammonium ions and reutilization of nitrogen in rice (Oryza sativa L.). J Exp Bot. 58, 2319–2327 (2007).
Coschigano, K. T., Melo-Oliveira, R., Lim, J. & Coruzzi, G. M. Arabidopsis gls mutants and distinct Fd-GOGAT genes. Implications for photorespiration and primary nitrogen assimilation. Plant Cell. 10, 741–752 (1998).
Jamai, A., Salome, P. A., Schilling, S. H., Weber, A. P. & McClung, C. R. Arabidopsis photorespiratory serine hydroxymethyltransferase activity requires the mitochondrial accumulation of ferredoxin-dependent glutamate synthase. Plant Cell. 21, 595–606 (2009).
Ishiyama, K., Hayakawa, T. & Yamaya, T. Expression of NADH-dependent glutamate synthase protein in the epidermis and exodermis of rice roots in response to the supply of ammonium ions. Planta. 204, 288–294 (1998).
Huang, Q. N. et al. Characterization and genetic analysis of a light- and temperature-sensitive spotted-leaf mutant in rice. J Integr Plant Biol. 53, 671–681 (2011).
Park, S. Y. et al. The senescence-induced staygreen protein regulates chlorophyll degradation. Plant Cell. 19, 1649–1664 (2007).
Zhou, Q. Y. et al. Knockdown of GDCH gene reveals reactive oxygen species-induced leaf senescence in rice. Plant Cell Environ. 36, 1476–1489 (2013).
Lee, R. H., Wang, C. H., Huang, L. T. & Chen, S. C. Leaf senescence in rice plants: cloning and characterization of senescence up-regulated genes. J Exp Bot. 52, 1117–1121 (2001).
Emanuelsson, O., Nielsen, H. & von Heijne, G. ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci. 8, 978–984 (1999).
Emanuelsson, O., Nielsen, H., Brunak, S. & Heijne, G. von . Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol. 300, 1005–1016 (2000).
Suzuki, A. & Rothstein, S. Structure and regulation of ferredoxin-dependent glutamase synthase from Arabidopsis thaliana. Cloning of cDNA expression in different tissues of wild-type and gltS mutant strains, and light induction. Eur J Biochem. 243, 708–718 (1997).
Sharkey, T. D. Estimating the rate of photorespiration in leaves. Physiol Plantarum. 73, 147–152 (1988).
Noctor, G. et al. Light-dependent modulation of foliar glutathione synthesis and associated amino acid metabolism in poplar overexpressing γ-glutamylcysteine synthetase. Planta. 202, 357–369 (1997).
Jones, D. P. Redefining oxidative stress. Antioxid Redox Signal. 8, 1865–1879 (2006).
Li, Z. et al. Fine mapping of the lesion mimic and early senescence 1 (lmes1) in rice (Oryza sativa). Plant Physiol Bioch. 80, 300–307 (2014).
Kliebenstein, D. J., Dietrich, R. A., Martin, A. C., Last, R. L. & Dangl, J. L. LSD1 regulates salicylic acid induction of copper zinc superoxide dismutase in Arabidopsis thaliana . Mol Plant Microbe Interact. 12, 1022–1026 (1999).
Dangl, J. L. & Jones, J. D. Plant pathogens and integrated defence responses to infection. Nature. 411, 826–833 (2001).
Torres, M. A., Dangl, J. L. & Jones, J. D. Arabidopsis gp91phox homologues AtrbohD and AtrbohF are required for accumulation of reactive oxygen intermediates in the plant defense response. Proc Natl Acad Sci USA 99, 517–522 (2002).
Brodersen, P. et al. Knockout of Arabidopsis accelerated-cell-death11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev. 16, 490–502 (2002).
Navabpour, S. et al. Expression of senescence-enhanced genes in response to oxidative stress. J Exp Bot. 54, 2285–2292 (2003).
Sagor, G. H. et al. Reducing cytoplasmic polyamine oxidase activity in Arabidopsis increases salt and drought tolerance by reducing reactive oxygen species production and increasing defense gene expression. Front Plant Sci. 7, 00214, 10.3389/fpls 00214 (2016).
Voss, I., Sunil, B., Scheibe, R. & Raghavendra, A. S. Emerging concept for the role of photorespiration as an important part of abiotic stress response. Plant Biol (Stuttg). 15, 713–722 (2013).
Jing, S. J., Zhou, X., Song, Y. & Yu, D. Q. Heterologous expression of OsWRKY23 gene enhances pathogen defense and dark-induced leaf senescence in Arabidopsis . Plant Growth Regul. 58, 181–190 (2009).
Chen, H. L. et al. The Fd-GOGAT1 mutant gene lc7 confers resistance to Xanthomonas Oryzae Pv. Oryzae in rice. Sci Rep. 6, 26411, 10.1038/srep 26411 (2016).
Xie, X. Z. et al. Phytochromes regulate SA and JA signaling pathways in rice and are required for developmentally controlled resistance to Magnaporthe grisea . Mol Plant. 4, 688–696 (2011).
Streb, P., Shang, W., Feierabend, J. & Bligny, R. Divergent strategies of photoprotection in high-mountain plants. Planta 207, 313–324 (1998).
Tamura, W. et al. Reverse genetics approach to characterize a function of NADH-glutamate synthase1 in rice plants. Amino Acids. 39, 1003–1012 (2010).
Tamura, W. et al. Disruption of a novel NADH-glutamate synthase2 gene caused marked reduction in spikelet number of rice. Front Plant Sci. 2, 57 (2011).
Hiei, Y. & Komari, T. Agrobacterium-mediated transformation of rice using immature embryos or calli induced from mature seed. Nat Protoc. 3, 824–834 (2008).
Wu, Z. M. et al. A chlorophyll-deficient rice mutant with impaired chlorophyllide esterification in chlorophyll biosynthesis. Plant Physiol. 145, 29–40 (2007).
Wang, Z. H. et al. Functional inactivation of UDP-N-acetylglucosamine pyrophosphorylase 1 (UAP1) induces early leaf senescence and defence responses in rice. J Exp Bot. 66, 973–987 (2015).
Thompson, J. D., Gibson, T. J. & Higgins, D. G. Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics. Chapter 2, 2–3 (2002).
Su, N. et al. Disruption of a rice pentatricopeptide repeat protein causes a seedling-specific albino phenotype and its utilization to enhance seed purity in hybrid rice production. Plant Physiol. 159, 227–238 (2012).
Chiu, W. et al. Engineered GFP as a vital reporter in plants. Curr Biol. 6, 325–330 (1996).
Wang, Y. C., Zhang, Y., Wang, Z., Zhang, X. Y. & Yang, S. H. A missense mutation in CHS1, a TIR-NB protein, induces chilling sensitivity in Arabidopsis . Plant J. 75, 553–565 (2013).
Manosalva, P. M., Bruce, M. & Leach, J. E. Rice 14-3-3 protein (GF14e) negatively affects cell death and disease resistance. Plant J. 68, 777–787 (2011).
Acknowledgements
This research was supported by grants from the National Basic Research Program of China (2011CB100102), the 863 Program (2014AA10A604-4), National Science and Technology Support Program (2013BAD01B02-16), Jiangsu Science and Technology Development Program (BE2014394), and Key Laboratory of Biology, Genetics and Breeding of Japonica Rice in Mid-lower Yangtze River, Ministry of Agriculture, P.R. China. We also acknowledge the support of the Jiangsu Collaborative Innovation Center for Modern Crop Production.
Author information
Authors and Affiliations
Contributions
Liting Sun, Yihua Wang, Ling-long Liu, Chunming Wang, Ling Jiang and Jianmin Wan designed research; Liting Sun, Yihua Wang, Ting Gan, Zhengyao Zhang, Di Wang, Mei Niu, Wuhua Long, Yunlong Wang, Xiaohui Li and Ming Zheng performed research; Liting Sun, Yihua Wang, Ling-long Liu, Chunming Wang, Ling Jiang and Jianmin Wan worte the paper. All authors final approval of the version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sun, L., Wang, Y., Liu, Ll. et al. Isolation and characterization of a spotted leaf 32 mutant with early leaf senescence and enhanced defense response in rice. Sci Rep 7, 41846 (2017). https://doi.org/10.1038/srep41846
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41846
This article is cited by
-
ATP-citrate lyase B (ACLB) negatively affects cell death and resistance to Verticillium wilt
BMC Plant Biology (2022)
-
A dominant gene Ihrl1 is tightly linked to and inhibits the gene Ndhrl1 mediating nitrogen-dependent hypersensitive reaction-like phenotype in wheat
Theoretical and Applied Genetics (2022)
-
A review of approaches to control bacterial leaf blight in rice
World Journal of Microbiology and Biotechnology (2022)
-
Primary metabolic processes as drivers of leaf ageing
Cellular and Molecular Life Sciences (2021)
-
Rice EARLY SENESCENCE 2, encoding an inositol polyphosphate kinase, is involved in leaf senescence
BMC Plant Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.