Abstract
Sarcosine oxidase catalyzes the oxidative demethylation of sarcosine to yield glycine, formaldehyde and hydrogen peroxide. In this study, we analyzed the transcription and regulation of the sox locus, including the sarcosine oxidase-encoding genes in Bacillus thuringiensis (Bt). RT-PCR analysis revealed that the sox locus forms two opposing transcriptional units: soxB (soxB/E/F/G/H/I) and soxR (soxR/C/D/A). The typical −12/−24 consensus sequence was located 15 bp and 12 bp from the transcriptional start site (TSS) of soxB and soxC, respectively. Promoter-lacZ fusion assays showed that the soxB promoter is controlled by the Sigma54 factor and is activated by the Sigma54-dependent transcriptional regulator SoxR. SoxR also inhibits its own expression. Expression from the PsoxCR promoter, which is responsible for the transcription of soxC, soxD, and soxA, is Sigma54-dependent and requires SoxR. An 11-bp inverted repeat sequence was identified as SoxR binding site upstream of the soxB TSS. Purified SoxR specifically bound a DNA fragment containing this region. Mutation or deletion of this sequence abolished the transcriptional activities of soxB and soxC. Thus, SoxR binds to the same sequence to activate the transcription of soxB and soxC. Sarcosine utilization was abolished in soxB and soxR mutants, suggesting that the sox locus is essential for sarcosine utilization.
Similar content being viewed by others
Introduction
Sarcosine is reportedly a potential oncometabolite where in prostate cancer sarcosine may serve as a possible sensitive tumor biomarker through its role in tumor progression and metastasis1. Sarcosine oxidase (SOX, EC 1.5.3.1) catalyzes the oxidative demethylation of sarcosine to yield glycine, formaldehyde and hydrogen peroxide, which as the nitrogen source for bacteria growth2. Sarcosine oxidases exist in monomeric, heterodimeric and heterotetrameric (alpha, beta, gamma and delta) forms3 and mediate creatine and glycine betaine metabolism in bacteria. In Arthrobacter sp. 1-IN and Arthrobacter globiformis, the glyA, soxBDAG and purU genes of the sarcosine oxidase operon and the dmg gene of the dimethylglycine oxidase operon work together to catabolize glycine betaine to produce serine4. In Arthrobacter sp. TE1826, the soxA gene, which encodes monomeric sarcosine oxidase (MSOX), forms a cluster with the upstream regulator gene soxR and the downstream crnA and creA genes, which encode creatininase and creatinase, respectively and this Sigma70 factor-regulated cluster participates in the metabolism of creatine to glycine5. The three enzymes encoded by this operon have been used for the diagnostic assessment of serum creatinine levels6. Other studies have shown that the soxR gene product SoxR, a member of the LysR family of regulatory proteins, is a negative regulator of soxA7. A similar genetic organization was observed in Bacillus sp. B-0618, where the gene encoding creatinase is located near the gene encoding MSOX8. In Corynebacterium sp. U-96 and Corynebacterium sp. P-1, the genetic organization of sox and its nearby genes is similar, which both includes glyA, soxBDAG and purU. However, the sdh gene separates soxG and purU in Corynebacterium sp. U-96. The different organization of the sox locus in these species reflects the utilization of sarcosine in different metabolic pathways, e.g., in the metabolism of sarcosine to pyruvate in Corynebacterium sp. U-96 and to serine in Corynebacterium sp. P-19,10.
The Bacillus cereus group of gram-positive, spore-forming bacteria, includes Bacillus cereus, the causative agent of food poisoning in humans; Bacillus thuringiensis (Bt), a pathogenic agent in insects; and Bacillus anthracis, the etiological agent of anthrax in mammals11,12. The organization of the sarcosine oxidase gene and nearby genes is similar in these groups12,13. A gene encoding a Sigma54-dependent transcriptional regulator separates the genes encoding the sarcosine oxidase units in reverse orientation, suggesting that the regulation of the sarcosine oxidase locus in the B. cereus group, which occurs through the Sigma54 factor, is different from the well-studied regulatory mechanisms of other bacteria.
The alternative Sigma factors are the promoter-recognition subunits of bacterial RNA polymerase holoenzymes14. Structural and functional studies have shown these factors can be divided into Sigma70 and Sigma54 classes. Sigma54 promoters have common features: (i) they are devoid of the typical −10/−35 sequences recognized by the Sigma70 factor15 and have strongly conserved −12/−24 regions16; and (ii) they require a positive regulator to stimulate isomerization of the closed complexes of RNA polymerase and the promoter to the corresponding open complexes17,18. Sigma54 plays an important role in the regulation of many metabolic pathways in bacteria19,20,21. The sigL gene in Bacillus subtilis encodes Sigma54 22,23 and was used to identify five Sigma54-dependent transcriptional regulators (or enhancer-binding proteins, EBPs), including LevR, RocR, AcoR, BkdR and YplP, which regulate the levanase operon, arginine metabolic pathway, acetoin catabolic pathway, branched-chain fatty acid synthetic pathway and the cold shock response, respectively. However, only two metabolic pathways are known to be controlled by Sigma54: the γ-aminobutyric acid (GABA)24 and l-lysine metabolism25 pathways in Bt HD73. Little is known about other metabolic pathways controlled by Sigma54 in the B. cereus group.
In this study, we focused on the organization and regulation of the sarcosine oxidase gene and nearby genes (HD73_3147-HD73_3138) in B. thuringiensis subsp. kurstaki strain HD73 (Bt HD73)26. The HD73_3147-HD73_3138 genes of the sox locus were separately designated as soxI, soxH, soxG, soxF, soxE, soxB, soxR, soxC, soxD and soxA. Three SoxR-regulated promoters were identified in the sox locus, two of which are Sigma54-dependent. The results of this study will provide new insight into the metabolic pathways controlled by Sigma54.
Results
Characterization of transcription units in the sox locus
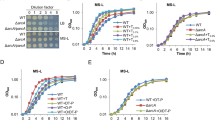
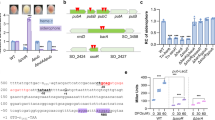
The nucleotide sequence of the sox locus (10,979 bp) of Bt HD73 is comprised of ten open reading frames (ORFs) and encodes ten proteins, which have been annotated as amino acid carrier protein (soxI, HD73_3147), aldehyde dehydrogenase (soxH, HD73_3146), dihydrodipicolinate synthase (soxG, HD73_3145), proline racemase (soxE, HD73_3143), sarcosine oxidase, β subunit (soxB, HD73_3142), Sigma54-dependent transcriptional activator (soxR, HD73_3141), hypothetical protein (soxC/D/F, HD73_3140/HD73_3139/HD73_3144) and sarcosine oxidase, α subunit (soxA, HD73_3138) (Fig. 1A). Identity of Bt Sox proteins to already known function proteins in other bacteria was described in Table S1. The transcriptional units in the sox locus were determined by reverse transcription (RT)-PCR. Products were detected for the ten ORFs in this cluster (Fig. 1B, amplicons 1–10). The products between neighboring genes in both the soxB and soxR orientations were amplified (Fig. 1B, amplicons 11–15 and 17–19); however, no positive signals were detected for amplicons either upstream of soxI or downstream of soxA (Fig. 1B, amplicons 16 and 20). These results suggest that the sox locus is composed of two opposite transcriptional units, soxB/E/F/G/H/I and soxR/C/D/A.
The sox locus in Bt HD73 chromosome.
Panel A, Map of the sox locus in Bt strain HD73. The gray arrows represent ORFs. The positions of fragments that were deleted from the chromosome to disrupt various genes are indicated. Dashed lines with small black arrows annotated with letters correspond to RT-PCR amplicons (see lanes in panel B). The full lines below the ORFs indicate operons. Panel B, RT-PCR analysis of the sox locus in Bt strain HD73. The RNA samples were prepared at T7 of stationary phase (7 hours after the end of the exponential phase) in SSM. The RT-PCR reactions labeled ‘c’ were performed with 500 ng RNA. The positive controls are labeled ‘+’: PCR with 100 ng genomic DNA. The negative controls are labeled ‘−’: RT-PCR with 500 ng RNA with heat-inactivated reverse transcriptase. The letters refer to the positions of the RT-PCR products, as represented in Fig. 1A.
The sequences GGCACGTCAATTGC and GGCATGATTTTTGC (double underline indicated −12/−24 region of consensus sequence) were located upstream of the soxB and soxC gene start codons (Fig. 2), respectively, that were similar to the −12/−24 consensus sequence (BYGGCMYRNNNYYGCW) of Sigma54-binding sites27. The presence of this motif indicates that a third promoter in the sox locus may be controlled by the Sigma54 factor to direct the transcription of soxC and its two downstream genes. DNA microarray data obtained from the HD73 wild-type strain and the sigL mutant strain [GEO: GSE48410] revealed that transcription of the six genes of the soxB operon is significantly higher in the wild-type strain than in the sigL mutant strain28, which is consistent with the RT-PCR analysis of the soxB operon transcriptional unit (Fig. 1B). Among the four genes of the soxR operon, transcription of the soxC, soxD and soxA genes is higher in the wild-type strain than in the sigL mutant. However, transcription of the soxR gene did not significantly differ between strains28, suggesting that a Sigma54-dependent promoter directs the transcription of soxC, soxD and soxA.
Nucleotide sequence of the intergenic region between the soxB and soxC genes.
The single solid underlined regions with asterisks represent transcriptional start codons. The putative ribosome-binding-site (RBS) is indicated by the single dashed underlined. The -12 and -24 sequences are double-underlined. Transcriptional start sites (TSSs) of the soxB, soxR and soxC genes are indicated numerically from the TSS (+1) and marked bold characters. The SoxR binding site maps 54 bp upstream of the TSS of soxB. A 11-bp repeat region (underlined, gray and arrow) maps 54 bp upstream of the soxB TSS. The sequence in the frame represents the soxR gene deletion, which is the same fragment deleted in soxR mutant.
Determination of the transcriptional start site of soxB, soxR and soxC
To determine the TSSs of soxB, soxR and soxC, 5′-RACE analysis was performed as described in the materials and methods. According to the sequences of 16 random clones, a C residue located 17 bp upstream from the soxR start codon was identified in eight and an A residue located 18 bp upstream of the start codon in the remaining eight. Thus, two TSSs were located 17 and 18 bp upstream of the ATG start codon of soxR. The TSSs of soxB and soxC were confirmed to be a single 5′-end nucleotide residue G located 28 bp and 29 bp upstream of the start codon according to the sequences of 11 random clones, respectively. Three typical ribosome-binding sites (RBSs) (-GGAGG-) were identified at an appropriate distance upstream of the start codon of soxB, soxR and soxC. Consistent with the results described above, the sequences upstream of soxB and soxC proved to be −12/−24 motifs.
The transcription and regulation of the soxB and soxR promoters
To characterize the transcription mechanism of the sox locus, the promoters of soxB and soxR were fused with lacZ (Fig. 3A,B) and the expression of PsoxB and PsoxR was assayed in HD73 wild-type, the sigL and soxR mutants (Fig. 3C,D). The results showed that the β-galactosidase activity of PsoxB in HD73 wild-type increased from T0 to T5 and remained high after T5. However, it was abolished from T0 to T3 and significantly reduced after T3 in the ∆soxR and ∆sigL mutants (Fig. 3C). The activity of PsoxB recovered in soxR complementary strain from T0–T8, but not reached to wild type level. These results suggest that the transcriptional activity of the soxB promoter is dependent on Sigma54 and activated by SoxR. The promoter PsoxR showed lower activity than PsoxB in HD73 from T0 to T7 and was lower in HD73 than in the ∆soxR mutant (Fig. 3D), suggesting transcription of the soxR operon is negatively autoregulated. The activity of PsoxR slightly recovered in soxR complementary strain from T0–T7, but not reached to wild type level. However, the activity of PsoxR decreased in ∆sigL mutant compared to that of HD73 wild-type (Fig. 3D), suggesting Sigma54 involved in autoregulation of soxR.
Transcriptional activity of the soxB and soxR promoters.
Panel A, soxB promoter region analysis. The indicated promoter region, 156 bp upstream and 121 bp downstream of the TSS, was fused to lacZ. Panel B, soxR promoter region analysis. The indicated promoter region, 109 bp upstream and 183 bp downstream of the TSS, was fused to lacZ. Panel C, β-galactosidase activity of PsoxB-lacZ in wild-type HD73 (■), the sigL (●) and soxR mutants (▲) and soxR complementary strain (△). Panel D, β-galactosidase activity of PsoxR-lacZ in wild-type HD73 (■), the soxR mutant (▲), the sigL (●) and soxR complementary strain (△). T0 is the end of exponential phase and Tn is n hours after T0. Each value represents the mean of at least three replicates.
Identification of a SoxR-binding site in the soxB promoter fragment
To determine whether SoxR binds to the soxB promoter, SoxR-His protein was expressed in E. coli and purified to near-homogeneity by Ni2+-affinity chromatography (Fig. S1). The ability of SoxR to bind to a DNA fragment containing PsoxB (245 bp) was examined by EMSA. FAM-labeled fragments containing the promoter regions of soxB were incubated with different amounts of SoxR and assayed for the formation of protein-DNA complexes. Slower-migrating probe-protein complexes were observed upon incubation with increasing amounts of SoxR (Fig. 4A). Competitive gel shift assays were performed with labeled DNA probes and about 100-fold of the unlabeled DNA targets (specific competitors) respectively. As shown in Fig. S1, 100-fold excess of soxB promoter probe could dissociate most of the SoxR from labeled soxB promoter probe. Thus, SoxR recognizes and specifically binds to sequences within the soxB promoter fragment.
Identification of the SoxR-binding site in the soxB promoter.
Panel A, Mobility shift assay of the soxB promoter fragment (245 bp) after interaction with SoxR. Lane 1, FAM-labeled PsoxB probe; lanes 2–7, incubation of the probe with increasing concentrations of purified SoxR indicated at the top of the figure. Each lane contained 5.4 ng of probe. Panel B, protection of a 29-bp sequence in the soxB promoter by SoxR, as revealed by DNase I footprinting protection assay. The fluorograms correspond to the DNA in the protection reactions (with 0 and 2.8 μg SoxR). Panel C, soxB promoter analysis. The indicated promoter regions with the wild-type or mutated SoxR binding site (underlined), PsoxB and PsoxBM were fused to lacZ. Panel D, β-galactosidase activity assay of the soxB promoter with the wild-type SoxR-binding site (■) and mutated SoxR-binding site (●). T0 is the end of exponential phase and Tn is n hours after T0. Each value represents the mean of at least three replicates.
To precisely determine the SoxR-binding site in the soxB promoter, DNase I footprinting assays were performed using the same soxB promoter fragment used in the EMSA. A fragment (5′-AAAATATTTTTTACAAATAAAAATATTTT-3′) were protected by SoxR binding (Fig. 4B) (corresponding to the shaded gray and underlined sequence in the soxB promoter region shown in Fig. 2). Moreover, an 11-bp repeat region mapped 54 bp upstream of the TSS of soxB (Fig. 2), with the 11-bp inverted repeat separated by 7 bp (shaded gray in Fig. 2).
To determine whether the proposed sequence is the SoxR binding site in vivo, a 278-bp fragment containing the binding site was mutated from the soxB promoter and the promoter carrying the mutation was fused to lacZ (PsoxBM) (as described in Methods) (Fig. 4C). The activity was sharply reduced in HD(PsoxBM) versus the wild-type HD73 carrying the soxB promoter fused to lacZ (PsoxB-lacZ) (Fig. 4D). This result suggests that disruption of the proposed SoxR-binding site prevents SoxB expression in vivo.
Identification and regulation of the soxC promoter
To identify the promoters of the soxC/D/A genes, a putative promoter fragment (PsoxC) located 126 bp upstream and 70 bp downstream of the TSS of soxC was fused with the lacZ reporter gene (Fig. 5A). This fusion showed no transcriptional activity (Fig. S2), even though a putative −12/24 motif upstream of soxC was identified, suggesting that this fragment did not contain a binding site for SoxR activation of the Sigma54-dependent promoter. PsoxCR, which contains PsoxC and PsoxR with the SoxR-binding site, was fused with the lacZ gene (Fig. 5A) and showed significantly higher transcriptional activity in comparison to PsoxR from T0 to T8 (Fig. 5B). However, the transcriptional activity of PsoxCR was greatly reduced in ∆soxR and ∆sigL mutants (Fig. 5B), suggesting that PsoxCR contains two promoter regions: a PsoxR promoter, which directs transcription of the soxR operon with low-level activity and a PsoxC promoter, controlled by Sigma54 and positively regulated by SoxR with high-level activity to direct the transcription of the soxC, soxD and soxA genes of the soxR operon.
Identification and regulation of the soxC promoter.
Panel A, soxC promoter analysis. The PsoxC region is located 126 bp upstream and 70 bp downstream of the soxC TSS. The PsoxCR region contains PsoxC and the fragment located 113 bp upstream and 23 bp downstream of the soxR TSS and contains a SoxR binding site. The PsoxCDR region contains PsoxCR and no SoxR binding site. These regions were fused to lacZ. Panel B, activity of PsoxCR site in wild-type HD73 (■) and the sigL (●) and soxR mutants (▲) and PsoxCDR promoter without the SoxR-binding site in wild-type HD73 (○). T0 is the end of exponential phase and Tn is n hours after T0. Each value represents the mean of at least three replicates.
To determine whether the SoxR-binding site is necessary for expression from PsoxCR, a 115-bp fragment containing the SoxR-binding sites was deleted from the PsoxCR promoter and the 5′-truncated promoter carrying the deletion was fused to lacZ (PsoxCDR-lacZ) (as described in Methods). The β-galactosidase activity of HD(PsoxCDR) strain was abolished in comparison to wild-type HD73 carrying the PsoxCR promoter (Fig. 5B). Thus, the SoxR-binding site is required for Sigma54-dependent activity of soxC promoter, suggesting that SoxR binds to the same sequence to activate the transcription of both soxB and soxC genes.
The sox locus is responsible for the utilization of sarcosine
The soxA and soxB genes in the sox locus were annotated as the sarcosine oxidase α and β subunits. Sarcosine oxidase catalyzes the oxidative demethylation of sarcosine to yield glycine and has been implicated in creatine and glycine betaine metabolism in some bacteria29,30,31. To evaluate the metabolic role of this locus, mutants with soxB disruptions were constructed. The growth of various strains was tested using sarcosine, proline, creatine, glycine betaine and glycine, as the sole nitrogen sources in glucose minimal medium. The results demonstrated that Bt HD73 utilizes sarcosine, proline, creatine and glycine betaine (Table 1). The disruption of soxR greatly reduced expression of the soxR and soxB operons. The doubling time of the soxR and soxB mutants exceeded 60 h in the presence of sarcosine as the sole nitrogen source and no effect on proline utilization was observed (Table 1). These results clearly indicated that the soxR and soxB operons are responsible for sarcosine utilization. All mutants grew in medium containing either creatine or glycine betaine as a nitrogen source.
Discussion
Transcription of the sox locus, which encodes sarcosine oxidase, is regulated in a Sigma54-dependent manner in Bt. Transcriptional regulation of the sox locus has been studied in only a few bacteria. The sarcosine-responsive transcription factor SouR regulates the soxBDAG operon in Pseudomonas aeruginosa32. Both of the putative promoter regions of soxA gene and the reverse-strand soxR gene possess −10 and −35 sequences in Arthrobacter sp. TE18265,7, suggesting that the sox locus is regulated by Sigma70 in this bacterium. Thus, regulation of the sox locus varies in Bt.
In this study, three promoters of two reversed operons were identified in the sox locus of Bt. PsoxB and PsoxCR were found to be regulated by the Sigma54 factor. The Sigma54-dependent promoter requires an EBP to trigger Sigma54 factor activity33,34. Sigma54-dependent loci typically contain an EBP-encoded gene that is directed by a Sigma54-independent promoter and a Sigma54-dependent gene or operon. For example, the levDEFG35, bkd36 and acoABCL37 operons in B. subtilis are transcribed from Sigma54-dependent promoters and are positively regulated by their EBP (LevR, BkdR and AcoR proteins); these EBP-encoded genes are regulated through a Sigma54-independent promoter. Few studies have shown that three promoters drive the transcription at the Sigma54-dependent gene locus (Fig. S3). For example, roc locus (the rocABC and rocDEF operons and the rocG gene) encode the relevant enzymes of the arginine pathway in B. subtilis23,38,39. There are three conserved −12/−24 motifs upstream of the rocG, rocA and rocD genes and they are expressed in the same direction that is positively regulated by RocR. The rocG-rocABC intergenic region acts as both a downstream activating sequence (DAS) and an upstream activating sequence (UAS) for RcoR protein binding40. A UAS was identified upstream from the translational start codon of the rocDEF operon, which is similar to the rocABC operon23. The sox locus in Bt also has three promoters (Fig. S3), two of them are regulated by the Sigma54 factor. The soxB and soxC genes have conserved −12/−24 motifs, but the promoter of the soxR gene is not a Sigma54-dependent promoter, but is negatively regulated by SoxR in an unknown manner. In contrast to the rocABC operon, the rocDEF operon and the rocG gene are in the same orientation and the soxB operon in the sox locus is located in the opposite direction of the soxR operon.
The binding site of SoxR is located far from the −12/−24 motif of soxC in Bt. The distance from the first G of the −12 element to the A of the translational start codon of the soxR gene is at least 1731 bp. A previous report indicated that the EBP binding sites retain partial activity and activate gene expression, even when located far from its promoter. For example, moving the binding sites for NRI more than 1000 bp does not diminish the ability of NRI to stimulate transcription of glnAp2 in the E. coli glnALG operon41. Expression of the rocG gene in Bacillus subtilis depends on the binding site for RocR, which is located 1.5 kb downstream of its promoter. Furthermore, this activating sequence can be moved as far as 15 kb downstream of the rocG promoter and still retain partial activity39. In this study, we also demonstrated that the binding site of SoxR retains activity for the soxC gene although it is located far from the −12/−24 motif of the soxC gene (at least 1.7 kb). We proposed that the EBP binding site contains a large number of A or T tracts that are in phase with the DNA helix pitch, which might cause sequence curvature, thereby facilitating the interaction of SoxR with a Sigma54-RNA polymerase, such as RocR40. However, the PsoxCRW promoter, which contains the soxR promoter, soxR gene and soxC promoter, had no activity in HD73 or the soxR mutant strains (Fig. S2). Expression of the soxR gene was negatively autoregulated through the promoter PsoxR, suggesting that the level of SoxR protein expression fine-tunes this regulation in Bt. Thus, PsoxCRW may show no promoter activity due to the overexpression of SoxR. It is also a possible reason in that the activity of PsoxB in a genetically complementary strain with soxR was lower than that in wild-type strain (Fig. 3C).
The activity of PsoxR decreased in ∆sigL mutant compared to that of HD73 wild-type (Fig. 3D), suggesting Sigma54 involved in autoregulation of soxR. In sigL mutant strain, Sigma54 could not interaction with SoxR and SoxR specific autoregulated its own promoter. Consequently, the activity of PsoxR decreased in sigL mutant. However, soxR promoter region has no typical −12/−24 conserved sequence, indicating that Sigma54 could not directly control the transcription of soxR. The promoters of soxB, soxC and soxR are regulated by SoxR and used the same SoxR binding site UAS, suggesting that SoxR has a precise regulatory mechanism. The transcription of soxB and soxC are controlled by Sigma54 through the interaction with SoxR binding with UAS. In wild type strain HD73, PsoxR was precisely autoregulated by SoxR interacted with Sigma54. However, the activity of PsoxR decreased by SoxR autoregulation binding with UAS without Sigma54 interaction in sigL mutant. It indicates that Sigma54 positively regulates PsoxR promoter through SoxR as its interaction protein and does not play a role as the sigma factor.
In this study, we showed that the sox locus of Bt HD73 is essential for sarcosine utilization. The soxA and soxB genes encode proteins with high sequence similarity to the sarcosine oxidase alpha and beta units42,43. Disruption of the soxB gene abolished sarcosine utilization in Bt. Conserved domain analysis showed that the SoxC and SoxD proteins contain a 2Fe-2S binding domain, with a 2Fe-2S cluster, which appears in amino acid oxidase proteins44 as well as sarcosine oxidase. Thus, the functions of the SoxC and SoxD proteins might be similar to that of sarcosine oxidase, although the functions of the proteins encoded by the soxI/H/G/F/E genes of the soxB operon remain unknown. All of these proteins showed low sequence similarity and shared no conserved domains with the enzymes involved in the metabolism of creatine, glycine betaine and sarcosine in Arthrobacter4,5 and Corynebacterium9,10.
The orthologs of the sox locus of Bt HD73 are conserved in the genome of B. cereus13,45,46. These genes share high sequence similarity and a similar organization as the sox locus. A similar organization and Sigma54-dependent transcription activator of the sox locus have been identified in other Bacillus cereus species (Fig. S4) and the domains of SoxR from Bt HD73 were conserved in the genomes of these strains. Further analysis of sox locus in these strains revealed a series of putative −12/−24 motifs (Fig. S5). All of these suggest Sigma54 regulates the expression of sarcosine oxidase in these bacteria.
We propose a transcription model for the sox locus in Bt HD73 (Fig. 6). The sox locus in Bt has two opposing operons, soxR (four genes) and soxB (six genes), which contain two Sigma54-regulated promoters, PsoxB and PsoxC. Sigma54 and SoxR regulate the sox locus. RNA polymerases containing Sigma54 recognize the conserved −12/−24 promoter sequence of the soxB and soxC genes and generate closed complexes, while SoxR stimulates the isomerization of the closed complexes to open complexes, thus activating the transcription of the soxB operon and the soxC/D/A genes. The sarcosine oxidase encoded by this locus catalyzes sarcosine to generate glycine. The expression of soxR, which is controlled by PsoxR, is negatively autoregulated.
Schematic representation of sox locus regulation in Bt strain HD73.
The hollow arrows (from left to right) in the middle of the figure indicate the organization of the sox locus in Bt HD73. The magnified section of the figure shows the promoters of soxB, soxR and soxC (which are regulated by SoxR), in which PsoxB, PsoxR and PsoxC represent the putative promoter regions (marked with angled arrows). The single arrows represent positive regulation, the double arrow between soxR and Sigma54 represents interdependence and the block arrow represents negative regulation. The dotted arrows indicate translation.
Methods
Bacterial strains, plasmids and growth conditions
The bacterial strains and plasmids used in this study are listed in Table S2. Bacillus thuringiensis subsp. kurstaki strain HD73 from the Centre OILB (Institut Pasteur, France) that is deposited in Bacillus Genetic Stock Center (BGSCID. 4D4) was used throughout the study (accession numbers CP004069)26. E. coli strain TG1 was used as the host for cloning experiments. The Dam−/Dcm− E. coli ET12567 strain (laboratory stock) was used to generate unmethylated DNA for the electrotransformation assay. Bt strains were transformed by electroporation, as described previously47. E. coli were cultured in Luria-Bertani (LB) medium, with 220 rpm shaking, at 37 °C. Bt was grown in LB medium, Schaeffer’s sporulation medium (SSM)48, or glucose minimal medium (GMM)49 supplemented with 40 mM of a given amino acid as the sole nitrogen source, with vigorous shaking (220 rpm) at 30 °C. The antibiotic concentrations used for bacterial selection were as follows: 100 μg/ml kanamycin and 10 μg/ml erythromycin for Bt and 100 μg/ml ampicillin for E. coli.
DNA manipulation techniques
PCR was performed using Taq and Primestar DNA polymerase (TaKaRa Biotechnology, Dalian, China). Amplified fragments were purified using Axygen purification kits (Silicon Valley, CA, USA). Bt chromosomal DNA was extracted with the Puregene kit (Gentra, Minneapolis, MN, USA). Restriction enzymes and T4 DNA ligase (TaKaRa Biotechnology, Dalian, China) were used according to the manufacturer’s instructions. Oligonucleotide primers (Table S3) were synthesized by Sangon (Shanghai, China). E. coli plasmid DNA was extracted using the Axygen Plasmid Extraction Kit. All constructs were confirmed by DNA sequencing (BGI, Beijing, China).
Total RNA isolation, reverse transcription PCR (RT-PCR)
Total RNA was extracted at stage T7 from Bt cells grown in SSM and the RT-PCR analysis was performed as described50 using primers RT-1 to RT-20. The 16S rRNA gene was PCR-amplified in all samples using the 16SrDNA5/16SrDNA3 primers to verify the absence of DNA contamination.
5′-RACE analysis
The extraction and purification of total RNA were performed as described above. cDNA synthesis and transcriptional start sites (TSSs) of soxB, soxR and soxC were determined using the SMARTerTM RACE cDNA Amplification Kit (Clontech, Mountain View, CA) according to manufacturer instructions. Gene-specific primers and universal primer mix were used to amplify the 5′ end of soxB, soxR and soxC mRNA.
Construction of soxB and soxR promoters with lacZ gene fusion
Fragments of the soxB (278 bp) and soxR (128 bp) promoters were PCR-amplified from strain HD73 DNA using primers PsoxB-F/PsoxB-R and PsoxR-F/PsoxR-R, respectively. The PstI-BamHI restriction fragments were then ligated into pHT304-18Z, which contains a promoterless lacZ gene51. The recombinant pHT-PsoxB and pHT-PsoxR plasmids were introduced into Bt HD73, ΔsigL and ΔsoxR mutant strains, to yield HD73(PsoxB), ΔsigL(PsoxB), ΔsoxR(PsoxB), HD73(PsoxR), ΔsigL(PsoxR) and ΔsoxR(PsoxR), which were selected by resistance to erythromycin and verified by PCR.
Construction of a PsoxC-lacZ fusion
Two DNA fragments of the soxC promoter were fused with lacZ. A 197-bp fragment located between −126 bp and +70 bp was PCR-amplified from strain HD73 with primers PsoxC-F/PsoxCR-R. Another 334-bp fragment was amplified from the ΔsoxR mutant strain with primers PsoxCR-F/PsoxCR-R. This fragment contained the putative SoxR binding site. The two PstI-BamHI restriction fragments were then ligated into the pHT304-18Z vector. The recombinant plasmids pHT-PsoxC and pHT-PsoxCR were introduced into the Bt HD73, ΔsigL and ΔsoxR mutant strains to yield HD73(PsoxC), ΔsigL(PsoxC), ΔsoxR(PsoxC), HD73(PsoxCR), ΔsigL(PsoxCR) and ΔsoxR(PsoxCR). Transformants were selected by resistance to erythromycin and verified by PCR.
Construction of the PsoxC-lacZ fusion bearing deletion of the SoxR-binding site
A fragment containing the 5′-truncated soxC promoters with the SoxR-binding site deleted was fused to lacZ. The construct was obtained as follows: The 219-bp fragment was PCR-amplified from pHT-PsoxCR with primers PsoxCDR-F/PsoxCR-R. The fragment did not contain the SoxR-binding site. The recombinant plasmid pHT-PsoxCDR was introduced into Bt HD73 to produce HD73(PsoxCDR), which were selected by resistance to erythromycin and verified by PCR. The β-galactosidase activity was determined as previously described52 and expressed as Miller units. Reported values represent averages from at least three independent assays.
Expression and purification of SoxR
The expression plasmid pETsoxR containing soxR from Bt strain HD73 was constructed by amplifying soxR with primers SoxR-F and SoxR-R and cloning into BamHI/SalI-digested pET21b. pETsoxR was transferred into E. coli BL21(DE3) and the transformants were grown to the exponential phase in LB medium supplemented with ampicillin at 37°C. The expression and purification of SoxR-His protein was performed as previously described53.
Gel mobility shift assays and DNase I footprinting assays
DNA fragments were PCR-amplified from HD73 genomic DNA using specific primers labeled with a 5′-end 6-FAM modification and confirmed by DNA sequencing. Electrophoresis mobility shift assays (EMSA) were performed as described54 to analyze the binding of purified SoxR to PsoxB DNA containing the putative SoxR binding site. The specificity of the shift was confirmed using poly (dI:dC) and GST protein, bovine serum albumin (BSA) and as negative controls. DNase I footprinting assays were performed based on a fluorescence labeling procedure53.
Construction of the PsoxB-lacZ fusion bearing mutation of SoxR-binding site
A fragment containing the soxB promoter with the SoxR-binding site mutated was cloned in fusion with the lacZ gene. The construct was obtained as follows: The fragment (278 bp) with the SoxR-binding site was mutated (A to G, the mutation site was indicated in Fig. 4) by gene synthesis (GENEWIZ, Suzhou, China) and ligated into pHT304-18Z. The recombinant plasmids named pHT-PsoxBM was introduced into Bt strain HD73, yielding the transformant strains HD(PsoxBM).
Construction of the soxR and soxB mutants
To construct the soxR deletion mutant, DNA fragments corresponding to the upstream and downstream regions of soxR were first PCR-amplified from genomic Bt HD73 DNA with the soxR-a/soxR-d and soxR-b/soxR-c primers. The amplified fragments were then fused via overlapping PCR using the soxR-a/soxR-b primers. The resultant 1257-bp fragment was then digested with BamHI and EcoRI and ligated into pMAD. The recombinant pMAD∆soxR plasmid was electroporated into host strains and erythromycin-sensitive transformants were selected. Transformants were verified by culturing at 39 °C–41 °C. Colonies lacking erythromycin resistance were selected and one mutant strain, ΔsoxR, was verified by PCR.
The upstream and downstream regions (fragments A and B, respectively) of the soxB gene were PCR-amplified from Bt with primers soxB-a/soxB-d and soxB-c/soxB-b. The kanamycin resistance cassette (kan) was PCR-amplified from pDG780 with primers soxB-kmF/soxB-kmR. Fragment A and kan were ligated by overlapping PCR with primers soxB-a and soxB-kmR. The amplification product was then integrated with fragment B in a second round of overlapping PCR using the soxB-a and soxB-b primers. The resultant PCR products were digested, purified and ligated with pMAD to yield pMADΔsoxB, which was used to transform the host strain by electroporation, followed by selection of erythromycin-sensitive transformants. Transformants were verified by culturing at 39 °C–41 °C. Colonies with kanamycin resistance but lacking erythromycin resistance were selected and one mutant strain, ΔsoxB, was verified by PCR.
Complementation of the soxR mutant
A DNA fragment containing soxR and the soxR promoter was amplified with CsoxR-1 and CsoxR-2 primers (Table S3) using Bt strain HD73 DNA as template. The PCR product (2,268 bp) was digested with PstI and XbaI and ligated into plasmid pHT161855. The resulting plasmid (pHT1618-soxR) was amplified in E. coli and introduced into the Bt mutant strain ΔsoxR(PsoxB-lacZ) and the new strain named CsoxR(PsoxB). This plasmid complements the soxR mutant strain and allows evaluation of the expression of the soxB promoter-lacZ fusion.
Additional Information
How to cite this article: Peng, Q. et al. Sox transcription in sarcosine utilization is controlled by Sigma54 and SoxR in Bacillus thuringiensis HD73. Sci. Rep. 6, 29141; doi: 10.1038/srep29141 (2016).
References
Kim, M. J., Jung, W. H. & Koo, J. S. Expression of sarcosine metabolism-related proteins in estrogen receptor negative breast cancer according to the androgen receptor and HER-2 status. International journal of clinical and experimental pathology 8, 7967–7977 (2015).
Suzuki, M. Purification and some properties of sarcosine oxidase from Corynebacterium sp. U-96. Journal of biochemistry 89, 599–607 (1981).
Willie, A., Edmondson, D. E. & Jorns, M. S. Sarcosine oxidase contains a novel covalently bound FMN. Biochemistry 35, 5292–5299, 10.1021/bi952995h (1996).
Meskys, R., Harris, R. J., Casaite, V., Basran, J. & Scrutton, N. S. Organization of the genes involved in dimethylglycine and sarcosine degradation in Arthrobacter spp.: implications for glycine betaine catabolism. J. Biol. Chem 268, 3390–3398 (2001).
Nishiya, Y., Toda, A. & Imanaka, T. Gene cluster for creatinine degradation in Arthrobacter sp. TE1826. Mol. Gen. Genet. 257, 581–586 (1998).
Tsuru, D. On the catabolism of creatinine and the related enzymes in microorganisms. Nucleic Acids Amino Acids 35, 31–37 (1977).
Nishiya, Y. & Imanaka, T. Analysis of a negative regulator, soxR, for the Arthrobacter sarcosine oxidase gene. J. Ferment. Bioeng. 81, 64–67 (1996).
Suzuki, K., Sagai, H., Imamura, S. & Sugiyama, M. Cloning, sequencing and overexpression in Escherichia coli of a sarcosine oxidase-containing gene linked to the Bacillus creatinase gene. J. Ferment. Bioeng. 77, 231–234 (1994).
Chlumsky, L. J., Zhang, L. & Jorns, M. S. Sequence analysis of sarcosine oxidase and nearby genes reveals homologies with key enzymes of folate one-carbon metabolism. J. Biol. Chem 270, 18252–18259 (1995).
Suzuki, H., Tamamura, R., Yajima, S., Kanno, M. & Suguro, M. Corynebacterium sp. U-96 contains a cluster of genes of enzymes for the catabolism of sarcosine to pyruvate. Biosci. Biotechnol. Biochem. 69, 952–956 (2005).
Helgason, E. et al. Bacillus anthracis, Bacillus cereus and Bacillus thuringiensis–one species on the basis of genetic evidence. Applied and environmental microbiology 66, 2627–2630 (2000).
Han, C. S. et al. Pathogenomic sequence analysis of Bacillus cereus and Bacillus thuringiensis isolates closely related to Bacillus anthracis. Journal of bacteriology 188, 3382–3390, 10.1128/JB.188.9.3382-3390.2006 (2006).
Ivanova, N. et al. Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature 423, 87–91, 10.1038/nature01582 (2003).
Fenton, M. S., Lee, S. J. & Gralla, J. D. Escherichia coli promoter opening and -10 recognition: mutational analysis of sigma70. The EMBO journal 19, 1130–1137, 10.1093/emboj/19.5.1130 (2000).
Lonetto, M., Gribskov, M. & Gross, C. The sigma 70 family: sequence conservation and evolutionary relationships. J. Bacteriol 174, 3843–3849 (1992).
Wigneshweraraj, S. et al. Modus operandi of the bacterial RNA polymerase containing the sigma54 promoter-specificity factor. Mol. Microbiol. 68, 538–546, 10.1111/j.1365-2958.2008.06181.x (2008).
Merrick, M. J. In a class of its own–the RNA polymerase sigma factor sigma 54 (sigma N). Mol. Microbiol 10, 903–909 (1993).
Buck, M., Gallegos, M. T., Studholme, D. J., Guo, Y. & Gralla, J. D. The bacterial enhancer-dependent sigma(54) (sigma(N)) transcription factor. J. Bacteriol 182, 4129–4136 (2000).
Zhao, K., Liu, M. & Burgess, R. R. Promoter and regulon analysis of nitrogen assimilation factor, sigma54, reveal alternative strategy for E. coli MG1655 flagellar biosynthesis. Nucl. Acids Res. 38, 1273–1283, 10.1093/nar/gkp1123 (2010).
Bernard, C. S., Brunet, Y. R., Gavioli, M., Lloubes, R. & Cascales, E. Regulation of type VI secretion gene clusters by sigma54 and cognate enhancer binding proteins. J. Bacteriol 193, 2158–2167, 10.1128/JB.00029-11 (2011).
Massimelli, M. J. et al. Choline catabolism, sigma(5)(4) factor and NtrC are required for the full expression of the Pseudomonas aeruginosa phosphorylcholine phosphatase gene. Microbiol. Res. 166, 380–390, 10.1016/j.micres.2010.07.004 (2011).
Debarbouille, M., Martin-Verstraete, I., Kunst, F. & Rapoport, G. The Bacillus subtilis sigL gene encodes an equivalent of sigma 54 from gram-negative bacteria. Proc. Natl. Acad. Sci. USA 88, 9092–9096 (1991).
Gardan, R., Rapoport, G. & Debarbouille, M. Expression of the rocDEF operon involved in arginine catabolism in Bacillus subtilis. J. Mol. Biol. 249, 843–856, 10.1006/jmbi.1995.0342 (1995).
Zhu, L. et al. Structure and regulation of the gab gene cluster, involved in the gamma-aminobutyric acid shunt, are controlled by a sigma54 factor in Bacillus thuringiensis. Journal of bacteriology 192, 346–355, 10.1128/JB.01038-09 (2010).
Zhang, Z. et al. Transcription of the lysine-2,3-aminomutase gene in the kam locus of Bacillus thuringiensis subsp. kurstaki HD73 is controlled by both sigma54 and sigmaK factors. Journal of bacteriology 196, 2934–2943, 10.1128/JB.01675-14 (2014).
Liu, G. et al. Complete genome sequence of Bacillus thuringiensis subsp. kurstaki strain HD73. Genome announcements 1, e0008013, 10.1128/genomeA.00080-13 (2013).
Francke, C. et al. Comparative analyses imply that the enigmatic Sigma factor 54 is a central controller of the bacterial exterior. BMC genomics 12, 385, 10.1186/1471-2164-12-385 (2011).
Peng, Q., Wang, G., Liu, G., Zhang, J. & Song, F. Identification of metabolism pathways directly regulated by sigma(54) factor in Bacillus thuringiensis. Frontiers in microbiology 6, 407, 10.3389/fmicb.2015.00407 (2015).
Tsuru, D. O., I. Yoshimoto & T. Creatinine decomposing enzymes in Pseudomonas putida. Agric. Biol. Chem. 40, 1011–1018 (1976).
Meskys, R. Screening and Investigation of Hydrogen-Peroxide-Producing Microbial Oxidase. Vilnius University, Vilnius, Lithuania. (1998).
Meskys, R., Rudomanskis, R., Semsnaits, R. & Trainys, M. Peculiarities of creatinine catabolism in some Arthrobacter strains. Biology 1, 61–64 (1997).
Willsey, G. G. & Wargo, M. J. Sarcosine Catabolism in Pseudomonas aeruginosa Is Transcriptionally Regulated by SouR. Journal of bacteriology 198, 301–310, 10.1128/JB.00739-15 (2016).
Rappas, M., Bose, D. & Zhang, X. Bacterial enhancer-binding proteins: unlocking sigma54-dependent gene transcription. Curr. Opin. Struct. Biol. 17, 110–116, 10.1016/j.sbi.2006.11.002 (2007).
Shingler, V. Signal sensory systems that impact sigma54 -dependent transcription. FEMS Microbiol. Rev. 35, 425–440, 10.1111/j.1574-6976.2010.00255.x (2011).
Debarbouille, M., Martin-Verstraete, I., Klier, A. & Rapoport, G. The transcriptional regulator LevR of Bacillus subtilis has domains homologous to both sigma 54- and phosphotransferase system-dependent regulators. Proc. Natl. Acad. Sci. USA 88, 2212–2216 (1991).
Debarbouille, M., Gardan, R., Arnaud, M. & Rapoport, G. Role of bkdR, a transcriptional activator of the sigL-dependent isoleucine and valine degradation pathway in Bacillus subtilis. J. Bacteriol. 181, 2059–2066 (1999).
Ali, N. O., Bignon, J., Rapoport, G. & Debarbouille, M. Regulation of the acetoin catabolic pathway is controlled by sigma L in Bacillus subtilis. J. Bacteriol. 183, 2497–2504, 10.1128/JB.183.8.2497-2504.2001 (2001).
Calogero, S. et al. RocR, a novel regulatory protein controlling arginine utilization in Bacillus subtilis, belongs to the NtrC/NifA family of transcriptional activators. J. Bacteriol 176, 1234–1241 (1994).
Belitsky, B. R. & Sonenshein, A. L. An enhancer element located downstream of the major glutamate dehydrogenase gene of Bacillus subtilis. Proc. Natl. Acad. Sci. USA 96, 10290–10295 (1999).
Ali, N. O. et al. Specificity of the interaction of RocR with the rocG-rocA intergenic region in Bacillus subtilis. Microbiology 149, 739–750 (2003).
Reitzer, L. J. & Magasanik, B. Transcription of glnA in E. coli is stimulated by activator bound to sites far from the promoter. Cell 45, 785–792 (1986).
Marchler-Bauer, A. et al. CDD: conserved domains and protein three-dimensional structure. Nucl. Acids Res. 41, 348–352 (2013).
Todone, F. et al. Active site plasticity in D-amino acid oxidase: a crystallographic analysis. Biochemistry 36, 5853–5860, 10.1021/bi9630570 (1997).
Martinez-Perez, O., Moreno-Ruiz, E., Floriano, B. & Santero, E. Regulation of tetralin biodegradation and identification of genes essential for expression of thn operons. J. Bacteriol. 186, 6101–6109, 10.1128/JB.186.18.6101-6109.2004 (2004).
Han, C. S. et al. Pathogenomic sequence analysis of Bacillus cereus and Bacillus thuringiensis isolates closely related to Bacillus anthracis. J. Bacteriol 188, 3382–3390, 10.1128/JB.188.9.3382-3390.2006 (2006).
Read, T. D. et al. The genome sequence of Bacillus anthracis Ames and comparison to closely related bacteria. Nature 423, 81–86, 10.1038/nature01586 (2003).
Lereclus, D., Arantes, O., Chaufaux, J. & Lecadet, M. Transformation and expression of a cloned delta-endotoxin gene in Bacillus thuringiensis. FEMS microbiology letters 51, 211–217 (1989).
Schaeffer, P., Millet, J. & Aubert, J. P. Catabolic repression of bacterial sporulation. Proceedings of the National Academy of Sciences of the United States of America 54, 704–711 (1965).
Debarbouille, M., Martin-Verstraete, I., Kunst, F. & Rapoport, G. The Bacillus subtilis sigL gene encodes an equivalent of sigma 54 from gram-negative bacteria. Proc Natl Acad Sci USA 88, 9092–9096 (1991).
Du, L. et al. Identification of the Promoter in the Intergenic Region between orf1 and cry8Ea1 Controlled by Sigma H Factor. Appl. Environ. Microbiol. 78, 4164–4168 (2012).
Agaisse, H. & Lereclus, D. Structural and functional analysis of the promoter region involved in full expression of the cryIIIA toxin gene of Bacillus thuringiensis. Molecular microbiology 13, 97–107 (1994).
Perchat, S. et al. A cell-cell communication system regulates protease production during sporulation in bacteria of the Bacillus cereus group. Molecular microbiology 82, 619–633, 10.1111/j.1365-2958.2011.07839.x (2011).
Peng, Q. et al. Activation of gab cluster transcription in Bacillus thuringiensis by gamma-aminobutyric acid or succinic semialdehyde is mediated by the Sigma 54-dependent transcriptional activator GabR. BMC microbiology 14, 306, 10.1186/s12866-014-0306-3 (2014).
Li, R. et al. PolY, a transcriptional regulator with ATPase activity, directly activates transcription of polR in polyoxin biosynthesis in Streptomyces cacaoi. Molecular microbiology 75, 349–364, 10.1111/j.1365-2958.2009.06968.x (2010).
Lereclus, D. & Arantes, O. spbA locus ensures the segregational stability of pTH1030, a novel type of gram-positive replicon. Molecular microbiology 6, 35–46 (1992).
Acknowledgements
This work was supported by a grant from the National Natural Science Foundation of China (Nos 31270111 and 31300085). The authors would like to thank Prof. Linquan Bai from Shanghai Jiao Tong University for his helpful suggestions and Dr. Didier Lereclus from the Institut National de la Recherche Agronomique for his critical review.
Author information
Authors and Affiliations
Contributions
F.S. and Q.P. designed the research. Q.P. drafted the manuscript. C.L., B.W., M.Y. and J.W. performed the experimental work. F.S. and J.Z. critically revised the manuscript for intellectual content. All authors read and approved the final version of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Peng, Q., Liu, C., Wang, B. et al. Sox transcription in sarcosine utilization is controlled by Sigma54 and SoxR in Bacillus thuringiensis HD73. Sci Rep 6, 29141 (2016). https://doi.org/10.1038/srep29141
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29141
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.